

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.2 - phase: 0 /pseudo
(461 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
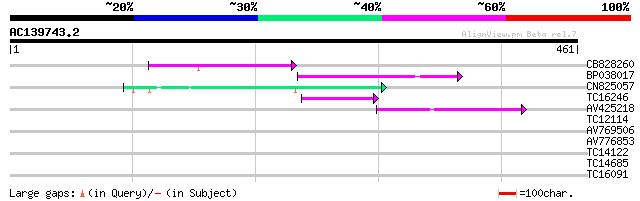
Score E
Sequences producing significant alignments: (bits) Value
CB828260 74 7e-14
BP038017 71 4e-13
CN825057 44 4e-05
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 44 7e-05
AV425218 42 2e-04
TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transp... 36 0.015
AV769506 28 2.4
AV776853 27 5.4
TC14122 similar to UP|Q9FE12 (Q9FE12) Peroxiredoxin precursor, c... 27 7.1
TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 27 9.3
TC16091 27 9.3
>CB828260
Length = 557
Score = 73.6 bits (179), Expect = 7e-14
Identities = 38/122 (31%), Positives = 64/122 (52%), Gaps = 2/122 (1%)
Frame = +3
Query: 114 LVCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQ--TKEWGLNILNGVHNHPMEP 171
L CE+ G ++ T ++K C F + G K ++W L ++ G HNH
Sbjct: 192 LGCEKHGKYVPYRDPDLVEGTRTQKTECPFRLKGRPMKNGTDRDWRLKVMEGAHNHEPAR 371
Query: 172 ALEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKA 231
+L GH GRL ++K+ V ++KS +LPR +L+ LK P +T + Q+Y +++ K+
Sbjct: 372 SLLGHNFVGRLNSEEKEQVEKMSKSWVLPRKMLLTLKENNPLNLTTISQIYGACKRLRKS 551
Query: 232 NR 233
R
Sbjct: 552 LR 557
>BP038017
Length = 570
Score = 70.9 bits (172), Expect = 4e-13
Identities = 42/136 (30%), Positives = 67/136 (48%), Gaps = 2/136 (1%)
Frame = +2
Query: 235 DKKSLQFLISKLEEHNYTYYSRTQLES-NTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKT 293
D ++L K++ N +Y QL+ N + ++FWA S +N F ++ D+ Y+
Sbjct: 167 DAQNLLNYFKKMQGKNPGFYYAIQLDDENRMINVFWADARSRSAYNYFGDAVIFDTMYRP 346
Query: 294 NMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKV-IVTD 352
N Y++P GV G + E E +F W + R LS+ + P V I TD
Sbjct: 347 NQYQVPFAPFTGVNHHGQNVLFGCALLLDESESSFTW---LFRTWLSAMNDRPPVSITTD 517
Query: 353 RDMSLMKAVAHVFPES 368
+D ++ AVA VFPE+
Sbjct: 518 QDRAIQAAVAQVFPET 565
>CN825057
Length = 721
Score = 44.3 bits (103), Expect = 4e-05
Identities = 52/227 (22%), Positives = 89/227 (38%), Gaps = 13/227 (5%)
Frame = +1
Query: 93 ANKAGFT--IVTQRSSLINPMF---RLVCERSGSHIVPEKKP-KHANTGSRKCGCLFMIS 146
A GFT I R S F + C R G + PE + + +K C +
Sbjct: 25 AKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYG--VTPESDGGSNRRSSVKKTDCKACMH 198
Query: 147 GYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGR-LKEDDKKIVRDLTKSKMLPRNILI 205
+ K +W ++ HNH + PAL H R +K +K + L R + +
Sbjct: 199 -VKRKPDGKWIIHEFIKEHNHELLPALAYHFRIHRNVKLAEKNNMDILHAVSERTRKMYV 375
Query: 206 HLKNQRPHCMTNVKQVYNERQQIWKA-----NRGDKKSLQFLISKLEEHNYTYYSRTQL- 259
+ Q C+ V + Q K + GD + + +++ N ++ L
Sbjct: 376 EMSRQSGGCLNIESLVGDLNDQFKKGQYLAMDEGDAQVMLEYFKHIQKENPNFFYSIDLN 555
Query: 260 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGV 306
E + +IFW SI + +F V+ D++Y + ++P VGV
Sbjct: 556 EEQRLRNIFWIDAKSINDYLSFNDVVSFDTSYIKSNEKLPFAPFVGV 696
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 43.5 bits (101), Expect = 7e-05
Identities = 20/63 (31%), Positives = 32/63 (50%)
Frame = +2
Query: 238 SLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYR 297
+L +L K + +Y T+ ++E++FW S +N F V+ DSTYK N Y
Sbjct: 161 ALSYLEGKADNDPMFFYKFTKTGDESLENLFWCDGVSRMDYNVFGDVIAFDSTYKKNKYN 340
Query: 298 MPM 300
P+
Sbjct: 341 KPL 349
>AV425218
Length = 419
Score = 42.4 bits (98), Expect = 2e-04
Identities = 29/122 (23%), Positives = 53/122 (42%)
Frame = +2
Query: 299 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLM 358
P+ +VGV + G ++ E E+FVW+ L +S P+ I+TD ++
Sbjct: 5 PLATLVGVNHHGQSVLFGCALLSSEDSESFVWLFQSLLHCMSGV--PPQGIITDHSEAMK 178
Query: 359 KAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRDVVKKIMNAWKAM 418
KA+ V P + C ++ + Q+ + +Y V + V+ + WK +
Sbjct: 179 KAIETVLPSTRHRWCLSYIMKKLPQKLLGYAQYESIRHHLQNVVYDAVVIDEFERNWKKI 358
Query: 419 VE 420
VE
Sbjct: 359 VE 364
>TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transposase-like
{Oryza sativa (japonica cultivar-group);}, partial (8%)
Length = 488
Score = 35.8 bits (81), Expect = 0.015
Identities = 21/93 (22%), Positives = 41/93 (43%)
Frame = +1
Query: 239 LQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRM 298
LQ+ KL E++ Y++ + + I ++FW + + F ++ +DSTY T+
Sbjct: 58 LQYFQRKLVENSPFYHAYQLDDEDQITNVFWVDARMLIDYGYFGDMVSLDSTYCTHSSNR 237
Query: 299 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWV 331
P+ G G + E E++ W+
Sbjct: 238 PLAVFSGFNHHRKAVIFGAALLYDETTESY*WL 336
>AV769506
Length = 345
Score = 28.5 bits (62), Expect = 2.4
Identities = 24/97 (24%), Positives = 41/97 (41%), Gaps = 13/97 (13%)
Frame = +1
Query: 82 RDELLEWVRRQANKAGFTIVTQRSSLINPMFR-----LVCERSGSHI-------VPEKKP 129
RD+L+ WV A + G+ + +S R L CE+ G ++ V ++ P
Sbjct: 52 RDDLINWVHGIAIENGYVTLITKSDYGGNESRKAYVMLGCEKHGKYVPYRDPDLVADEDP 231
Query: 130 KHANTGSRKCGCLFMISGYQSKQTKE-WGLNILNGVH 165
+T C CL SK + + W + ++G H
Sbjct: 232 SAQSTLEPACDCLNHHHKQGSKPSPQLWPPSTVDGPH 342
>AV776853
Length = 625
Score = 27.3 bits (59), Expect = 5.4
Identities = 24/95 (25%), Positives = 35/95 (36%), Gaps = 9/95 (9%)
Frame = +1
Query: 111 MFRLVCERSGSHIVPEKKPKHANTGSRK--------CGCLFMISGYQSKQTKEWGLNILN 162
M + VC R G + KH N RK CL + + + +W +
Sbjct: 352 MRQFVCNRQGL-----RSKKHYNRTDRKRDHKPVTHTNCLAKLRVHLDYKIGKWKVVSFE 516
Query: 163 GVHNHPMEPALEGHILAG-RLKEDDKKIVRDLTKS 196
HNH + PA H + R+ D K D+ S
Sbjct: 517 ECHNHELTPARFVHFIPPYRVMNDADKAKGDMLHS 621
>TC14122 similar to UP|Q9FE12 (Q9FE12) Peroxiredoxin precursor, complete
Length = 1107
Score = 26.9 bits (58), Expect = 7.1
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = +1
Query: 31 KPPNFEAKTNIDEASVHVEASKYVGGSGVL 60
K P+FEA+ D+ + V+ S+Y+G V+
Sbjct: 298 KAPDFEAEAVFDQEFIKVKLSEYIGKKYVI 387
>TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (38%)
Length = 1323
Score = 26.6 bits (57), Expect = 9.3
Identities = 23/115 (20%), Positives = 50/115 (43%), Gaps = 2/115 (1%)
Frame = +1
Query: 79 SKNRDELLEWVRRQANKAGFTIVTQRSSLINPMFRLVCERSGSHIVPEKKPKHANTGSRK 138
+++R +++ +RR A+ GF V ++ + R V H PE++ T
Sbjct: 325 NESRAAVVDQIRRAASSVGFFQVVNHGVPLDLLRRTVAAIKAFHEQPEEERARVYTREMG 504
Query: 139 CGCLFM--ISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVR 191
G ++ + +QSK W + + ++ ++ + + E DK++VR
Sbjct: 505 TGVSYISNVDLFQSK-AASWRDTLQLRMGPTAVDQSMIPEVCRQEVMEWDKEVVR 666
>TC16091
Length = 627
Score = 26.6 bits (57), Expect = 9.3
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Frame = +3
Query: 86 LEWVRRQANKA-GFTI--VTQRSSLINPMFRLVCERSGSHIVPEKKPKHAN 133
L WV + G TI VT R S + R+VC G+H+ PE+ +HA+
Sbjct: 147 LPWVSTSGSGPNGRTISGVTYRYS--SNQIRIVCACHGTHLTPEEFVRHAS 293
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,922,536
Number of Sequences: 28460
Number of extensions: 104021
Number of successful extensions: 505
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 502
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 503
length of query: 461
length of database: 4,897,600
effective HSP length: 93
effective length of query: 368
effective length of database: 2,250,820
effective search space: 828301760
effective search space used: 828301760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC139743.2


