

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
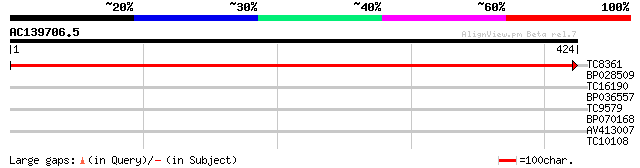
Query= AC139706.5 - phase: 0
(424 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8361 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial... 780 0.0
BP028509 32 0.20
TC16190 weakly similar to UP|Q9LQ75 (Q9LQ75) T1N6.22 protein, pa... 28 2.2
BP036557 27 6.4
TC9579 similar to UP|DCUP_TOBAC (Q42967) Uroporphyrinogen decarb... 27 8.3
BP070168 27 8.3
AV413007 27 8.3
TC10108 similar to GB|AAO63297.1|28950747|BT005233 At2g45360 {Ar... 27 8.3
>TC8361 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial (92%)
Length = 1523
Score = 780 bits (2015), Expect = 0.0
Identities = 391/424 (92%), Positives = 403/424 (94%)
Frame = +3
Query: 1 MANLSMTQLKTRNSASRFSFNFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPN 60
MA+LSMTQ++T SAS FS N KV RPSRV F+VFASE QV EPDLSV VNGLHMPN
Sbjct: 72 MASLSMTQVRTGKSASGFSLNCPGKVQARPSRVGFRVFASEAQVPEPDLSVTVNGLHMPN 251
Query: 61 PFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGE 120
PFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSLDAAKVINVTPRYARLRA ANGSAKGE
Sbjct: 252 PFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLDAAKVINVTPRYARLRAGANGSAKGE 431
Query: 121 IIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGID 180
IIGW+NIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGID
Sbjct: 432 IIGWENIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGID 611
Query: 181 AIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARV 240
AIEINFSCPHGMPERKMGAAVGQDCALLEEVC WINAKATVPVWAKMTPNITDIS+PARV
Sbjct: 612 AIEINFSCPHGMPERKMGAAVGQDCALLEEVCRWINAKATVPVWAKMTPNITDISEPARV 791
Query: 241 ALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKM 300
ALSSGC GVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIAL KVMSIAKM
Sbjct: 792 ALSSGC*GVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALAKVMSIAKM 971
Query: 301 MKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFM 360
MKSEFDSE+YSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKL +EL+DFM
Sbjct: 972 MKSEFDSEDYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLCVELQDFM 1151
Query: 361 KKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESES 420
KKHNFTSIEDFRG SLQYFTTHTDLV RQQEAI+QRKAI+KGLQSD+DWTGDGFVKESES
Sbjct: 1152KKHNFTSIEDFRGTSLQYFTTHTDLVQRQQEAIRQRKAIRKGLQSDRDWTGDGFVKESES 1331
Query: 421 MVSN 424
MVSN
Sbjct: 1332MVSN 1343
>BP028509
Length = 551
Score = 32.0 bits (71), Expect = 0.20
Identities = 24/82 (29%), Positives = 42/82 (50%), Gaps = 3/82 (3%)
Frame = -3
Query: 95 DAAKVINVTPRYARLRANANGSAKGEIIGWQNI-ELISDRPLETMLKEFKQ--LKEEYPD 151
D+ +++NV+ +L NG AKG + N+ E D L+ LK+F+Q L+ +
Sbjct: 549 DSPRIVNVSSSLGKLECVPNGWAKGVLSDVDNLTEEAVDGVLKRFLKDFQQGSLESKGWP 370
Query: 152 RILIASIMEEYNKAAWEELIDR 173
+ L A IM + A+ ++ R
Sbjct: 369 KTLSAYIMSKAAMNAYTRILAR 304
>TC16190 weakly similar to UP|Q9LQ75 (Q9LQ75) T1N6.22 protein, partial (41%)
Length = 1221
Score = 28.5 bits (62), Expect = 2.2
Identities = 27/98 (27%), Positives = 46/98 (46%), Gaps = 3/98 (3%)
Frame = +1
Query: 95 DAAKVINVTPRYARLRANANGSAKGEIIGWQNI-ELISDRPLETMLKEFK--QLKEEYPD 151
D+ +++NV+ +L NG AKG + N+ E D L+ L++F+ L+ +
Sbjct: 16 DSPRIVNVSSFLGKLECVPNGWAKGVLSDADNLTEETVDGVLKKFLQDFQDGSLEGKGWP 195
Query: 152 RILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCP 189
+I A IM + A+ ++ R T IN CP
Sbjct: 196 KIFSAYIMSKAAMNAYTRILARKYPT----FCINSVCP 297
>BP036557
Length = 369
Score = 26.9 bits (58), Expect = 6.4
Identities = 17/73 (23%), Positives = 37/73 (50%)
Frame = +2
Query: 140 KEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGA 199
K K L + +P+++ I +E A+W++ +DR ++ ++I + H R + +
Sbjct: 116 KPNKTLTKLFPNKVTIHMPQDETLLASWKDQLDR----DVETLKIKGNLHH---LRTVLS 274
Query: 200 AVGQDCALLEEVC 212
G +C L+ +C
Sbjct: 275 RCGMECEGLDTLC 313
>TC9579 similar to UP|DCUP_TOBAC (Q42967) Uroporphyrinogen decarboxylase,
chloroplast precursor (URO-D) (UPD) , partial (31%)
Length = 552
Score = 26.6 bits (57), Expect = 8.3
Identities = 11/34 (32%), Positives = 20/34 (58%)
Frame = +3
Query: 35 FKVFASEGQVSEPDLSVKVNGLHMPNPFVIGSGP 68
+KVF +G + D+ ++G+++P V G GP
Sbjct: 396 WKVFKPDGVILFSDILTPLSGMNIPFDIVKGKGP 497
>BP070168
Length = 434
Score = 26.6 bits (57), Expect = 8.3
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -1
Query: 56 LHMPNPFVIGSGPPGTNYTVMKRAF 80
L +PNP ++ GPP + + M AF
Sbjct: 374 LAIPNPLLVAIGPPSDHVSTMSCAF 300
>AV413007
Length = 416
Score = 26.6 bits (57), Expect = 8.3
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +2
Query: 19 SFNFSKKVYPRPSRVDFKVFASEG 42
SFN+ KK+ P+ F VF EG
Sbjct: 293 SFNYDKKIAPKEGTKPFIVFIGEG 364
>TC10108 similar to GB|AAO63297.1|28950747|BT005233 At2g45360 {Arabidopsis
thaliana;}, partial (38%)
Length = 391
Score = 26.6 bits (57), Expect = 8.3
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -3
Query: 56 LHMPNPFVIGSGPPGTNYTVMKRAF 80
L +PNP ++ GPP + + M AF
Sbjct: 281 LAIPNPLLVAIGPPSDHVSTMSCAF 207
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,217,532
Number of Sequences: 28460
Number of extensions: 74988
Number of successful extensions: 352
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 349
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 352
length of query: 424
length of database: 4,897,600
effective HSP length: 93
effective length of query: 331
effective length of database: 2,250,820
effective search space: 745021420
effective search space used: 745021420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC139706.5


