

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139600.8 + phase: 0
(318 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
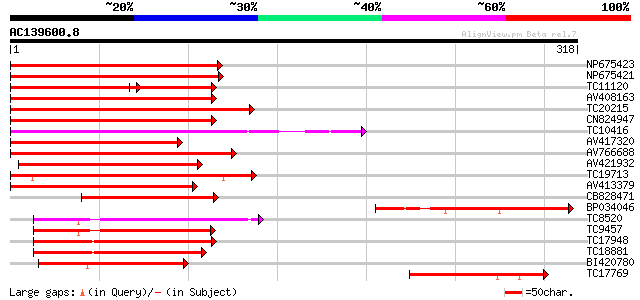
Score E
Sequences producing significant alignments: (bits) Value
NP675423 transcription factor MYB101 [Lotus japonicus] 188 9e-49
NP675421 transcription factor MYB103 [Lotus japonicus] 178 1e-45
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 116 3e-43
AV408163 167 3e-42
TC20215 similar to AAS10104 (AAS10104) MYB transcription factor,... 166 4e-42
CN824947 164 2e-41
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 160 2e-40
AV417320 158 1e-39
AV766688 156 4e-39
AV421932 147 3e-36
TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%) 144 3e-35
AV413379 142 6e-35
CB828471 118 2e-27
BP034046 113 4e-26
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 108 9e-25
TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 107 3e-24
TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose... 104 2e-23
TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose... 102 1e-22
BI420780 93 7e-20
TC17769 similar to UP|Q40175 (Q40175) Transcription factor, part... 68 2e-12
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 188 bits (478), Expect = 9e-49
Identities = 82/119 (68%), Positives = 102/119 (84%)
Frame = +1
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCD+ G+KKGPWTPEED ILV YIQ+HG G+W+A+P GL RC KSCRLRWTNY
Sbjct: 1 MGRTPCCDEIGLKKGPWTPEEDAILVDYIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNY 180
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL 119
LRP IKRG FT++EE++II+L A+LGN+W+AIA +LP RTDN+IKN+WNT+LKKKL ++
Sbjct: 181 LRPDIKRGKFTEEEEQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKKLLQM 357
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 178 bits (451), Expect = 1e-45
Identities = 79/120 (65%), Positives = 99/120 (81%)
Frame = +1
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M+R P D+ G+KKGPW+ EED +LV YIQ+HG G+W+A+P GL RC KSCRLRWTNY
Sbjct: 1 MMRTPSIDESGMKKGPWSQEEDKVLVDYIQKHGHGSWRALPQLAGLNRCGKSCRLRWTNY 180
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRP IKRG F+D+EE++II+L A LGN+WA IA++LP RTDN+IKN WNT+LKKKLK L+
Sbjct: 181 LRPDIKRGKFSDEEEQLIINLHASLGNKWATIASHLPGRTDNEIKNLWNTHLKKKLKLLQ 360
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 116 bits (291), Expect(2) = 3e-43
Identities = 50/74 (67%), Positives = 59/74 (79%)
Frame = +1
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCD+ G+KKGPWTPEED LV +IQ+HG G+W+A+P GL RC KSCRLRWTNY
Sbjct: 67 MGRSPCCDENGLKKGPWTPEEDQKLVEHIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNY 246
Query: 61 LRPGIKRGNFTDQE 74
LRP IKRG F+ +E
Sbjct: 247 LRPDIKRGKFSPEE 288
Score = 75.1 bits (183), Expect(2) = 3e-43
Identities = 30/49 (61%), Positives = 42/49 (85%)
Frame = +2
Query: 68 GNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL 116
GNF ++E+ I+HL ++LGN+W+ IA +LP RTDN+IKN+WNT+LKKKL
Sbjct: 269 GNFLQKKEQTILHLHSILGNKWSTIATHLPGRTDNEIKNFWNTHLKKKL 415
>AV408163
Length = 425
Score = 167 bits (422), Expect = 3e-42
Identities = 71/116 (61%), Positives = 92/116 (79%)
Frame = +2
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K + KG WT EED L+SYI+ HG G W+++P GLLRC KSCRLRW NY
Sbjct: 77 MGRSPCCEKAHINKGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 256
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL 116
LRP +KRGNFT++E+++II L +LLGN+W+ IA LP RTDN+IKNYWNT++++KL
Sbjct: 257 LRPDLKRGNFTEEEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKL 424
>TC20215 similar to AAS10104 (AAS10104) MYB transcription factor, partial
(35%)
Length = 464
Score = 166 bits (421), Expect = 4e-42
Identities = 72/137 (52%), Positives = 101/137 (73%)
Frame = +2
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K G+K+G W+ EED IL +I+++G G+WK++P N GLLRC KSCRLRW NY
Sbjct: 29 MGRAPCCEKVGLKRGRWSEEEDKILTDFIKQNGEGSWKSLPKNAGLLRCGKSCRLRWINY 208
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LR +KRGN T +EE++I+ L A LGNRW+ IA +LP RTDN+IKNYWN++L++K+
Sbjct: 209 LREDVKRGNITSEEEEIIVKLHAALGNRWSVIAGHLPGRTDNEIKNYWNSHLRRKIYCFM 388
Query: 121 TSTSSESCLGHDEFSVS 137
S + + L + +V+
Sbjct: 389 RSLNDQQSLPPIDMNVA 439
>CN824947
Length = 738
Score = 164 bits (414), Expect = 2e-41
Identities = 69/116 (59%), Positives = 90/116 (77%)
Frame = +1
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K G+K+G WT EED +L YIQ G G+W+++P N GLLRC KSCRLRW NY
Sbjct: 151 MGRAPCCEKVGLKRGRWTAEEDELLTKYIQASGEGSWRSLPKNAGLLRCGKSCRLRWINY 330
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL 116
LR +KRGN +D+EE +I+ L A GNRW+ IA+++P RTDN+IKNYWN++L +KL
Sbjct: 331 LRGDLKRGNISDEEESLIVKLHASFGNRWSLIASHMPGRTDNEIKNYWNSHLSRKL 498
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 160 bits (406), Expect = 2e-40
Identities = 87/201 (43%), Positives = 120/201 (59%), Gaps = 1/201 (0%)
Frame = +2
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R P CDK G++KG WT EED L++Y+ +G NW+ +P GL RC KSCRLRW NY
Sbjct: 92 MARTPSCDKSGMRKGTWTAEEDKKLIAYVTRYGCWNWRQLPKFAGLARCGKSCRLRWLNY 271
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRP IKRGNFT +EE++II + LGNRW+ IAA LP RTD+++KN+W+T LK+ + +
Sbjct: 272 LRPNIKRGNFTQEEEEIIIRMHKKLGNRWSTIAAELPGRTDSEVKNHWHTTLKRTVHEQN 451
Query: 121 TSTSSESCLGHDEFSVSQPIARGQWERRLQTDIHMAKKALSEALQPEKSTSSSNLMLPLE 180
T T+ E+ + + + + P+ + L T P +TS S + L
Sbjct: 452 TVTNEETKVSNSK-NERDPVPKSGIASLLVT--------------PPPATSISGSIGAL- 583
Query: 181 SNFSSESSF-CSTKPTTQSLS 200
S FS+ S F C+T T S S
Sbjct: 584 SPFSTSSEFSCTTSDLTTSSS 646
>AV417320
Length = 369
Score = 158 bits (399), Expect = 1e-39
Identities = 70/97 (72%), Positives = 82/97 (84%)
Frame = +3
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
MVR PCC+K G+KKGPWT EED IL+SYIQ+HG GNW+A+P + GLLRC KSCRLRW NY
Sbjct: 78 MVRAPCCEKIGLKKGPWTSEEDQILISYIQKHGHGNWRALPKHAGLLRCGKSCRLRWINY 257
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLP 97
LRP IKRGNFT +EE++II + LLGNRW+AIAA LP
Sbjct: 258 LRPDIKRGNFTAEEEELIIKMHELLGNRWSAIAAKLP 368
>AV766688
Length = 608
Score = 156 bits (395), Expect = 4e-39
Identities = 69/127 (54%), Positives = 93/127 (72%)
Frame = +2
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R P CDK G++KG WT EED L++Y+ +G NW+ +P GL RC KSCRLRW NY
Sbjct: 77 MARTPSCDKSGLRKGTWTAEEDRKLIAYVTRYGCWNWRQLPKFAGLARCGKSCRLRWLNY 256
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRP IKRG+FT +EE++II + LGNRW+ IAA LP RTDN++KN+W+T LK+++++ E
Sbjct: 257 LRPDIKRGHFTQEEEEIIIRMHKNLGNRWSIIAAELPGRTDNEVKNHWHTSLKRRVQQNE 436
Query: 121 TSTSSES 127
+ S S
Sbjct: 437 EAKVSNS 457
>AV421932
Length = 402
Score = 147 bits (370), Expect = 3e-36
Identities = 64/103 (62%), Positives = 78/103 (75%)
Frame = +3
Query: 6 CCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNYLRPGI 65
CC+++ VK+G W+PEED L+ YI HG G W VP GL RC KSCRLRW NYLRP I
Sbjct: 93 CCNQQKVKRGLWSPEEDEKLIRYITTHGYGCWSEVPEKAGLQRCGKSCRLRWINYLRPDI 272
Query: 66 KRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYW 108
+RG FT +EEK+II L ++GNRWA IA++LP RTDN+IKNYW
Sbjct: 273 RRGRFTPEEEKLIISLHGVVGNRWAHIASHLPGRTDNEIKNYW 401
>TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%)
Length = 532
Score = 144 bits (362), Expect = 3e-35
Identities = 69/143 (48%), Positives = 92/143 (64%), Gaps = 5/143 (3%)
Frame = +3
Query: 1 MVRPPCCDKEG--VKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWT 58
M + PC V+KGPWT EED+IL++YI HG G W + + GL R KSCRLRW
Sbjct: 66 MDKKPCNSSHDPEVRKGPWTIEEDLILINYIANHGEGVWNTLAKSAGLKRTGKSCRLRWL 245
Query: 59 NYLRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKK 118
NYLRP ++RGN T +E+ +I+ L A GNRW+ IA +LP RTDN+IKN+W T ++K +K+
Sbjct: 246 NYLRPDVRRGNITPEEQLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKHIKQ 425
Query: 119 ---LETSTSSESCLGHDEFSVSQ 138
L+ SS H + S SQ
Sbjct: 426 AESLQQQQSSSEINDHHQTSTSQ 494
>AV413379
Length = 416
Score = 142 bits (359), Expect = 6e-35
Identities = 63/105 (60%), Positives = 79/105 (75%)
Frame = +3
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K KG WT EED L++YI+ HG G W+++P + GLLRC KSCRLRW NY
Sbjct: 102 MGRSPCCEKAHTNKGAWTKEEDDRLIAYIRAHGEGCWRSLPKSAGLLRCGKSCRLRWINY 281
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIK 105
LRP +KRGNFT QE+++II L +LLGN+W+ IA L RTDN+IK
Sbjct: 282 LRPDLKRGNFTPQEDELIIKLHSLLGNKWSLIAGRLAGRTDNEIK 416
>CB828471
Length = 525
Score = 118 bits (295), Expect = 2e-27
Identities = 54/77 (70%), Positives = 63/77 (81%)
Frame = +1
Query: 41 PTNTGLLRCSKSCRLRWTNYLRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRT 100
P GLLRC KSCRLRW NYLRP IKRGNF+ +EE II+L LLGNRW+AIAA LP RT
Sbjct: 1 PKQAGLLRCGKSCRLRWANYLRPDIKRGNFSREEEDEIINLHELLGNRWSAIAARLPGRT 180
Query: 101 DNDIKNYWNTYLKKKLK 117
DN+IKN W+T+LKK+L+
Sbjct: 181 DNEIKNVWHTHLKKRLQ 231
>BP034046
Length = 490
Score = 113 bits (283), Expect = 4e-26
Identities = 69/118 (58%), Positives = 82/118 (69%), Gaps = 7/118 (5%)
Frame = -2
Query: 206 DNIARLLKGWMKNKPKEGSNGNNTNVTQNYETSASSEG---MEKGSTS-VELSETFESLF 261
DNIARLLKGWMK PK GS N +T ++SEG + KGS + VEL+ETFESLF
Sbjct: 489 DNIARLLKGWMKKSPK-GSCSKTLN-----DTDSTSEGYSIVAKGSNNCVELNETFESLF 328
Query: 262 GY-ESFDSSNSDS--TTLFQDESKPENIGEIMPFSMLEKWLLDDGACQEKVGLSEINY 316
GY ES DS NS+S TLFQDESKP +G MPFS+LE+WLLDD QEK+ L + N+
Sbjct: 327 GYNESLDSLNSESPEATLFQDESKPAGVGAEMPFSLLEQWLLDDAGYQEKLSLLQ*NH 154
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 108 bits (271), Expect = 9e-25
Identities = 57/133 (42%), Positives = 76/133 (56%), Gaps = 4/133 (3%)
Frame = +1
Query: 14 KGPWTPEEDIILVSYIQEHGPGNW----KAVPTNTGLLRCSKSCRLRWTNYLRPGIKRGN 69
KGPW+PEED L ++ HGP NW K++P +G KSCRLRW N L P ++
Sbjct: 184 KGPWSPEEDEALQKLVERHGPRNWSLISKSIPGRSG-----KSCRLRWCNQLSPQVEHRA 348
Query: 70 FTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLETSTSSESCL 129
FT +E+ II A GN+WA IA L RTDN IKN+WN+ LK+K + + +
Sbjct: 349 FTPEEDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRKCSSIFAADDLAAGG 528
Query: 130 GHDEFSVSQPIAR 142
G +F QP+ R
Sbjct: 529 GGGDF-CPQPLKR 564
>TC9457 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (43%)
Length = 860
Score = 107 bits (266), Expect = 3e-24
Identities = 52/106 (49%), Positives = 67/106 (63%), Gaps = 4/106 (3%)
Frame = +3
Query: 14 KGPWTPEEDIILVSYIQEHGPGNW----KAVPTNTGLLRCSKSCRLRWTNYLRPGIKRGN 69
KGPW+PEED L +++HGP NW K++P +G KSCRLRW N L P ++
Sbjct: 78 KGPWSPEEDEALQRLVEKHGPRNWSLISKSIPGRSG-----KSCRLRWCNQLSPQVEHRA 242
Query: 70 FTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKK 115
FT +E+ II A GN+WA IA L RTDN IKN+WN+ LK+K
Sbjct: 243 FTAEEDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK 380
>TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (50%)
Length = 487
Score = 104 bits (259), Expect = 2e-23
Identities = 50/103 (48%), Positives = 65/103 (62%)
Frame = +2
Query: 14 KGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNYLRPGIKRGNFTDQ 73
KGPW+ EED IL ++ HG NW + R KSCRLRW N L P ++ F+ Q
Sbjct: 170 KGPWSAEEDRILTRLVERHGARNWSLISRYIKG-RSGKSCRLRWCNQLSPTVEHRPFSVQ 346
Query: 74 EEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL 116
E++ II A GNRWAAIA LP RTDN +KN+WN+ LK+++
Sbjct: 347 EDETIIAAHARFGNRWAAIARLLPGRTDNAVKNHWNSTLKRRV 475
>TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (28%)
Length = 346
Score = 102 bits (253), Expect = 1e-22
Identities = 48/97 (49%), Positives = 60/97 (61%)
Frame = +1
Query: 14 KGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNYLRPGIKRGNFTDQ 73
KGPW+PEED L +Q HGP NW A+ + R KSCRLRW N L P ++ FT +
Sbjct: 55 KGPWSPEEDEALTRLVQVHGPKNWTAISKSIPG-RSGKSCRLRWCNQLSPEVEHRPFTPE 231
Query: 74 EEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNT 110
E+ II A GN+WA IA L RTDN +KN+WN+
Sbjct: 232 EDSTIIRAHARCGNKWATIARILNGRTDNAVKNHWNS 342
>BI420780
Length = 475
Score = 92.8 bits (229), Expect = 7e-20
Identities = 43/86 (50%), Positives = 60/86 (69%), Gaps = 2/86 (2%)
Frame = +1
Query: 17 WTPEEDIILVSYIQEHGPGNWKAVPT--NTGLLRCSKSCRLRWTNYLRPGIKRGNFTDQE 74
W+ EED +L +Y+Q++GP W V NT L R +KSC RW NYL+PGIK+G+ T +E
Sbjct: 211 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 390
Query: 75 EKMIIHLQALLGNRWAAIAAYLPQRT 100
++++I LQA GN+W IAA +P RT
Sbjct: 391 QRLVILLQANYGNKWKKIAAEVPGRT 468
>TC17769 similar to UP|Q40175 (Q40175) Transcription factor, partial (8%)
Length = 545
Score = 68.2 bits (165), Expect = 2e-12
Identities = 43/83 (51%), Positives = 52/83 (61%), Gaps = 5/83 (6%)
Frame = +2
Query: 225 NGNNTNVTQNYETSASSEGMEK-GSTSVELSETFESLFGYESFDSSNSD--STTLFQDES 281
N N N+ +ASS G + STS ELSETFESLFG+ES +S NSD S + E
Sbjct: 23 NSFNYNLNAAGTDTASSSGAKGISSTSAELSETFESLFGFESLESPNSDQFSPSNLSPER 202
Query: 282 KPE--NIGEIMPFSMLEKWLLDD 302
KP+ IG MP S+LEKWLLD+
Sbjct: 203 KPDVNVIGPEMPLSLLEKWLLDE 271
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.128 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,035,795
Number of Sequences: 28460
Number of extensions: 83090
Number of successful extensions: 423
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 412
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 414
length of query: 318
length of database: 4,897,600
effective HSP length: 90
effective length of query: 228
effective length of database: 2,336,200
effective search space: 532653600
effective search space used: 532653600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC139600.8


