

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
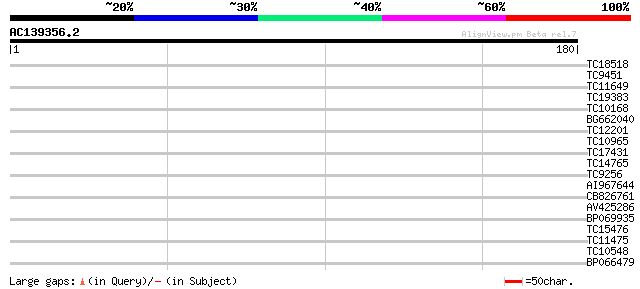
Score E
Sequences producing significant alignments: (bits) Value
TC18518 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 28 1.2
TC9451 homologue to UP|O24457 (O24457) Pyruvate dehydrogenase E1... 27 1.6
TC11649 similar to UP|Q9M4G3 (Q9M4G3) CMP-KDO synthetase precurs... 27 2.7
TC19383 27 2.7
TC10168 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase... 27 2.7
BG662040 26 4.6
TC12201 25 6.0
TC10965 25 6.0
TC17431 similar to UP|Q44395 (Q44395) Ti plasmid pTi15955 T-DNA ... 25 6.0
TC14765 homologue to UP|Q8T5T7 (Q8T5T7) Porin, partial (4%) 25 6.0
TC9256 homologue to UP|ERD2_PETHY (Q9ZTN2) ER lumen protein reta... 25 6.0
AI967644 25 6.0
CB826761 25 6.0
AV425286 25 6.0
BP069935 25 7.8
TC15476 similar to UP|THRC_SOLTU (Q9MT28) Threonine synthase, ch... 25 7.8
TC11475 25 7.8
TC10548 similar to UP|Q940V0 (Q940V0) T23O15.3/T23O15.3, partial... 25 7.8
BP066479 25 7.8
>TC18518 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(10%)
Length = 623
Score = 27.7 bits (60), Expect = 1.2
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = +3
Query: 133 LGCCKLEQLKYFCKYNNHH 151
LGCCKLE YFC+ + H
Sbjct: 423 LGCCKLESF-YFCRTESSH 476
>TC9451 homologue to UP|O24457 (O24457) Pyruvate dehydrogenase E1 alpha
subunit (At1g01090/T25K16_8) , partial (52%)
Length = 1082
Score = 27.3 bits (59), Expect = 1.6
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = -2
Query: 129 LGQMLGCCKLEQLKYFCKYNNHHRT 153
L Q+LG C EQ + FC YN T
Sbjct: 586 LTQLLGHCTSEQNQTFCPYNTSRHT 512
>TC11649 similar to UP|Q9M4G3 (Q9M4G3) CMP-KDO synthetase precursor ,
partial (33%)
Length = 582
Score = 26.6 bits (57), Expect = 2.7
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +3
Query: 138 LEQLKYFCKYNNHHRTGAKNRVLYLTYF 165
LEQ K+ C N++H+T +RV Y+ +F
Sbjct: 315 LEQ*KHLCIINHYHKTSQIDRV-YIPFF 395
>TC19383
Length = 558
Score = 26.6 bits (57), Expect = 2.7
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = -2
Query: 118 KKKNINWLFLLLGQMLGCCKLEQLKYFCK 146
K++ N L L + ++ GC KLE LK+ C+
Sbjct: 536 KQQQHNQLCLSMEKVKGC*KLENLKWACR 450
>TC10168 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, partial
(21%)
Length = 717
Score = 26.6 bits (57), Expect = 2.7
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = +2
Query: 63 QMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMN 96
Q+P L+TK Q LVT +H+R P+IL++
Sbjct: 290 QLPSKLKTK-----QALVTTSTRVHNRNPRILIS 376
>BG662040
Length = 393
Score = 25.8 bits (55), Expect = 4.6
Identities = 17/48 (35%), Positives = 23/48 (47%), Gaps = 1/48 (2%)
Frame = +2
Query: 118 KKKNINWLFLLLGQMLGCCKLEQLKYFCKYNNHHRTG-AKNRVLYLTY 164
K +IN FL C+L+ + YFC HR G ++VLY Y
Sbjct: 152 KAPHINLNFLRFDSFP*LCRLQAVLYFC-----HRAG*GLHQVLYHNY 280
>TC12201
Length = 503
Score = 25.4 bits (54), Expect = 6.0
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = -3
Query: 140 QLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLD 172
+LKYF Y N + K + + +LCQQL+
Sbjct: 366 KLKYFINYENMNTITVKVPITLSSCRKLCQQLN 268
>TC10965
Length = 507
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/18 (61%), Positives = 14/18 (77%)
Frame = +1
Query: 122 INWLFLLLGQMLGCCKLE 139
IN LFLLLGQ+L C ++
Sbjct: 55 INLLFLLLGQILRCIMVQ 108
>TC17431 similar to UP|Q44395 (Q44395) Ti plasmid pTi15955 T-DNA region,
partial (12%)
Length = 737
Score = 25.4 bits (54), Expect = 6.0
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = -1
Query: 140 QLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLD 172
+LKYF Y N + K + + +LCQQL+
Sbjct: 629 KLKYFINYENMNTITVKVPITLSSCRKLCQQLN 531
>TC14765 homologue to UP|Q8T5T7 (Q8T5T7) Porin, partial (4%)
Length = 1114
Score = 25.4 bits (54), Expect = 6.0
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -1
Query: 137 KLEQLKYFCKYNNHHRTGAKNRV 159
KL +KY+CKY H T +N +
Sbjct: 1114 KL*TIKYYCKYQKIHYTEFQNYI 1046
>TC9256 homologue to UP|ERD2_PETHY (Q9ZTN2) ER lumen protein retaining
receptor (HDEL receptor) (PGP169-12), complete
Length = 1009
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/16 (68%), Positives = 11/16 (68%)
Frame = +2
Query: 2 TANHTHVPPRRRSAPR 17
TA HTH P RRSA R
Sbjct: 41 TATHTHTPIGRRSAQR 88
>AI967644
Length = 244
Score = 25.4 bits (54), Expect = 6.0
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 2/39 (5%)
Frame = -2
Query: 27 WATNHRA--KIHNLNHLLQNKIFSITGDVQCKSCQTKFQ 63
W T+ +A KIHNL +L + T V + +TKFQ
Sbjct: 159 WTTHCKAYPKIHNLRGVLGVSLRLKTNTVAAATTKTKFQ 43
>CB826761
Length = 417
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = +3
Query: 5 HTHVPPRRRSAPRSGPIPIP 24
H PPRRRS S P+P P
Sbjct: 33 HLDQPPRRRSPCVSSPLPDP 92
>AV425286
Length = 320
Score = 25.4 bits (54), Expect = 6.0
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = -1
Query: 5 HTHVPPRRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQ 43
H H P + SA +PI ++ KIH L+H LQ
Sbjct: 278 HIHECPHKLSA-----VPIELLYLVG*HQKIHALDHHLQ 177
>BP069935
Length = 376
Score = 25.0 bits (53), Expect = 7.8
Identities = 12/25 (48%), Positives = 15/25 (60%), Gaps = 1/25 (4%)
Frame = -3
Query: 2 TANHT-HVPPRRRSAPRSGPIPIPF 25
T NH H PRRR++PR G + F
Sbjct: 296 TQNHDDHSGPRRRASPRIGAATVMF 222
>TC15476 similar to UP|THRC_SOLTU (Q9MT28) Threonine synthase, chloroplast
precursor (TS) , partial (19%)
Length = 658
Score = 25.0 bits (53), Expect = 7.8
Identities = 11/28 (39%), Positives = 17/28 (60%), Gaps = 1/28 (3%)
Frame = -3
Query: 83 MNTMHDRAP-KILMNPRLPRCAHCNQEN 109
M++ H +P L NPR+P+ C+Q N
Sbjct: 158 MSSAHHNSPISRLDNPRVPQLKQCSQSN 75
>TC11475
Length = 561
Score = 25.0 bits (53), Expect = 7.8
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = +2
Query: 147 YNNHHRTGAKNRVLYLTYF 165
Y HRTG VLY+ YF
Sbjct: 488 YVKFHRTGLTTLVLYIVYF 544
>TC10548 similar to UP|Q940V0 (Q940V0) T23O15.3/T23O15.3, partial (10%)
Length = 760
Score = 25.0 bits (53), Expect = 7.8
Identities = 12/30 (40%), Positives = 15/30 (50%)
Frame = +3
Query: 4 NHTHVPPRRRSAPRSGPIPIPFIWATNHRA 33
NH H+ R R RS PIP+ + N A
Sbjct: 642 NHIHLM*RVRWVERSNTFPIPYSYQRNRCA 731
>BP066479
Length = 445
Score = 25.0 bits (53), Expect = 7.8
Identities = 18/71 (25%), Positives = 34/71 (47%)
Frame = -3
Query: 36 HNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILM 95
HN HLL + ++ ++ ++C Q P LR +F V S+ L + + + P + +
Sbjct: 395 HNFIHLLSL*VLALGLNIWPRTCIMVLQFPCHLREQF-VYSK*LNSWSKLVGQQTPCVNL 219
Query: 96 NPRLPRCAHCN 106
N ++ CN
Sbjct: 218 NFQIRN*ELCN 186
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,385,567
Number of Sequences: 28460
Number of extensions: 71552
Number of successful extensions: 630
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 625
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 629
length of query: 180
length of database: 4,897,600
effective HSP length: 84
effective length of query: 96
effective length of database: 2,506,960
effective search space: 240668160
effective search space used: 240668160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC139356.2


