

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.8 + phase: 0 /pseudo
(481 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
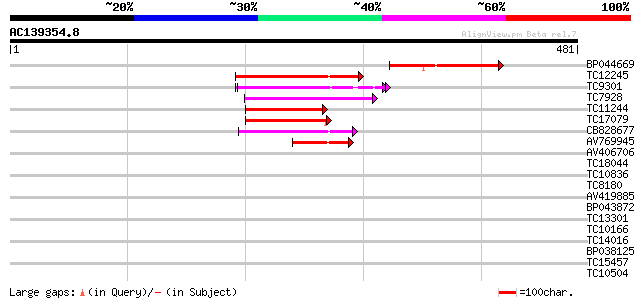
Score E
Sequences producing significant alignments: (bits) Value
BP044669 126 9e-30
TC12245 similar to UP|Q84K11 (Q84K11) Type 5 protein serine/thre... 79 2e-15
TC9301 similar to UP|Q7Y0Z0 (Q7Y0Z0) TPR1, complete 63 1e-10
TC7928 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like... 55 3e-08
TC11244 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (32%) 54 7e-08
TC17079 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (38%) 50 6e-07
CB828677 44 4e-05
AV769945 41 4e-04
AV406706 39 0.002
TC18044 similar to UP|Q9LDC0 (Q9LDC0) FKBP-like protein (Genomic... 39 0.002
TC10836 similar to GB|AAL60196.1|18139887|AF441079 O-linked N-ac... 38 0.004
TC8180 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cel... 35 0.021
AV419885 34 0.060
BP043872 31 0.51
TC13301 homologue to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyru... 29 1.5
TC10166 weakly similar to UP|Q8W0T8 (Q8W0T8) Gb protein, partial... 29 1.5
TC14016 similar to UP|Q38931 (Q38931) Rof1, partial (12%) 28 2.5
BP038125 28 2.5
TC15457 similar to UP|Q9LV01 (Q9LV01) Gb|AAC14404.1, partial (21%) 27 5.6
TC10504 similar to GB|AAN18208.1|23308477|BT000642 At2g29670/T27... 27 7.4
>BP044669
Length = 548
Score = 126 bits (316), Expect = 9e-30
Identities = 66/102 (64%), Positives = 76/102 (73%), Gaps = 5/102 (4%)
Frame = -2
Query: 323 GSRSHAAPASFGSMPFNPSNLASMFMA----AANGGQGSHSQEGQEDANSSGANEPEIRF 378
G RSH P F SMPFNP+NLASMFM A N QG H QE +EDAN+SGA+EPEIR
Sbjct: 547 GPRSHGVPPPFTSMPFNPNNLASMFMNMAPNATNAHQGPHPQE-REDANTSGADEPEIRA 371
Query: 379 GG-NVNVNQDQIPQELRGAFQSVMHMFSGNAPPGQPHDQMNG 419
G N ++N +QIPQ+L GAFQS+M MFSG+ PPGQPHDQ NG
Sbjct: 370 GSSNFSMNFEQIPQDLSGAFQSMMEMFSGSVPPGQPHDQTNG 245
>TC12245 similar to UP|Q84K11 (Q84K11) Type 5 protein serine/threonine
phosphatase 62 kDa isoform, partial (26%)
Length = 559
Score = 79.0 bits (193), Expect = 2e-15
Identities = 38/109 (34%), Positives = 68/109 (61%)
Frame = +2
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A ++++Y AIELY AI + ++AVY NRA A+ ++ Y AI D+ ++IE+DP Y+K
Sbjct: 173 AFKARKYAQAIELYTEAIELDSQNAVYLANRAFAHLRLEEYGSAILDATKAIEVDPKYAK 352
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R G A+ G +++A+ K F++ ++ PN+ + ++ E +M+
Sbjct: 353 GYYRRGAAHLGLGKFKEAL-KDFQQVKRMCPNDPDATKKLKECEKAVMK 496
>TC9301 similar to UP|Q7Y0Z0 (Q7Y0Z0) TPR1, complete
Length = 1288
Score = 62.8 bits (151), Expect = 1e-10
Identities = 36/130 (27%), Positives = 70/130 (53%)
Frame = +3
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y +A+ Y +I Y NRAA YT++ E ++D+ + IE+DP + K Y
Sbjct: 567 KQQKYPEAVGHYTESIRRNPNDPRAYSNRAACYTKLGAMPEGLKDAEKCIELDPTFVKGY 746
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRS 313
+R G Y ++ A++ +++ L+LDPNN+ + + I K M++ ++A +
Sbjct: 747 TRKGAVQYFMKDHEKALET-YREGLKLDPNNQELLDGI----GKCMQQINKASRG-DLTP 908
Query: 314 QEFQNHYARG 323
+E + A+G
Sbjct: 909 EELKERQAKG 938
Score = 43.5 bits (101), Expect = 8e-05
Identities = 30/129 (23%), Positives = 61/129 (47%)
Frame = +3
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + KQ+ AI+ Y+ A+ + ++ + NRAA Y ++ +Y + I+D +++E
Sbjct: 144 AYKKKQFDTAIQHYSKALELDDEDISFLTNRAAVYLEMGKYEDCIKDCDKAVERGRELRA 323
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS 311
Y + A +G A+ K K + DP E+ ++ + EH+ + + + Q +
Sbjct: 324 DYKMVARALTRKGT---ALVKMAKCSADFDPAIETFQK--ALTEHRNPDTLKKLNEAQKA 488
Query: 312 RSQEFQNHY 320
+ Q Y
Sbjct: 489 KKDLEQKEY 515
>TC7928 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like, partial
(52%)
Length = 765
Score = 55.1 bits (131), Expect = 3e-08
Identities = 33/114 (28%), Positives = 57/114 (49%), Gaps = 1/114 (0%)
Frame = +2
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
+AIE AI++ SA+ Y NRA+ Y ++ + AI+D+ ++EI+P+ +K Y G+A
Sbjct: 404 EAIESLTDAISLNPTSAIMYGNRASVYIKMKKPNAAIRDANAALEINPDSAKGYKTRGMA 583
Query: 260 YYAQGNYRDAI-DKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSR 312
G + +A D L D +V + + HK+ E R + + R
Sbjct: 584 RALLGQWEEAAKDLHVASKLDYDEEISAVLKKVEPNAHKIEEHRRKYEDLHKER 745
>TC11244 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (32%)
Length = 417
Score = 53.5 bits (127), Expect = 7e-08
Identities = 29/69 (42%), Positives = 42/69 (60%)
Frame = +2
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A+EL AI + K A Y +RA A ++N +TEAI D+ ++IE++P+ SKAY R G A
Sbjct: 92 AVELLTQAIDVDPKHAELYADRAQANIKLNNFTEAIADANKAIELNPSLSKAYLRKGTAC 271
Query: 261 YAQGNYRDA 269
Y+ A
Sbjct: 272 MKLEEYQTA 298
>TC17079 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (38%)
Length = 676
Score = 50.4 bits (119), Expect = 6e-07
Identities = 27/76 (35%), Positives = 47/76 (61%), Gaps = 3/76 (3%)
Frame = +1
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++LY AI + +A + +RA A+ ++N +TEA+ D+ ++I++DP+ KAY R G A
Sbjct: 154 AVDLYTEAINLDPINANLFADRAQAHLKLNAFTEAVSDANKAIQLDPSLPKAYLRKGTAC 333
Query: 261 YAQGNY---RDAIDKG 273
Y + A++KG
Sbjct: 334 IKLEEYHTAKVALEKG 381
>CB828677
Length = 520
Score = 44.3 bits (103), Expect = 4e-05
Identities = 24/101 (23%), Positives = 50/101 (48%)
Frame = +3
Query: 195 SKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYS 254
S ++ DA Y + + V YCNRA ++++ + +++QD +++ I PNY+KA
Sbjct: 204 SGKFSDACTAYGEGLKYDNSNYVLYCNRAICWSKLGLWEQSVQDCNQALNIQPNYTKALL 383
Query: 255 RLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAE 295
R + + + + K ++ PN+ V ++ A+
Sbjct: 384 RRAASNAKLERWAEVV-KDYQALKHELPNDNDVAVSLHQAQ 503
>AV769945
Length = 284
Score = 41.2 bits (95), Expect = 4e-04
Identities = 20/51 (39%), Positives = 33/51 (64%)
Frame = -1
Query: 241 RSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENI 291
+S+E+ P Y+ A++ LGL + A+G +A F+KALQ DP ++ K N+
Sbjct: 266 KSLELQPGYAPAFNNLGLVFVAEGLLEEA-KYCFEKALQSDPLLDAAKSNL 117
>AV406706
Length = 433
Score = 38.9 bits (89), Expect = 0.002
Identities = 24/77 (31%), Positives = 40/77 (51%), Gaps = 1/77 (1%)
Frame = +2
Query: 195 SKQYFDAIELYNCAIAIYEKSAVY-YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
SK + AI + + ++ + A+ CNRA Y+Q+ + I+D R++++DP AY
Sbjct: 203 SKDWSKAIRILDSLVS--QSGAIQDICNRAFCYSQLELHKHVIKDCDRALQLDPVLLHAY 376
Query: 254 SRLGLAYYAQGNYRDAI 270
G A A G DA+
Sbjct: 377 ILKGHALSALGRKSDAV 427
>TC18044 similar to UP|Q9LDC0 (Q9LDC0) FKBP-like protein (Genomic DNA,
chromosome 3, P1 clone: MIL23), partial (41%)
Length = 867
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/76 (34%), Positives = 39/76 (51%)
Frame = +1
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKAL 278
+ N AA + ++NRY EAI + D N KA R G A A G DA + + KA
Sbjct: 187 HLNMAACFIKLNRYEEAIGHCSIVLGEDENNVKALFRRGKARAALGQ-TDAAREDYLKAR 363
Query: 279 QLDPNNESVKENIRVA 294
+ P ++++ I+VA
Sbjct: 364 KYAPEDKAIARRIKVA 411
>TC10836 similar to GB|AAL60196.1|18139887|AF441079 O-linked N-acetyl
glucosamine transferase {Arabidopsis thaliana;} ,
partial (16%)
Length = 833
Score = 37.7 bits (86), Expect = 0.004
Identities = 28/79 (35%), Positives = 40/79 (50%), Gaps = 1/79 (1%)
Frame = +2
Query: 192 AMQSKQYFD-AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYS 250
A++ K D AI Y AI A + N A+AY + R TEA Q +++ I+P
Sbjct: 596 ALKEKGNIDLAIRYYLIAIEFRPNFADAWSNLASAYMRKGRLTEAAQCCRQALAINPLMV 775
Query: 251 KAYSRLGLAYYAQGNYRDA 269
A+S LG AQG ++A
Sbjct: 776 DAHSNLGNLMKAQGLVQEA 832
>TC8180 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein), partial (3%)
Length = 575
Score = 35.4 bits (80), Expect = 0.021
Identities = 20/68 (29%), Positives = 37/68 (54%)
Frame = +2
Query: 225 AYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNN 284
A +Q + A+ D S+++ P KA+S LG Y + +A K +++ALQL+P
Sbjct: 5 ATSQRGKVDPALADLTESVKLSPRNVKAWSLLGECYEGKKMEEEA-KKAYQEALQLEPEL 181
Query: 285 ESVKENIR 292
+ +E ++
Sbjct: 182 SAAQEALK 205
>AV419885
Length = 108
Score = 33.9 bits (76), Expect = 0.060
Identities = 14/35 (40%), Positives = 25/35 (71%)
Frame = +1
Query: 221 NRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSR 255
NRAA + ++ ++ ++I+DS +++ I PNY KA R
Sbjct: 4 NRAACWFKLGQWEKSIEDSNQALNIQPNYMKATLR 108
>BP043872
Length = 491
Score = 30.8 bits (68), Expect = 0.51
Identities = 20/73 (27%), Positives = 35/73 (47%)
Frame = -1
Query: 228 QINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESV 287
++N + EA + R + ++ KA R A + D + KKAL++DP+N V
Sbjct: 488 KLNDFIEAEKLCTRVLNLESTNVKALYRRAEALMQLADL-DLAELDIKKALEVDPDNREV 312
Query: 288 KENIRVAEHKLME 300
K + + K+ E
Sbjct: 311 KLQYKTLKEKVKE 273
>TC13301 homologue to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvate
kinase-like protein), partial (13%)
Length = 618
Score = 29.3 bits (64), Expect = 1.5
Identities = 10/33 (30%), Positives = 21/33 (63%)
Frame = +3
Query: 209 IAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLR 241
+ +++K+A+Y CN A + R +++ D+LR
Sbjct: 111 VFLFQKAAIYKCNMAGKPVVVTRVVDSMTDNLR 209
>TC10166 weakly similar to UP|Q8W0T8 (Q8W0T8) Gb protein, partial (5%)
Length = 980
Score = 29.3 bits (64), Expect = 1.5
Identities = 37/189 (19%), Positives = 74/189 (38%)
Frame = +3
Query: 264 GNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQNHYARG 323
GN+ D K +DPN + K ++ H++ ++ + N NSR + + G
Sbjct: 345 GNHSDEDIYAMLKECSMDPNETTQKLLLQDTFHEVKRKKDKRKENLNSR-ESVEPRGRPG 521
Query: 324 SRSHAAPASFGSMPFNPSNLASMFMAAANGGQGSHSQEGQEDANSSGANEPEIRFGGNVN 383
++ G+ F+P N+A + G S G++ +GA++ + +
Sbjct: 522 NQGRGPRGGRGN--FSPHNVAH------DAGGSKSSVTGKD----NGAHQVSEKVVPPLP 665
Query: 384 VNQDQIPQELRGAFQSVMHMFSGNAPPGQPHDQMNGSHEDANSSEDNEPDIQFGGNVNLN 443
+Q+ I + S N P G+ A+S+ + G+ N +
Sbjct: 666 ASQEIISKGKSSGTSSA--SIIANGPTIVASGTAGGASRSASSAGSGDRTGPSSGDNNNS 839
Query: 444 SDQIPQELS 452
S+ +P + S
Sbjct: 840 SNALPSDSS 866
>TC14016 similar to UP|Q38931 (Q38931) Rof1, partial (12%)
Length = 506
Score = 28.5 bits (62), Expect = 2.5
Identities = 23/91 (25%), Positives = 45/91 (49%)
Frame = +2
Query: 251 KAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQN 310
KA R AY + D + +KAL++DPN+ V+ E+K ++E+ + H +
Sbjct: 17 KALYRRAQAYIPLADL-DLAEFDIQKALEIDPNHRDVQ-----LEYKTLKEKMKEFHQKE 178
Query: 311 SRSQEFQNHYARGSRSHAAPASFGSMPFNPS 341
++ + N +++ S+ + S S P P+
Sbjct: 179 AKF--YGNMFSQMSQLDSLDQS*TSFPGCPT 265
>BP038125
Length = 435
Score = 28.5 bits (62), Expect = 2.5
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = +1
Query: 268 DAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
D + KKAL+++P+N VK ++ + K+ E
Sbjct: 43 DLAEMDIKKALEIEPDNRDVKIEYKILKEKVRE 141
>TC15457 similar to UP|Q9LV01 (Q9LV01) Gb|AAC14404.1, partial (21%)
Length = 1147
Score = 27.3 bits (59), Expect = 5.6
Identities = 20/71 (28%), Positives = 33/71 (46%)
Frame = +3
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKAL 278
Y RAA ++ EAI + R+I+ P+ + R Y + +Y + + + AL
Sbjct: 360 YRYRAAVLMDDHKEAEAIAELSRAIDFKPDLQLLHLRAAF-YDSMSDYVSTV-RDCEAAL 533
Query: 279 QLDPNNESVKE 289
LDPN+ E
Sbjct: 534 CLDPNHTETLE 566
>TC10504 similar to GB|AAN18208.1|23308477|BT000642 At2g29670/T27A16.23
{Arabidopsis thaliana;}, partial (14%)
Length = 544
Score = 26.9 bits (58), Expect = 7.4
Identities = 12/44 (27%), Positives = 25/44 (56%)
Frame = +2
Query: 241 RSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNN 284
R+IE++P ++AYS+ + N ++ + +A+ +PNN
Sbjct: 53 RAIEVEPPDAEAYSKYATFLWKVKNDLWGAEETYLEAISAEPNN 184
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,320,193
Number of Sequences: 28460
Number of extensions: 90078
Number of successful extensions: 468
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 453
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 462
length of query: 481
length of database: 4,897,600
effective HSP length: 94
effective length of query: 387
effective length of database: 2,222,360
effective search space: 860053320
effective search space used: 860053320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC139354.8


