

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
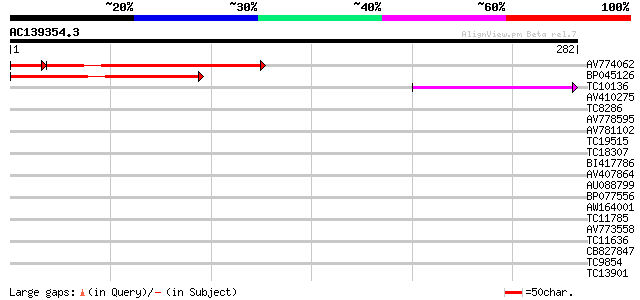
Query= AC139354.3 + phase: 0
(282 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV774062 152 4e-40
BP045126 124 2e-29
TC10136 42 1e-04
AV410275 32 0.093
TC8286 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabid... 27 0.13
AV778595 31 0.21
AV781102 31 0.21
TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, pa... 30 0.35
TC18307 similar to UP|Q7XYK9 (Q7XYK9) Plastid protein SufE (Frag... 30 0.35
BI417786 30 0.35
AV407864 30 0.46
AU088799 30 0.46
BP077556 30 0.60
AW164001 30 0.60
TC11785 homologue to UP|Q9ZVC2 (Q9ZVC2) At2g41370 protein, parti... 30 0.60
AV773558 30 0.60
TC11636 30 0.60
CB827847 29 0.79
TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1 (AT3g1... 29 0.79
TC13901 similar to UP|Q9FYU7 (Q9FYU7) UDP-glucose:sinapate gluco... 28 1.3
>AV774062
Length = 478
Score = 152 bits (383), Expect(2) = 4e-40
Identities = 76/109 (69%), Positives = 83/109 (75%)
Frame = +2
Query: 19 VAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLL 78
VAW+AVE HH H+ P D NCPSDQD E L+ ENRRLRNLLDQNLKLL
Sbjct: 176 VAWSAVECGHHLRHHNSPA--------GADQNCPSDQDSELLRSENRRLRNLLDQNLKLL 331
Query: 79 NNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETASGGNQFP 127
+NLSES SFL+NCPPDLH RL ATV+SDEYLTRLK LQ+ETA GGNQFP
Sbjct: 332 HNLSESPSFLNNCPPDLHTRLVATVKSDEYLTRLKFLQQETAGGGNQFP 478
Score = 28.9 bits (63), Expect(2) = 4e-40
Identities = 13/18 (72%), Positives = 15/18 (83%)
Frame = +1
Query: 1 MRTHNAVEMAKTVLEVAD 18
M H+A+EM KTVLEVAD
Sbjct: 121 MTGHHALEMTKTVLEVAD 174
>BP045126
Length = 542
Score = 124 bits (310), Expect = 2e-29
Identities = 62/96 (64%), Positives = 71/96 (73%)
Frame = +1
Query: 1 MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 60
M H+A+EM KTVLEVADVAW+AVE HH H+ P D NCPSDQD E L
Sbjct: 106 MTGHHALEMTKTVLEVADVAWSAVECGHHLRHHNSPATA--------DQNCPSDQDSELL 261
Query: 61 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLH 96
+ ENRRLRNLLDQNLKLL+NLSES SFL+NCPPD++
Sbjct: 262 RSENRRLRNLLDQNLKLLHNLSESPSFLNNCPPDVY 369
>TC10136
Length = 547
Score = 42.0 bits (97), Expect = 1e-04
Identities = 22/82 (26%), Positives = 40/82 (47%)
Frame = +2
Query: 201 KARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLALAGLYQTRSLLKVA 260
++R ++P LQ LS+ F+ K K+ W ++ Y +++WG G+Y +L+ A
Sbjct: 50 QSRTLTPIPLQEALSQTFSLKKKKGKLRKAWDGSQVIYNVASWGATAIGIYPNPVILRAA 229
Query: 261 AKGVHSGGKLALKMKSLKMAAP 282
K + + K+ L A P
Sbjct: 230 TKAFWTSCHVISKLL*LVDAVP 295
>AV410275
Length = 431
Score = 32.3 bits (72), Expect = 0.093
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = +2
Query: 26 YHHHHHRHHDPVVTHD 41
+HHHHH HH P + H+
Sbjct: 53 HHHHHHHHHGPQIHHN 100
>TC8286 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;}, partial (6%)
Length = 671
Score = 26.9 bits (58), Expect(2) = 0.13
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = +2
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNN 50
YHHHHH+H H + + N
Sbjct: 317 YHHHHHQHQQQHHFHQQQDQNVSGN 391
Score = 23.5 bits (49), Expect(2) = 0.13
Identities = 8/27 (29%), Positives = 15/27 (54%)
Frame = +2
Query: 8 EMAKTVLEVADVAWTAVEYHHHHHRHH 34
E+ + LE+ + + + +HHHH H
Sbjct: 260 EVYRPELEIGESSSSMPAGYHHHHHQH 340
>AV778595
Length = 594
Score = 31.2 bits (69), Expect = 0.21
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +3
Query: 25 EYHHHHHRHHDPVVTHDDKY 44
++HHHHH HH V ++ Y
Sbjct: 90 KHHHHHHHHHQVVAVNNKSY 149
Score = 30.8 bits (68), Expect = 0.27
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +3
Query: 27 HHHHHRHHDPVVTHDDKYDDKDNNCPS 53
HHHHH HH VV ++K N P+
Sbjct: 93 HHHHHHHHHQVVAVNNKSYCYGNRTPN 173
>AV781102
Length = 567
Score = 31.2 bits (69), Expect = 0.21
Identities = 8/16 (50%), Positives = 11/16 (68%)
Frame = +1
Query: 19 VAWTAVEYHHHHHRHH 34
+ W + +HHHHH HH
Sbjct: 292 LGWLRLHHHHHHHHHH 339
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = +1
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 316 HHHHHHHHH 342
>TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, partial (3%)
Length = 504
Score = 30.4 bits (67), Expect = 0.35
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = -1
Query: 24 VEYHHHHHRHHDP 36
+ +HHHHHRHH P
Sbjct: 135 LHHHHHHHRHHLP 97
>TC18307 similar to UP|Q7XYK9 (Q7XYK9) Plastid protein SufE (Fragment),
partial (9%)
Length = 773
Score = 30.4 bits (67), Expect = 0.35
Identities = 8/13 (61%), Positives = 11/13 (84%)
Frame = +2
Query: 25 EYHHHHHRHHDPV 37
++HHHHH HH P+
Sbjct: 188 QHHHHHHHHHHPL 226
>BI417786
Length = 513
Score = 30.4 bits (67), Expect = 0.35
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +2
Query: 26 YHHHHHRHHDPVVTH 40
+HHHHH HH V+ H
Sbjct: 122 HHHHHHHHHHHVLLH 166
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = +2
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 119 HHHHHHHHH 145
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = +2
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 116 HHHHHHHHH 142
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = +2
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 113 HHHHHHHHH 139
>AV407864
Length = 414
Score = 30.0 bits (66), Expect = 0.46
Identities = 9/15 (60%), Positives = 10/15 (66%)
Frame = +1
Query: 26 YHHHHHRHHDPVVTH 40
+HHHHH HH P H
Sbjct: 1 HHHHHHHHHPPNNAH 45
>AU088799
Length = 436
Score = 30.0 bits (66), Expect = 0.46
Identities = 10/13 (76%), Positives = 10/13 (76%)
Frame = +1
Query: 27 HHHHHRHHDPVVT 39
HHHHHRH P VT
Sbjct: 277 HHHHHRHQHPSVT 315
Score = 29.6 bits (65), Expect = 0.60
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = +3
Query: 24 VEYHHHHHRHHDPVVTHDDKYDDKD 48
+++HHHHH HH+ + + KD
Sbjct: 114 IKHHHHHHHHHES*LPYMQPEPSKD 188
>BP077556
Length = 298
Score = 29.6 bits (65), Expect = 0.60
Identities = 8/11 (72%), Positives = 10/11 (90%)
Frame = +1
Query: 25 EYHHHHHRHHD 35
++HHHHH HHD
Sbjct: 160 KWHHHHHHHHD 192
>AW164001
Length = 447
Score = 29.6 bits (65), Expect = 0.60
Identities = 8/14 (57%), Positives = 9/14 (64%)
Frame = -3
Query: 21 WTAVEYHHHHHRHH 34
W +HHHHH HH
Sbjct: 142 WQVRHHHHHHHHHH 101
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = -3
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 121 HHHHHHHHH 95
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = -3
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 118 HHHHHHHHH 92
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/11 (63%), Positives = 9/11 (81%)
Frame = -3
Query: 26 YHHHHHRHHDP 36
+HHHHH H +P
Sbjct: 115 HHHHHHHHQNP 83
Score = 26.9 bits (58), Expect = 3.9
Identities = 7/9 (77%), Positives = 8/9 (88%)
Frame = -3
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 124 HHHHHHHHH 98
>TC11785 homologue to UP|Q9ZVC2 (Q9ZVC2) At2g41370 protein, partial (42%)
Length = 644
Score = 29.6 bits (65), Expect = 0.60
Identities = 14/39 (35%), Positives = 19/39 (47%), Gaps = 2/39 (5%)
Frame = +1
Query: 26 YHHHHHRHHDPVVTHD--DKYDDKDNNCPSDQDLEALKL 62
+HHHHH HHD D D+ + D+E +KL
Sbjct: 466 HHHHHHGHHDMGAAADLEDQKIRRMRRALDSSDVELVKL 582
Score = 29.6 bits (65), Expect = 0.60
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = +1
Query: 24 VEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQN 74
+ +HHHHH HH H D + DLE K+ RR+R LD +
Sbjct: 451 IPHHHHHHHHHG----HHD--------MGAAADLEDQKI--RRMRRALDSS 561
>AV773558
Length = 469
Score = 29.6 bits (65), Expect = 0.60
Identities = 9/14 (64%), Positives = 10/14 (71%)
Frame = +3
Query: 26 YHHHHHRHHDPVVT 39
+HHHHH HH P T
Sbjct: 333 HHHHHHHHHHPHYT 374
Score = 28.5 bits (62), Expect = 1.3
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = +3
Query: 26 YHHHHHRHHDPVVTH 40
+HHHHH H+ P H
Sbjct: 345 HHHHHHPHYTPTQNH 389
Score = 28.1 bits (61), Expect = 1.8
Identities = 11/27 (40%), Positives = 13/27 (47%)
Frame = +3
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCP 52
+HHHHH HH H Y N+ P
Sbjct: 324 HHHHHHHHHH---HHHPHYTPTQNHHP 395
>TC11636
Length = 655
Score = 29.6 bits (65), Expect = 0.60
Identities = 8/18 (44%), Positives = 14/18 (77%)
Frame = -2
Query: 27 HHHHHRHHDPVVTHDDKY 44
HHHHH HH+ ++T ++ +
Sbjct: 474 HHHHHHHHNQLLT*EEMF 421
>CB827847
Length = 503
Score = 29.3 bits (64), Expect = 0.79
Identities = 11/33 (33%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -2
Query: 19 VAWTAVEYHHHHHRHHD-PVVTHDDKYDDKDNN 50
+A +++HH HH H + D++YDD D +
Sbjct: 133 IAVNRLKFHHQHHHHQECEKRLSDEEYDDDDGD 35
>TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1
(AT3g17100/K14A17_22_), partial (43%)
Length = 1108
Score = 29.3 bits (64), Expect = 0.79
Identities = 8/11 (72%), Positives = 9/11 (81%)
Frame = +2
Query: 26 YHHHHHRHHDP 36
+HHHHH HH P
Sbjct: 17 HHHHHHHHHHP 49
Score = 27.7 bits (60), Expect = 2.3
Identities = 11/32 (34%), Positives = 15/32 (46%), Gaps = 7/32 (21%)
Frame = +2
Query: 26 YHHHHHRHHD-------PVVTHDDKYDDKDNN 50
+HHHHH HH PV T + D++
Sbjct: 11 HHHHHHHHHHHHPSPLIPVTTFISVFSSSDSH 106
>TC13901 similar to UP|Q9FYU7 (Q9FYU7) UDP-glucose:sinapate
glucosyltransferase , partial (16%)
Length = 558
Score = 28.5 bits (62), Expect = 1.3
Identities = 9/20 (45%), Positives = 11/20 (55%)
Frame = +1
Query: 25 EYHHHHHRHHDPVVTHDDKY 44
E+HHHHH H V+ Y
Sbjct: 214 EHHHHHHPHQSATVSSSSTY 273
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.131 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,193,586
Number of Sequences: 28460
Number of extensions: 75345
Number of successful extensions: 2402
Number of sequences better than 10.0: 147
Number of HSP's better than 10.0 without gapping: 1285
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1910
length of query: 282
length of database: 4,897,600
effective HSP length: 89
effective length of query: 193
effective length of database: 2,364,660
effective search space: 456379380
effective search space used: 456379380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC139354.3


