

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138453.15 + phase: 0 /pseudo/partial
(195 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
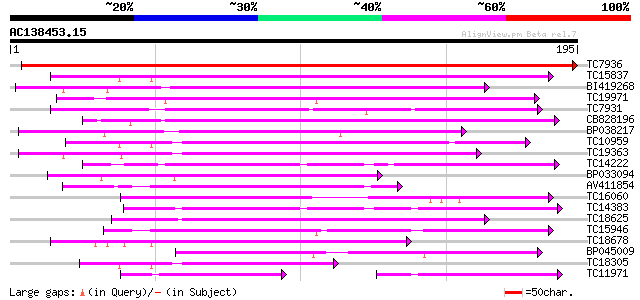
Score E
Sequences producing significant alignments: (bits) Value
TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, pa... 323 1e-89
TC15837 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyge... 96 3e-21
BI419268 92 5e-20
TC19971 similar to UP|Q84V01 (Q84V01) Anthocyanidin synthase, pa... 91 1e-19
TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial... 83 4e-17
CB828196 82 6e-17
BP038217 79 5e-16
TC10959 weakly similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/... 73 4e-14
TC19363 similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30... 69 4e-13
TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carb... 66 5e-12
BP033094 65 6e-12
AV411854 65 1e-11
TC16060 weakly similar to UP|Q9SHK4 (Q9SHK4) F12K11.6, partial (8%) 65 1e-11
TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-ca... 63 3e-11
TC18625 weakly similar to UP|O48631 (O48631) Ethylene-forming-en... 62 9e-11
TC15946 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 58 9e-10
TC18678 weakly similar to UP|Q9LY48 (Q9LY48) Leucoanthocyanidin ... 57 2e-09
BP045009 57 2e-09
TC18305 weakly similar to UP|Q9SB33 (Q9SB33) SRG1-like protein, ... 57 3e-09
TC11971 weakly similar to UP|Q43419 (Q43419) ACC oxidase, partia... 40 4e-09
>TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(94%)
Length = 1492
Score = 323 bits (828), Expect = 1e-89
Identities = 154/191 (80%), Positives = 170/191 (88%)
Frame = +3
Query: 5 KTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEA 64
KTL LAQ+ TLESSF+RD ERPKVAYNNFSNEIP+ISLAGID+VD R E CNKIVEA
Sbjct: 159 KTLTTLAQQNTLESSFVRDEDERPKVAYNNFSNEIPVISLAGIDEVDDRRSEICNKIVEA 338
Query: 65 CENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEP 124
CENWGIFQ+VDHGVD +L+S MT LAK FF LPPEEKLRFD++GGKKGGFIVSSHLQGE
Sbjct: 339 CENWGIFQVVDHGVDTELVSHMTTLAKEFFALPPEEKLRFDMTGGKKGGFIVSSHLQGES 518
Query: 125 VRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
V+DWRE++ YFSYPI+ RDYSRWP+ P GWK VTE+YSEKLM L+CKLLEVLSEAMGLEK
Sbjct: 519 VQDWREIVTYFSYPIRNRDYSRWPDTPAGWKAVTEEYSEKLMGLACKLLEVLSEAMGLEK 698
Query: 185 EALTKACVDMD 195
EALTKACVDMD
Sbjct: 699 EALTKACVDMD 731
>TC15837 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (50%)
Length = 679
Score = 96.3 bits (238), Expect = 3e-21
Identities = 56/181 (30%), Positives = 94/181 (50%), Gaps = 8/181 (4%)
Frame = +2
Query: 15 TLESSFIRDVSERPKVAYNNFS---NEIPIISLAGI-----DDVDGLRIETCNKIVEACE 66
++ +I+ +++RP + ++ S N IPII L G+ D R T +I EAC+
Sbjct: 89 SIPERYIKPLTDRPSINSSSSSSDANNIPIIDLRGLLYHGDPDHPEARASTLRQISEACK 268
Query: 67 NWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVR 126
WG FQIV+HGV L+ + FF LP E K ++ S G+ ++ +
Sbjct: 269 EWGFFQIVNHGVSPDLMDMARETWRKFFHLPMELKQQYANSPKTYEGYGSRLGVEKGAIL 448
Query: 127 DWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEA 186
DW + P+ +DY++WP P +EV ++Y +L+ L +L++VLS + LE++
Sbjct: 449 DWSDYYFLHYLPLSLKDYNKWPALPPSCREVFDEYGRELVKLCGRLMKVLSLNLELEEDF 628
Query: 187 L 187
L
Sbjct: 629 L 631
>BI419268
Length = 536
Score = 92.4 bits (228), Expect = 5e-20
Identities = 53/166 (31%), Positives = 87/166 (51%), Gaps = 3/166 (1%)
Frame = +1
Query: 3 PVKTLKYLAQEKTLE--SSFIRDVSERPKVAY-NNFSNEIPIISLAGIDDVDGLRIETCN 59
PV ++ + ++ L+ + R E K Y ++ S+EIPII + + D E
Sbjct: 43 PVPNVQEMVRKNPLQVPKRYERSEEEMEKENYKSHLSSEIPIIDFSLLSDGSK---EELL 213
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
K+ A + WG FQ+V+HG+ +L+ M L FF LP EEK ++ + G+ +S
Sbjct: 214 KLDTALKEWGFFQVVNHGIKTELMQRMKELTAEFFGLPVEEKNKYAMPPDDIQGYGHTSV 393
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKL 165
+ E + DW + +I+ YP K R WP PEG+K++ E YS ++
Sbjct: 394 VSDEQILDWCDQLIFLVYPTKFRKLQFWPETPEGFKDMIEAYSSEI 531
>TC19971 similar to UP|Q84V01 (Q84V01) Anthocyanidin synthase, partial (53%)
Length = 595
Score = 91.3 bits (225), Expect = 1e-19
Identities = 54/169 (31%), Positives = 85/169 (49%), Gaps = 3/169 (1%)
Frame = +1
Query: 17 ESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDG-LRIETCNKIVEACENWGIFQIVD 75
E + I DV E K ++P I L ID D +R + K+ +A E WG+ +V+
Sbjct: 100 ELANIGDVFEEEK----KVGPQVPTIDLKDIDSPDEFVRAKCREKLRKAAEEWGVMHLVN 267
Query: 76 HGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEPVRDWREMMI 133
HG+ +L++++ FF LP EEK ++ D + G G+ +W +
Sbjct: 268 HGIPDELLNQLKSAGAEFFSLPVEEKEKYANDQAAGNVQGYGSKLANNASGQLEWEDYFF 447
Query: 134 YFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGL 182
+ +P +RD S WP P + EVT Y+ +L L+ K+LEVLS +GL
Sbjct: 448 HLIFPEDKRDLSIWPKTPSYYTEVTSDYARRLRVLASKILEVLSLELGL 594
>TC7931 weakly similar to UP|Q43640 (Q43640) Dioxygenase, partial (29%)
Length = 1477
Score = 82.8 bits (203), Expect = 4e-17
Identities = 49/172 (28%), Positives = 90/172 (51%), Gaps = 3/172 (1%)
Frame = +1
Query: 15 TLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIV 74
T+ S+++ RP +P++ L G D R ET +I +A E +G FQ++
Sbjct: 61 TVPLSYVQPPESRPGTVTVASGKAVPVVDLGGHD-----RAETLKQIFKASEEYGFFQVI 225
Query: 75 DHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQ---GEPVRDWREM 131
+HGV +LI + + K F +P EEKL + S G ++ + + + ++ W++
Sbjct: 226 NHGVSNELIEDTLNIFKEFHAMPAEEKLS-ESSRDPNGSCMLYTSREINNKDCIQFWKDT 402
Query: 132 MIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLE 183
+ + P + WP KP ++EV E+Y+++L L ++LE+L E +GL+
Sbjct: 403 LRHPCTP-SEEFMEFWPLKPARYREVVEKYTKELRELGSQILEMLCEGLGLD 555
>CB828196
Length = 555
Score = 82.0 bits (201), Expect = 6e-17
Identities = 43/166 (25%), Positives = 92/166 (54%), Gaps = 2/166 (1%)
Frame = +3
Query: 26 ERPKVAYNNFSNEIP--IISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLI 83
ERP + YN+ + +P II L+ + D ++ ++ +AC+ WG FQ+++HGV+ L+
Sbjct: 3 ERPAL-YNSSTTPLPLPIIDLSKLLSKDH-KVPELERLHQACKEWGFFQLINHGVNTSLL 176
Query: 84 SEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRD 143
++ R + FF LP EEK +F+ G G+ + + +W ++ F +P+++R
Sbjct: 177 EDVKRGVQEFFHLPMEEKKKFEQRQGDAEGYGQVFVVSEDQKLEWGDVFFMFLFPLEKRK 356
Query: 144 YSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
+P P +++ + Y ++ L +LE+++ A+ ++ + +T+
Sbjct: 357 PYLFPKLPSKFRDDLDTYYTEVKKLGIHILELMANALKIDTKEITE 494
>BP038217
Length = 514
Score = 79.0 bits (193), Expect = 5e-16
Identities = 43/157 (27%), Positives = 83/157 (52%), Gaps = 3/157 (1%)
Frame = +1
Query: 4 VKTLKYLAQEKTLESSFIRDVSERPKVA-YNNFSNEIPIISLAGIDDVDGLRIETCNKIV 62
V+++ +++ ++ + F+R +E+P + E+PII L G D+V L ++IV
Sbjct: 55 VQSVAAQSKDASIPAMFVRSETEQPGITTVRGVELEVPIIDLNGTDEVKVL-----SEIV 219
Query: 63 EACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKG--GFIVSSHL 120
EA + WG+FQ+V+H + ++I+++ + K FF+LP EEK + G G+
Sbjct: 220 EASKEWGMFQVVNHEIPSEVIAKLQAVGKEFFELPQEEKEVYGKIEGSDSLEGYGTKLQK 399
Query: 121 QGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEV 157
+ + W + + + +P +Y WP P ++EV
Sbjct: 400 EVNGKKGWVDHLFHIIWPTSSINYRFWPKNPASYREV 510
>TC10959 weakly similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (21%)
Length = 634
Score = 72.8 bits (177), Expect = 4e-14
Identities = 44/163 (26%), Positives = 86/163 (51%), Gaps = 3/163 (1%)
Frame = +3
Query: 20 FIRDVSERPKVAYNNFS-NEIPIISLAGI--DDVDGLRIETCNKIVEACENWGIFQIVDH 76
++R ERP ++ + ++PII L+ + ++ G +E ++ AC+ WG FQ+++H
Sbjct: 159 YVRPQYERPILSTTVPNLRQVPIIDLSKLLSKNLKGPELE---RLHHACKEWGFFQLINH 329
Query: 77 GVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFS 136
GV L+ + R + FF+LP EEK++F G+ G+ + +W +M+ +
Sbjct: 330 GVSTSLLENVKRGVREFFNLPMEEKMKFQQRQGETEGYGQLLVFSEDQKLEWGDMLYMLT 509
Query: 137 YPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
P + R PN P ++ E Y ++ +S ++LE+++ A
Sbjct: 510 LPPEMRKPHLLPNLP--FRNDLEAYYAEMKKVSMQILELMANA 632
>TC19363 similar to UP|Q39224 (Q39224) SRG1 mRNA (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (23%)
Length = 552
Score = 69.3 bits (168), Expect = 4e-13
Identities = 44/162 (27%), Positives = 82/162 (50%), Gaps = 3/162 (1%)
Frame = +2
Query: 4 VKTLKYLAQEKTLE--SSFIRDVSERPKVAYNNFSN-EIPIISLAGIDDVDGLRIETCNK 60
V +++ LA++ +E ++R + VA N+ S+ ++P+I L + D ++ K
Sbjct: 74 VPSVQELAKQHMIELPEQYLRPNQDPINVAPNSTSSPQVPVIDLNKLLSEDATELQ---K 244
Query: 61 IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHL 120
+ AC+ WG FQ+++HGV L+ + + FF LP EEK +F + GF + +
Sbjct: 245 LHSACKEWGFFQLINHGVKPSLVENVKIGVQDFFSLPMEEKKKFWQTPEDIEGFGQAFVV 424
Query: 121 QGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYS 162
+ DW ++ + P R+ +PN P+ ++ E YS
Sbjct: 425 SEDQKLDWADLFFINTLPSDARNPRLFPNIPQPLRDNVESYS 550
>TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carboxylate
oxidase homolog (ACC oxidase) , partial (97%)
Length = 1597
Score = 65.9 bits (159), Expect = 5e-12
Identities = 47/164 (28%), Positives = 82/164 (49%)
Frame = +1
Query: 26 ERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISE 85
ER + NF P++ ++ ++ + R+ T I +ACENWG F++V+H + + +
Sbjct: 28 EREEKKMANF----PVVDMSKLNTEE--RVATMELIKDACENWGFFELVNHEISTEFMDT 189
Query: 86 MTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYS 145
+ RL K + E+ RF KG V S + DW +F + + S
Sbjct: 190 VERLTKEHYKRFMEQ--RFKEMVATKGLETVQSEIDD---LDWES--TFFLRHLPSSNIS 348
Query: 146 RWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
P+ E +++V ++++ KL L+ LL++L E +GLEK L K
Sbjct: 349 EVPDLDEDYRKVMKEFAVKLEKLAENLLDLLCENLGLEKGYLKK 480
>BP033094
Length = 446
Score = 65.5 bits (158), Expect = 6e-12
Identities = 32/119 (26%), Positives = 64/119 (52%), Gaps = 4/119 (3%)
Frame = +1
Query: 14 KTLESSFIRDVSERPKV---AYNNFSNEIPIISLAGIDDVDGLRI-ETCNKIVEACENWG 69
KT+ F+RD++ERP + A ++ +++P+I + + + + K+ ACE WG
Sbjct: 88 KTIPQRFVRDMTERPTLDDTALSSPQSDLPVIDFSKLSKGNKEEVLSELXKLAIACEEWG 267
Query: 70 IFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDW 128
FQ+++H +D L+ + +++ FF P EEK ++ ++ G G+ + + DW
Sbjct: 268 FFQVINHEIDLNLLESIENMSREFFMQPLEEKQKYPMAPGTVQGYGQAXVFSEDQKLDW 444
>AV411854
Length = 427
Score = 64.7 bits (156), Expect = 1e-11
Identities = 38/117 (32%), Positives = 60/117 (50%)
Frame = +3
Query: 19 SFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGV 78
+FI+ RP+ + EIPII D+ R E +KI +ACE WG FQ+++HGV
Sbjct: 6 AFIQSTEHRPRATFAEVG-EIPII------DLSESREELISKIGKACEEWGFFQVINHGV 164
Query: 79 DQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYF 135
+ +++ A+ FF+ EEK + G+ H + VRDW+E+ Y+
Sbjct: 165 PSEASTKVELEARKFFEQSIEEKKKVKRDEVNAIGYYDGEHTKN--VRDWKEVFDYY 329
>TC16060 weakly similar to UP|Q9SHK4 (Q9SHK4) F12K11.6, partial (8%)
Length = 835
Score = 64.7 bits (156), Expect = 1e-11
Identities = 47/165 (28%), Positives = 73/165 (43%), Gaps = 16/165 (9%)
Frame = +2
Query: 39 IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPP 98
+PII L I + LR ++I ACE WG F + +HG+ L+ +M + F + P
Sbjct: 362 VPIIDLKDIHNNPALRAAVIDQIRSACEEWGFFLVTNHGIPTTLLVDMIDGIRRFHEQVP 541
Query: 99 EEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRD-YSRW---------- 147
E + ++ RD +YFS RD YS W
Sbjct: 542 EVRKQY-------------------YTRDLTSKTLYFSNASLYRDKYSNWRDTVGCLMAP 664
Query: 148 -PNKPEG----WKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
P KPE ++++ +Y ++ +L K+LE+LSEA+GL L
Sbjct: 665 KPPKPEELPHVFRDIIIEYFNQVKALGYKMLELLSEALGLNPSFL 799
>TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-carboxylic
acid oxidase, complete
Length = 1249
Score = 63.2 bits (152), Expect = 3e-11
Identities = 44/151 (29%), Positives = 79/151 (52%)
Frame = +3
Query: 40 PIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPE 99
P+ISL ++ + I+ +I +ACENWG F++V+HG+ ++ + RL K + + E
Sbjct: 51 PVISLEKLNGEERKDIKL--QIKDACENWGFFELVNHGIPHDVMDTVERLTKEHYRICME 224
Query: 100 EKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTE 159
+ RF KG V + ++ DW P + + S P+ + +++V +
Sbjct: 225 Q--RFKELMASKGLEAVQTEVKD---MDWESTFHLRHLP--ESNISEIPDLTDEYRKVMK 383
Query: 160 QYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
++ ++ LS LL++L E +GLEK L KA
Sbjct: 384 DFALRIEKLSEDLLDLLCENLGLEKGYLKKA 476
>TC18625 weakly similar to UP|O48631 (O48631) Ethylene-forming-enzyme-like
dioxygenase, partial (33%)
Length = 599
Score = 61.6 bits (148), Expect = 9e-11
Identities = 35/130 (26%), Positives = 59/130 (44%)
Frame = +3
Query: 36 SNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFD 95
S IP++ L + + E K+ A WG FQ ++HG+ + ++ ++K FFD
Sbjct: 207 SEGIPVVDLHLLTSPSTAQQELA-KLHYALSTWGCFQAINHGMPSSFLDKVREVSKQFFD 383
Query: 96 LPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWK 155
LP EEK ++ G+ L DW + + P QR+ WP KP ++
Sbjct: 384 LPKEEKQKYAREPNGLEGYGNDQILIQNQRLDWTDRVYLKVQPEDQRNLKVWPQKPNEFR 563
Query: 156 EVTEQYSEKL 165
+Y++ L
Sbjct: 564 STIFEYTKNL 593
>TC15946 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (56%)
Length = 1252
Score = 58.2 bits (139), Expect = 9e-10
Identities = 42/158 (26%), Positives = 72/158 (44%), Gaps = 3/158 (1%)
Frame = +1
Query: 33 NNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKG 92
N+ + +P+I D+ E KI AC WG FQ+++HG+ ++ EM +
Sbjct: 211 NSKLSSVPLI------DLTDEHSEVIGKIRSACHEWGFFQVINHGIPTSVLDEMIDGIRR 372
Query: 93 FFDLPPEEKLRF---DLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPN 149
F + E + +F DL KK + + L +WR+ + P D +
Sbjct: 373 FHEQDSEVRKQFHSRDLK--KKVMYYTNISLFSGQAANWRDTFGFAVAP----DPPKPDE 534
Query: 150 KPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
P +++ +YS+K+ L +LE+LSE +GL L
Sbjct: 535 IPLVCRDIVMEYSKKIKELGFTILELLSEGLGLNPSYL 648
>TC18678 weakly similar to UP|Q9LY48 (Q9LY48) Leucoanthocyanidin
dioxygenase-like protein, partial (28%)
Length = 451
Score = 57.4 bits (137), Expect = 2e-09
Identities = 40/136 (29%), Positives = 63/136 (45%), Gaps = 12/136 (8%)
Frame = +3
Query: 15 TLESSFIRDVSERP-KVAY-NNFSNE---------IPIISLAGI-DDVDGLRIETCNKIV 62
++ FI+ S+RP K + NN ++E IP+I L + D L ET ++
Sbjct: 39 SIPERFIKPTSQRPTKTTFTNNHASEVHDDENTINIPVIDLQHVYGDDSRLCEETLKRVS 218
Query: 63 EACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQG 122
EAC WG FQ+V+HGV +L+ + + FF LP E K + S G+ ++
Sbjct: 219 EACREWGFFQVVNHGVSHELMKHAREVWREFFHLPHEVKEDYANSPTTYEGYGSRLGVKK 398
Query: 123 EPVRDWREMMIYFSYP 138
+ DW + P
Sbjct: 399 GAILDWSDYFFLHYMP 446
>BP045009
Length = 594
Score = 57.0 bits (136), Expect = 2e-09
Identities = 33/129 (25%), Positives = 65/129 (49%), Gaps = 3/129 (2%)
Frame = +1
Query: 58 CNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIV 116
CNK+ EACE WG F+I++H + L++EM + DLP E K+R D++ G
Sbjct: 64 CNKLREACEKWGCFRIINHSIPATLMAEMKTVIGTLLDLPMEIKMRNIDVAAG------- 222
Query: 117 SSHLQGEPVRDWREMMIYFSYPIKQ--RDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLE 174
S ++ E + + Q D+ + +++ E+Y + + L+ K+ +
Sbjct: 223 SGYMAPSAANPLYEALGLYDLGSSQAVHDFCSQLDVSPHQRQIMEKYGKAIHDLAAKVGQ 402
Query: 175 VLSEAMGLE 183
++E++G++
Sbjct: 403 KMAESLGIQ 429
>TC18305 weakly similar to UP|Q9SB33 (Q9SB33) SRG1-like protein, partial
(16%)
Length = 441
Score = 56.6 bits (135), Expect = 3e-09
Identities = 30/91 (32%), Positives = 51/91 (55%), Gaps = 2/91 (2%)
Frame = +1
Query: 25 SERPKVAYNNFS-NEIPIISLAGI-DDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKL 82
++ P V +N S ++P+I + + DG +E ++ AC+ WG FQ+++HGV+ L
Sbjct: 160 NQEPPVIFNTISLPQVPVIDFNKLFSEDDGTELE---RLDHACKEWGFFQLINHGVNPSL 330
Query: 83 ISEMTRLAKGFFDLPPEEKLRFDLSGGKKGG 113
+ +M + FF+LP EEK F G+ G
Sbjct: 331 VEKMKMDVQKFFNLPKEEKKVFSQKPGEIEG 423
>TC11971 weakly similar to UP|Q43419 (Q43419) ACC oxidase, partial (69%)
Length = 730
Score = 40.4 bits (93), Expect(2) = 4e-09
Identities = 20/57 (35%), Positives = 34/57 (59%)
Frame = +2
Query: 39 IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFD 95
IPII + ++ R ET + EAC+NWG F I +HG+D+ L+ ++ +L ++
Sbjct: 8 IPIIDFSTLNG--DKRGETMALLHEACKNWGCFLIENHGIDKSLMEKVKQLINTHYE 172
Score = 35.4 bits (80), Expect(2) = 4e-09
Identities = 20/64 (31%), Positives = 34/64 (52%)
Frame = +3
Query: 127 DWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEA 186
DW + P + + + N E ++ +QY EKL+ L+ L ++SE +GLEK+
Sbjct: 249 DWESAFFIWHRP--KSNIKQISNLSEELRQTMDQYIEKLIQLAEDLSGLMSENLGLEKDY 422
Query: 187 LTKA 190
+ KA
Sbjct: 423 IKKA 434
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,311,854
Number of Sequences: 28460
Number of extensions: 43286
Number of successful extensions: 241
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 233
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 233
length of query: 195
length of database: 4,897,600
effective HSP length: 85
effective length of query: 110
effective length of database: 2,478,500
effective search space: 272635000
effective search space used: 272635000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC138453.15


