

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.19 - phase: 0
(881 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
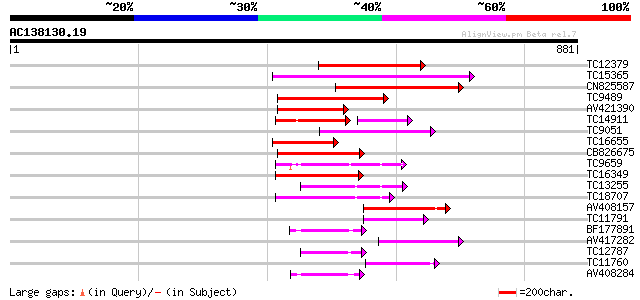
Score E
Sequences producing significant alignments: (bits) Value
TC12379 homologue to UP|O04327 (O04327) Cell division protein Ft... 277 5e-75
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 185 2e-47
CN825587 174 4e-44
TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase... 152 3e-37
AV421390 125 2e-29
TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), pa... 91 5e-29
TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 ... 122 2e-28
TC16655 homologue to UP|O99018 (O99018) Chloroplast protease pre... 119 2e-27
CB826675 116 2e-26
TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATP... 116 2e-26
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 109 2e-24
TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%) 104 5e-23
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 101 5e-22
AV408157 101 6e-22
TC11791 UP|PRSA_LYCES (P54776) 26S protease regulatory subunit 6... 82 4e-16
BF177891 80 1e-15
AV417282 79 4e-15
TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (T... 77 2e-14
TC11760 homologue to UP|PRS4_ORYSA (P46466) 26S protease regulat... 69 3e-12
AV408284 64 8e-11
>TC12379 homologue to UP|O04327 (O04327) Cell division protein FtsH isolog,
partial (15%)
Length = 505
Score = 277 bits (709), Expect = 5e-75
Identities = 143/165 (86%), Positives = 151/165 (90%)
Frame = +1
Query: 481 SISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDAT 540
S+S S + + + + S R+LYQEAKENAPSVVFIDELDAVGR+RGLIKGSGGQERDAT
Sbjct: 4 SLSLSLSLSLSLSLSLSLSRALYQEAKENAPSVVFIDELDAVGRERGLIKGSGGQERDAT 183
Query: 541 LNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVH 600
LNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG IGRIEILKVH
Sbjct: 184 LNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRIEILKVH 363
Query: 601 ARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTE 645
ARKKPIA DVDY VASMTDGMVGAELANIVEVAAINMMRD+RTE
Sbjct: 364 ARKKPIAGDVDYMAVASMTDGMVGAELANIVEVAAINMMRDTRTE 498
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 185 bits (470), Expect = 2e-47
Identities = 104/315 (33%), Positives = 171/315 (54%), Gaps = 2/315 (0%)
Frame = +3
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLL 467
+E+ D + + GL + E++E+++ H E++ G+ P G+LL GPPG GKTLL
Sbjct: 300 VEKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLL 479
Query: 468 AKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRG 527
A+AVA F +S S+ V+ Y+G G+ VR L+ A+E+APS++F+DE+D++G R
Sbjct: 480 ARAVAHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARM 659
Query: 528 LI-KGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIP 586
G G E T+ +LL LDGFE ++ + +TNR DILD AL+RPGR DRKI P
Sbjct: 660 ESGSGXGDSEVQRTMLELLNQLDGFEASNKIKVLMATNRIDILDQALLRPGRIDRKIEFP 839
Query: 587 KPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEV 646
P R +ILK+H+R+ + +D + +A +G GAEL + A + +R+ R V
Sbjct: 840 NPNEESRQDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHV 1019
Query: 647 STDDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAG 706
+ +D A + ++ ++ W+ + + + + ++ ++I AG
Sbjct: 1020TQEDFEMAVAKVMKKETEKNMSLRKLWK*FVFHVSFLCTLSCKDEKRTHLRVVSIYSHAG 1199
Query: 707 RELGYVRTMLESINF 721
YV ++ I F
Sbjct: 1200FHFAYVDQVMYQIMF 1244
>CN825587
Length = 663
Score = 174 bits (442), Expect = 4e-44
Identities = 92/201 (45%), Positives = 132/201 (64%), Gaps = 2/201 (0%)
Frame = +2
Query: 506 AKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNR 565
AK AP +VFIDE+DAVGR+RG G G ER+ T+NQLL +DGF G VI +A+TNR
Sbjct: 2 AKSKAPCIVFIDEIDAVGRQRGAGLGGGNDEREQTINQLLTEMDGFSGNSGVIVLAATNR 181
Query: 566 PDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGA 625
PD+LD AL+RPGRFDR++ + +P GR++IL+VH+R K +A+DVD++ +A T G GA
Sbjct: 182 PDVLDSALLRPGRFDRQVTVDRPDVAGRVKILQVHSRGKALAKDVDFDKIARRTPGFTGA 361
Query: 626 ELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAM 683
+L N++ AAI R E+S D++ A + G ++K S+EK + VA +EA
Sbjct: 362 DLQNLMNEAAILAARRDLKEISKDEISDALERIIAGP-EKKNAVVSEEKKKLVAYHEAGH 538
Query: 684 AVAAMNLPNFDNIEYITIAPR 704
A+ +P +D + I+I PR
Sbjct: 539 ALVGALMPEYDPVAKISIIPR 601
>TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase subunit
RPT2a, partial (69%)
Length = 916
Score = 152 bits (383), Expect = 3e-37
Identities = 81/173 (46%), Positives = 109/173 (62%), Gaps = 1/173 (0%)
Frame = +1
Query: 417 FTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL E++E V+ TH E+Y G+K P G++L G PG GKTLLAKAVA
Sbjct: 379 YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 558
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F + S+ ++ Y+G G VR L++ A + +PS+VFIDE+DAVG KR G +
Sbjct: 559 SATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRYDAHSGGER 738
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
E T+ +LL LDGF+ RG+V I +TNR + LDPAL+RPGR DRKI P P
Sbjct: 739 EIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDRKIEFPLP 897
>AV421390
Length = 409
Score = 125 bits (315), Expect = 2e-29
Identities = 63/110 (57%), Positives = 80/110 (72%)
Frame = +2
Query: 417 FTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAG 476
F DV G + ELEE+V++ + + R G K+P GILL G PG GKTLLAKA+AGEAG
Sbjct: 68 FKDVKGCDDAKQELEEVVEYLRNPAKFTRLGGKLPKGILLTGAPGTGKTLLAKAIAGEAG 247
Query: 477 VNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKR 526
V FF + S+F E++VGVGA RVRSL+Q AK+ AP ++FIDE+DAVG R
Sbjct: 248 VPFFYRAGSEFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGSTR 397
>TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), partial (32%)
Length = 1418
Score = 90.5 bits (223), Expect(2) = 5e-29
Identities = 44/117 (37%), Positives = 72/117 (60%), Gaps = 1/117 (0%)
Frame = +2
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+VK+ DV GL ++ + + V+ H +++ G++ G+LL GPPG GKTLLAKAVA
Sbjct: 212 NVKWEDVGGLEDVKKSILDTVQLPLLHKDLFAS-GLRKRSGVLLYGPPGTGKTLLAKAVA 388
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLI 529
E +NF S+ + + +Y+G VR ++Q+A+ P V+F DELD++ G++
Sbjct: 389 TECSLNFLSVKGPELINMYIGESEKNVRDIFQKARSARPCVIFFDELDSLAPASGML 559
Score = 55.1 bits (131), Expect(2) = 5e-29
Identities = 32/89 (35%), Positives = 50/89 (55%), Gaps = 3/89 (3%)
Frame = +3
Query: 541 LNQLLVCLDGF-EGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG-RIEILK 598
L+Q+L +DG + ++ I ++NRPD++DPAL+RPGRFD+ +++ R +LK
Sbjct: 597 LSQMLAEIDGLSDSSQDLFIIGASNRPDLIDPALLRPGRFDKLLYVGVNSDASYRERVLK 776
Query: 599 VHARKKPIAEDVD-YEIVASMTDGMVGAE 626
RK + EDV Y I GA+
Sbjct: 777 ALTRKFKLHEDVSLYSIAKKCPPNFTGAD 863
>TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (46%)
Length = 867
Score = 122 bits (306), Expect = 2e-28
Identities = 64/180 (35%), Positives = 98/180 (53%)
Frame = +2
Query: 482 ISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATL 541
+ S+ V+ YVG GA VR L+Q A+ +VF DE+DA+G R G E T+
Sbjct: 8 VIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAIGGARFDDGVGGDNEVQRTM 187
Query: 542 NQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHA 601
+++ LDGF+ RG + + +TNRPD LDPAL+RPGR DRK+ P R +I K+H
Sbjct: 188 LEIVNQLDGFDARGNIKVLMATNRPDTLDPALLRPGRLDRKVEFGLPDLESRTQIFKIHT 367
Query: 602 RKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERG 661
R D+ +E+++ + GA++ ++ A + +R R V+ D L A +G
Sbjct: 368 RTMNCERDIRFELLSRLCPNSTGADIRSVCTEAGMYAIRARRKTVTEKDFLDAVNKVIKG 547
>TC16655 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (39%)
Length = 934
Score = 119 bits (299), Expect = 2e-27
Identities = 59/102 (57%), Positives = 76/102 (73%)
Frame = +3
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + + E+V+F E + G +IP G+LL GPPG GKTLLA
Sbjct: 627 MEPNTGVTFDDVAGVDEAKQDFMEVVEFLKKPERFTAVGARIPKGVLLIGPPGTGKTLLA 806
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENA 510
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENA
Sbjct: 807 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENA 932
>CB826675
Length = 551
Score = 116 bits (290), Expect = 2e-26
Identities = 62/136 (45%), Positives = 87/136 (63%), Gaps = 1/136 (0%)
Frame = +1
Query: 417 FTDVAGLGKIRLEL-EEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ DV GL K EL E IV TH E +++ GV+ P G+LL GPPG GKTL+A+A A +
Sbjct: 133 YNDVGGLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLMARACAAQT 312
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F ++ Q V++++G GA VR +Q AKE +P ++FIDE+DA+G KR + SG +
Sbjct: 313 NATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEVSGDR 492
Query: 536 ERDATLNQLLVCLDGF 551
E T+ +LL LDGF
Sbjct: 493 EVQRTMLELLNQLDGF 540
>TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (74%)
Length = 1060
Score = 116 bits (290), Expect = 2e-26
Identities = 81/213 (38%), Positives = 122/213 (57%), Gaps = 10/213 (4%)
Frame = +1
Query: 414 DVKFTDVAGLGKIRLELEEIV-------KFFTHGEMYRRRGVKIPGGILLCGPPGVGKTL 466
++K+ DVAGL + L+E V +FFT +RR + LL GPPG GK+
Sbjct: 451 NIKWNDVAGLESAKQSLQEAVILPVKFPQFFTG----KRRPWR---AFLLYGPPGTGKSY 609
Query: 467 LAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKR 526
LAKAVA EA FFS+S+S ++G V +L+Q A+E+APS++F+DE+D++ +R
Sbjct: 610 LAKAVATEADSTFFSVSSSDLGSKWMGESEKLVSNLFQMARESAPSIIFVDEIDSLCGQR 789
Query: 527 GLIKGSGGQERDATLNQLLVCLDGFEGRGE-VITIASTNRPDILDPALVRPGRFDRKIFI 585
G +G+ + +LLV + G + V+ +A+TN P LD A+ R RFD++I+I
Sbjct: 790 G--EGNESEASRRIKTELLVQMQGVGNNDQKVLVLAATNTPYALDQAIRR--RFDKRIYI 957
Query: 586 PKPGFIGRIEILKVHARKKP--IAEDVDYEIVA 616
P P R + KVH P +AE D+E +A
Sbjct: 958 PLPDVKARQHMFKVHLGDTPHNLAES-DFEHLA 1053
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 109 bits (272), Expect = 2e-24
Identities = 57/138 (41%), Positives = 84/138 (60%), Gaps = 1/138 (0%)
Frame = +3
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
DV + D+ G + E+ E V+ TH E+Y++ G+ P G+LL GPPG GKT+LAKAVA
Sbjct: 486 DVTYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAVA 665
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
F + S+FV+ Y+G G VR +++ AKENAP+++FIDE+DA+ R +
Sbjct: 666 NHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAIATARFDAQTG 845
Query: 533 GGQERDATLNQLLVCLDG 550
+E L +LL +DG
Sbjct: 846 ADREVQRILMELLNQMDG 899
>TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%)
Length = 532
Score = 104 bits (260), Expect = 5e-23
Identities = 63/166 (37%), Positives = 96/166 (56%), Gaps = 1/166 (0%)
Frame = +1
Query: 453 GILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPS 512
GILL GPPG GKT+LAKAVA E FF+ISAS V + G V+ L+Q A+ +APS
Sbjct: 40 GILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPS 219
Query: 513 VVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGE-VITIASTNRPDILDP 571
+F+DE+DA+ +RG + R +LL+ +DG E V +A+TN P LD
Sbjct: 220 TIFLDEIDAIISQRGEARSEHEASR-RLKTELLIQMDGLTRTDELVFVLAATNLPWELDA 396
Query: 572 ALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVAS 617
A++R R +++I +P P R+ + + +P E + Y+++ +
Sbjct: 397 AMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVN 528
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 101 bits (252), Expect = 5e-22
Identities = 64/189 (33%), Positives = 106/189 (55%), Gaps = 3/189 (1%)
Frame = +3
Query: 413 VDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAV 471
+ V F D+ + +I+ L+E+V ++++ +K GILL GPPG GKT+LA A+
Sbjct: 186 IGVTFGDIGAMDEIKESLQELVMLPLRRPDLFKGGLLKPCRGILLFGPPGTGKTMLAXAI 365
Query: 472 AGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKG 531
A EAG +F ++S S + G VR+L+ A + AP+++F+DE+D++ +R +
Sbjct: 366 ANEAGASFINVSMSTITSKWFGEDEKNVRALFSLAAKVAPTIIFVDEVDSMLGQRTRVGE 545
Query: 532 SGGQERDATLNQLLVCLDG-FEGRGE-VITIASTNRPDILDPALVRPGRFDRKIFIPKPG 589
+ N+ + DG GE ++ +A+TNRP LD A++R RF+R+I + P
Sbjct: 546 HEAMRK--IKNEFMTHWDGLLTAPGEQILVLAATNRPFDLDEAIIR--RFERRIMVGLPS 713
Query: 590 FIGRIEILK 598
R ILK
Sbjct: 714 VENREMILK 740
>AV408157
Length = 423
Score = 101 bits (251), Expect = 6e-22
Identities = 60/139 (43%), Positives = 84/139 (60%), Gaps = 4/139 (2%)
Frame = +3
Query: 551 FEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKK--PIAE 608
F+ VI + +TNR D+LDPAL RPGRFDR + + P IGR ILKVH +K P+A+
Sbjct: 3 FDSNSAVIVLGATNRADVLDPALRRPGRFDRVVMVEAPDRIGREAILKVHVSRKELPLAK 182
Query: 609 DVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER 668
DVD + +A MT G GA+LAN+V AA+ R ++ V D +QA + G +++K
Sbjct: 183 DVDLDDIARMTTGFTGADLANLVNEAALLAGRKNKVVVERIDFIQAVERSIAG-IEKKTA 359
Query: 669 SKEKWEQ--VAINEAAMAV 685
+ E+ VA +EA AV
Sbjct: 360 KLQGSEKAVVARHEAGHAV 416
>TC11791 UP|PRSA_LYCES (P54776) 26S protease regulatory subunit 6A homolog
(TAT-binding protein homolog 1) (TBP-1)
(Mg(2+)-dependent ATPase 1) (LEMA-1), partial (29%)
Length = 574
Score = 82.0 bits (201), Expect = 4e-16
Identities = 44/101 (43%), Positives = 60/101 (58%)
Frame = +2
Query: 550 GFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAED 609
GF + IA+TNR DILDPAL+R GR DRKI P P R IL++H+RK + D
Sbjct: 2 GFSSDDRIKVIAATNRADILDPALMRSGRLDRKIEFPHPTEEARARILQIHSRKMNVNPD 181
Query: 610 VDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDD 650
V++E +A TD GA+L + A + +R TEV+ +D
Sbjct: 182 VNFEELARSTDDFNGAQLKAVCVEAGMLALRRDATEVNHED 304
>BF177891
Length = 390
Score = 80.5 bits (197), Expect = 1e-15
Identities = 49/123 (39%), Positives = 69/123 (55%), Gaps = 3/123 (2%)
Frame = +1
Query: 435 KFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGV 494
+ FTHG + R VK GILL GPPG GKTL+AKA+A EAG NF S+S S + G
Sbjct: 46 ELFTHGNLLRP--VK---GILLFGPPGTGKTLMAKALATEAGANFISVSGSTLTSKWFGD 210
Query: 495 GASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDAT---LNQLLVCLDGF 551
++L+ A + AP ++F+DE+D++ RG G E +AT N+ + DG
Sbjct: 211 AEKLTKALFSFASKLAPVIIFVDEVDSLLGARG-----GAFEHEATRRMRNEFMAAWDGL 375
Query: 552 EGR 554
+
Sbjct: 376 RSK 384
>AV417282
Length = 407
Score = 78.6 bits (192), Expect = 4e-15
Identities = 52/134 (38%), Positives = 69/134 (50%), Gaps = 2/134 (1%)
Frame = +3
Query: 573 LVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVE 632
LVRPGRFDR + +P P GR +IL+ H K A+DVD I+A T G GA+LAN+V
Sbjct: 3 LVRPGRFDRHVVVPNPDVEGRRQILESHMSKVLKADDVDLMIIARGTPGFSGADLANMVN 182
Query: 633 VAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNL 690
+AA+ D VS DL A G +RK S E + A +E A+ A++
Sbjct: 183 IAALKAAMDGAKAVSMHDLEFAKDKIMMGS-ERKSAVISDESRKMTAFHEGGHALVAIHT 359
Query: 691 PNFDNIEYITIAPR 704
+ TI PR
Sbjct: 360 DGALPVHKATIVPR 401
>TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (TAT-binding
homolog 4), partial (20%)
Length = 389
Score = 76.6 bits (187), Expect = 2e-14
Identities = 43/103 (41%), Positives = 62/103 (59%), Gaps = 1/103 (0%)
Frame = +3
Query: 453 GILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPS 512
GILL GPPG GKT+LAKAVA EAG NF +IS S + G G V++++ A + +PS
Sbjct: 87 GILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKISPS 266
Query: 513 VVFIDELDAVGRKRGLIKGSGGQERDATL-NQLLVCLDGFEGR 554
V+F+DE+D + +R + G E + N+ +V DG +
Sbjct: 267 VIFVDEVDXMLXRR---ENPGEHEAMRKMXNEFMVHWDGLPSK 386
>TC11760 homologue to UP|PRS4_ORYSA (P46466) 26S protease regulatory subunit
4 homolog (TAT-binding protein homolog 2), partial (27%)
Length = 555
Score = 68.9 bits (167), Expect = 3e-12
Identities = 40/114 (35%), Positives = 63/114 (55%)
Frame = +3
Query: 554 RGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYE 613
RG+V I +T+ + LDPAL+RPGR DRKI P P R I ++H + +A+DV+ E
Sbjct: 3 RGDVKVILATHSIESLDPALLRPGRIDRKIEFPLPDIKTRRRIFQIHTSRMTLADDVNLE 182
Query: 614 IVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKE 667
D GA++ I A + +R+ R +V+ D +A +++ M +KE
Sbjct: 183 EFVMTKDEFSGADIKAICTEAGLLALRERRMKVTHADFKKA---KDKVMFKKKE 335
>AV408284
Length = 425
Score = 64.3 bits (155), Expect = 8e-11
Identities = 39/119 (32%), Positives = 67/119 (55%), Gaps = 4/119 (3%)
Frame = +1
Query: 437 FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGA 496
F+HG++ + G+LL GPPG GKT+LAKA+A E+G F ++ S + + G
Sbjct: 64 FSHGKLLGPQK-----GVLLYGPPGTGKTMLAKAIAKESGAVFINVRISNLMSKWFGDAQ 228
Query: 497 SRVRSLYQEAKENAPSVVFIDELDA-VGRKRGLIKGSGGQERDATLN---QLLVCLDGF 551
V +++ A + P+++FIDE+D+ +G++R + +A LN + + DGF
Sbjct: 229 KLVAAVFSLAHKLQPAIIFIDEVDSFLGQRR-------TTDHEALLNMKTEFMALWDGF 384
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,131,337
Number of Sequences: 28460
Number of extensions: 145567
Number of successful extensions: 1048
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 1026
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1041
length of query: 881
length of database: 4,897,600
effective HSP length: 98
effective length of query: 783
effective length of database: 2,108,520
effective search space: 1650971160
effective search space used: 1650971160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC138130.19


