

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.17 + phase: 0 /pseudo
(626 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
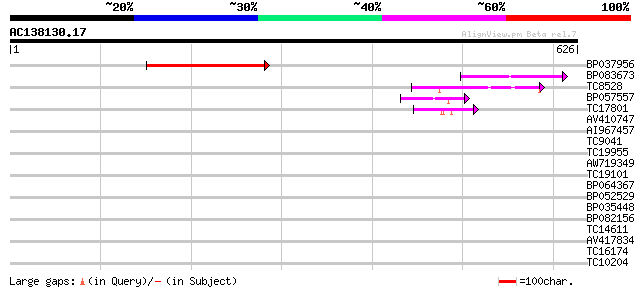
Sequences producing significant alignments: (bits) Value
BP037956 131 3e-31
BP083673 59 2e-09
TC8528 weakly similar to GB|AAO42978.1|28416895|BT004732 At3g139... 56 2e-08
BP057557 52 4e-07
TC17801 weakly similar to GB|AAO42978.1|28416895|BT004732 At3g13... 41 7e-04
AV410747 36 0.021
AI967457 34 0.080
TC9041 similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kinase Xa21-b... 32 0.31
TC19955 32 0.40
AW719349 31 0.68
TC19101 weakly similar to UP|Q93WC9 (Q93WC9) Adenylate isopenten... 29 2.0
BP064367 29 2.0
BP052529 29 2.0
BP035448 29 2.0
BP082156 28 3.4
TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding p... 28 3.4
AV417834 28 4.4
TC16174 similar to UP|Q940N7 (Q940N7) AT4g05150/C17L7_70, partia... 28 5.8
TC10204 homologue to GB|BAB17639.1|11094242|AB044952 cytosolic p... 28 5.8
>BP037956
Length = 476
Score = 131 bits (330), Expect = 3e-31
Identities = 80/136 (58%), Positives = 88/136 (63%)
Frame = +1
Query: 152 ESC*TSGRILFK*STRKGIQHCIMLVIKDTLRLYGSY*VAIQNLLYNTTTMVIHHCILQ* 211
ESC* SG+ L K* T I +CIMLVIK T RL+GSY* IQ LLYNTTTMV HHC Q*
Sbjct: 4 ESC*ISGQXLIK*LTTMEIHYCIMLVIKGTRRLHGSY*DVIQILLYNTTTMVTHHCTWQ* 183
Query: 212 LRGKSQPLTILW**VLRTFTIPQEKKRQYFI*L*DMDVMTL*FSW*ELPMVQI*CTDKIN 271
GKSQ L L VL FTI QEKKRQYF *L* D M +SW + PMV I C KI+
Sbjct: 184 *MGKSQSLRTLCQAVLHHFTISQEKKRQYFT*L*GTDAMMHLYSWCKFPMVPIYCIVKID 363
Query: 272 MATLSYISQFQAVAIR 287
+A + Y Q AIR
Sbjct: 364 LAIVLYTLQL*VAAIR 411
>BP083673
Length = 437
Score = 59.3 bits (142), Expect = 2e-09
Identities = 31/118 (26%), Positives = 65/118 (54%)
Frame = -1
Query: 498 IALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEM 557
IALF SL+VV+V S++ + + ++ I +K+MW+A + ++A ++V++ +E
Sbjct: 428 IALFISLAVVVVQTSVVVIESQAKKQMMAIINKLMWLACVLVSVAFLALSFVVV--GKEE 255
Query: 558 QWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKESGDGTAESDKESEDSD 615
+WL++ + +G + T ++ ++ H + S+ R RK S + + S + SD
Sbjct: 254 KWLAIGVTIIGTTIMATTLGTMCYWVIRHRIEASNLRNIRKSSLESKSRSFSQPALSD 81
>TC8528 weakly similar to GB|AAO42978.1|28416895|BT004732 At3g13950
{Arabidopsis thaliana;}, partial (17%)
Length = 788
Score = 55.8 bits (133), Expect = 2e-08
Identities = 45/171 (26%), Positives = 79/171 (45%), Gaps = 24/171 (14%)
Frame = +2
Query: 444 NARNTIVLVAVLIATVTFAAGISPPGGVYQ-------------------EGPKKGISMAG 484
N R ++ L A +IAT+ F +PPGGV+Q + P + +
Sbjct: 146 NMRGSVSLTASIIATMAFQLATNPPGGVFQANGGDSVAKIKSCLDNDAIQCPGEAVLAVV 325
Query: 485 ETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYV 544
+ +F N I+ +SLSV ++LVS IP + + +L+I M +++ + Y+
Sbjct: 326 NEDDYSLFLTFNTISFISSLSVCLLLVSGIPLKHRVIIWILSIG---MCISITSLALTYL 496
Query: 545 AATWVILPHNQEMQWLSVVLLAL--GGGFLGTIFISLSVML---VEHWLRK 590
A ++ P++ W +V LAL G LG + + L + + V W+ K
Sbjct: 497 VAASMVTPNH---VWGNVFALALIIWIGLLGIVSVYLIIRMIYSVGKWITK 640
>BP057557
Length = 497
Score = 51.6 bits (122), Expect = 4e-07
Identities = 32/80 (40%), Positives = 47/80 (58%), Gaps = 4/80 (5%)
Frame = -1
Query: 432 KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMA----GETS 487
K ++H L NA N+ +VAVLIATV FAA + P G Y E ++ S+ +
Sbjct: 407 KKLKKLHISGLNNAINSATVVAVLIATVAFAAIFTVP-GQYVEAKEQSFSLGPANIANNA 231
Query: 488 AFKVFAISNIIALFTSLSVV 507
AF +F + +I+ALF SL+V+
Sbjct: 230 AFMIFFVFDILALFISLAVI 171
>TC17801 weakly similar to GB|AAO42978.1|28416895|BT004732 At3g13950
{Arabidopsis thaliana;}, partial (22%)
Length = 423
Score = 40.8 bits (94), Expect = 7e-04
Identities = 30/91 (32%), Positives = 43/91 (46%), Gaps = 19/91 (20%)
Frame = +2
Query: 446 RNTIVLVAVLIATVTFAAGISPPGGVYQEG---PK---------KGISMAGET------- 486
R + + A +IAT+TF +PPGGV+Q PK I GE
Sbjct: 149 RGGLSMTASIIATMTFQLATNPPGGVFQANGGVPKTDARICWHNNTIQCPGEAVLGVVYR 328
Query: 487 SAFKVFAISNIIALFTSLSVVIVLVSIIPFR 517
+ F I N ++ +SLSV ++LVS IP +
Sbjct: 329 HTYTNFLICNTLSFISSLSVCLLLVSGIPLK 421
>AV410747
Length = 312
Score = 35.8 bits (81), Expect = 0.021
Identities = 15/30 (50%), Positives = 22/30 (73%)
Frame = +3
Query: 444 NARNTIVLVAVLIATVTFAAGISPPGGVYQ 473
N R ++ L+A +IAT+TF +PPGGV+Q
Sbjct: 150 NMRGSLSLMASIIATMTFQLATNPPGGVFQ 239
>AI967457
Length = 278
Score = 33.9 bits (76), Expect = 0.080
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +3
Query: 107 AFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGH 150
A A GHLD+V LL S + + +GFD H+AA +GH
Sbjct: 3 ALFTAAERGHLDVVKELLKHSNLKKKNRSGFDP--LHIAASQGH 128
>TC9041 similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kinase Xa21-binding
protein 3, partial (43%)
Length = 848
Score = 32.0 bits (71), Expect = 0.31
Identities = 25/120 (20%), Positives = 53/120 (43%)
Frame = +1
Query: 6 FDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSA 65
FDA+ ++ ++ +E +L + +PLH+A+ G IE++S ++ V
Sbjct: 181 FDAVANGELEVVEAMAEEDVTVLEHTLGRSRLSPLHVAAVNGRIEVLSMLLDR-NVKVDV 357
Query: 66 ENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLN 125
N++ +TP+ A + L++ + ++ A GHLD + +L+
Sbjct: 358 LNRHKQTPLMLAVIHGRTGCVEKLIDAGANILMFDSLRRRTCLHYAAYFGHLDCLKAILS 537
>TC19955
Length = 548
Score = 31.6 bits (70), Expect = 0.40
Identities = 11/28 (39%), Positives = 19/28 (67%)
Frame = -1
Query: 584 VEHWLRKSSWRKKRKESGDGTAESDKES 611
++HW+ K ++KRKE G+G E K++
Sbjct: 251 IQHWI*KKVKKEKRKEQGEGVGERKKQA 168
>AW719349
Length = 143
Score = 30.8 bits (68), Expect = 0.68
Identities = 12/40 (30%), Positives = 21/40 (52%), Gaps = 2/40 (5%)
Frame = -3
Query: 401 YNPYYFSPTNLVKQKHH--HNKGKIENVNHTKRKHYHEMH 438
+ PY+ S L+ HH H+ +++H+ R H+H H
Sbjct: 123 FPPYFHSHHTLLHSLHHSHHSSSYFHSLHHSLRNHHHSHH 4
>TC19101 weakly similar to UP|Q93WC9 (Q93WC9) Adenylate
isopentenyltransferase (Cytokinin synthase)
(AT3g63110/T20O10_210) , partial (37%)
Length = 880
Score = 29.3 bits (64), Expect = 2.0
Identities = 21/60 (35%), Positives = 34/60 (56%), Gaps = 5/60 (8%)
Frame = +1
Query: 328 KRSTPSSFSLELDNTSSPSPTSR-HSLSRRYISKEMEVLTEMVSYDCIS----PPPVSES 382
K STP+ F +TSSP+ + R +++ S E++ LT+ S IS PPP+++S
Sbjct: 538 KSSTPTRFKSTTASTSSPTKSPRKNNVESPTTSSELKTLTQ-TSPPAISVTVPPPPLTQS 714
>BP064367
Length = 518
Score = 29.3 bits (64), Expect = 2.0
Identities = 22/65 (33%), Positives = 33/65 (49%), Gaps = 3/65 (4%)
Frame = -2
Query: 328 KRSTPSSFSLELDNTSSP--SPTSRHSLSRR-YISKEMEVLTEMVSYDCISPPPVSESTE 384
+R++ S S + ++SSP S +S SL + Y + S DC SPPP + S
Sbjct: 475 RRNSSSLDSGDKSSSSSPPNSTSSXTSLPKNAYCDASKTSPSVGTSPDCTSPPPPNTSMS 296
Query: 385 SISPQ 389
SI P+
Sbjct: 295 SIKPK 281
>BP052529
Length = 394
Score = 29.3 bits (64), Expect = 2.0
Identities = 24/63 (38%), Positives = 33/63 (52%), Gaps = 5/63 (7%)
Frame = +3
Query: 337 LELDNTSSP----SPTSRHSLSRRYISKE-MEVLTEMVSYDCISPPPVSESTESISPQPQ 391
L+LDN P +PT HS R +IS E ME + E+ PPP+S S+ +SP +
Sbjct: 3 LKLDNLLHPRKP*NPT--HS-ERNFISGEKMEKVPEIEDPADAEPPPLSPSSPDVSPSIE 173
Query: 392 VSE 394
E
Sbjct: 174 TVE 182
>BP035448
Length = 509
Score = 29.3 bits (64), Expect = 2.0
Identities = 22/75 (29%), Positives = 35/75 (46%), Gaps = 6/75 (8%)
Frame = -1
Query: 33 DDTFSAPLHLASKYGCIEMVSEIVKLCPD------MVSAENKNMETPIHEACRQENVKVL 86
D+ + PLH A G ++V I+ M+ + + +TP+H A R E+ V+
Sbjct: 461 DEDGAIPLHDACAGGFTQIVQLILHSANGTEHIKRMLESIDSEGDTPLHHAARGEHADVI 282
Query: 87 MLLLEVNPTAACKLN 101
LLL N + K N
Sbjct: 281 RLLLS-NGASTTKTN 240
>BP082156
Length = 515
Score = 28.5 bits (62), Expect = 3.4
Identities = 17/56 (30%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Frame = +3
Query: 370 SYDCISPPPVSESTESISPQPQVSERFENGTYNPYYFS-PTNLVKQKHHHNKGKIE 424
S+ +SP P S S+ S+ P P E+ ++ FS K +HH+K +E
Sbjct: 291 SWTLLSPFPASFSSWSLPPPPSSHSSQEDNQHHQTCFSFQEKKKKTNYHHHKKTVE 458
>TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding protein,
partial (20%)
Length = 760
Score = 28.5 bits (62), Expect = 3.4
Identities = 10/32 (31%), Positives = 18/32 (56%)
Frame = -2
Query: 416 HHHNKGKIENVNHTKRKHYHEMHKEALLNARN 447
HHH++ ++++ HYH H +AL A +
Sbjct: 333 HHHHQSHHLHISYEHSPHYHHHHHQALEGAHD 238
>AV417834
Length = 420
Score = 28.1 bits (61), Expect = 4.4
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = +2
Query: 323 IRDGGKRSTPSSFSLELDNTSSPSPTS 349
I++GG TP+SF +D SPSP +
Sbjct: 338 IQEGGSAPTPTSFPQIIDKHFSPSPAA 418
>TC16174 similar to UP|Q940N7 (Q940N7) AT4g05150/C17L7_70, partial (20%)
Length = 491
Score = 27.7 bits (60), Expect = 5.8
Identities = 19/68 (27%), Positives = 30/68 (43%)
Frame = +2
Query: 326 GGKRSTPSSFSLELDNTSSPSPTSRHSLSRRYISKEMEVLTEMVSYDCISPPPVSESTES 385
GG+ P S L T SP P + S++RR+ S S +P +S +T +
Sbjct: 113 GGEWQHPHSTHLS*TLTQSPPPLAPTSMTRRHASASWSASAARSS---PAPTTISSATSA 283
Query: 386 ISPQPQVS 393
+P+ S
Sbjct: 284 ATPESSPS 307
>TC10204 homologue to GB|BAB17639.1|11094242|AB044952 cytosolic
phosphoglucose isomerase {Arabidopsis thaliana;} ,
partial (36%)
Length = 734
Score = 27.7 bits (60), Expect = 5.8
Identities = 13/38 (34%), Positives = 21/38 (55%), Gaps = 3/38 (7%)
Frame = -2
Query: 562 VVLLALGGGFLGTIFISLSVML---VEHWLRKSSWRKK 596
VV + +GG FLG +F+ ++ + RK SWR +
Sbjct: 115 VVAVGIGGSFLGPLFVHTALQTDPEAIEFARKRSWRPR 2
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.330 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,518,681
Number of Sequences: 28460
Number of extensions: 154036
Number of successful extensions: 1337
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 1309
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1331
length of query: 626
length of database: 4,897,600
effective HSP length: 96
effective length of query: 530
effective length of database: 2,165,440
effective search space: 1147683200
effective search space used: 1147683200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC138130.17


