

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.15 + phase: 0
(174 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
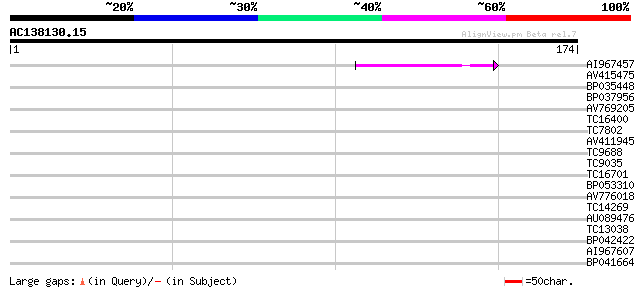
Score E
Sequences producing significant alignments: (bits) Value
AI967457 40 3e-04
AV415475 30 0.23
BP035448 30 0.30
BP037956 27 1.5
AV769205 27 2.5
TC16400 similar to UP|O48783 (O48783) At2g26680 protein, partial... 27 2.5
TC7802 homologue to UP|Q32904 (Q32904) Light harvesting protein,... 26 3.3
AV411945 26 4.3
TC9688 25 5.6
TC9035 similar to UP|Q9FLZ8 (Q9FLZ8) Emb|CAB87751.1 (AT5g39360/M... 25 5.6
TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome ... 25 5.6
BP053310 25 7.3
AV776018 25 7.3
TC14269 weakly similar to UP|Q13484 (Q13484) Ankyrin G119, parti... 25 7.3
AU089476 25 7.3
TC13038 25 7.3
BP042422 25 9.6
AI967607 25 9.6
BP041664 25 9.6
>AI967457
Length = 278
Score = 39.7 bits (91), Expect = 3e-04
Identities = 21/44 (47%), Positives = 26/44 (58%)
Frame = +3
Query: 107 AFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
A F A GHLD+V LL S + + +GFD HIAAS+GH
Sbjct: 3 ALFTAAERGHLDVVKELLKHSNLKKKNRSGFDP--LHIAASQGH 128
>AV415475
Length = 427
Score = 30.0 bits (66), Expect = 0.23
Identities = 20/62 (32%), Positives = 33/62 (52%)
Frame = +2
Query: 28 LNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLM 87
L + TD TPLH+AS +G + + +V D V+A++ TP+ +A + V+
Sbjct: 158 LVKATDYDKRTPLHVASLHGWLAVAKCLVEFGAD-VNAQDRWKNTPLADAEGAKRTAVIE 334
Query: 88 LL 89
LL
Sbjct: 335 LL 340
>BP035448
Length = 509
Score = 29.6 bits (65), Expect = 0.30
Identities = 23/85 (27%), Positives = 38/85 (44%), Gaps = 6/85 (7%)
Frame = -1
Query: 23 VREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPD------MVSAENENMETPIHE 76
++ G + D+ PLH A G ++V I+ M+ + + +TP+H
Sbjct: 491 IQRGANLEAQDEDGAIPLHDACAGGFTQIVQLILHSANGTEHIKRMLESIDSEGDTPLHH 312
Query: 77 ACRQENVKVLMLLLEVNPTAACKLN 101
A R E+ V+ LLL N + K N
Sbjct: 311 AARGEHADVIRLLLS-NGASTTKTN 240
>BP037956
Length = 476
Score = 27.3 bits (59), Expect = 1.5
Identities = 16/56 (28%), Positives = 24/56 (42%)
Frame = +3
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVN 93
TPLHLA G + ++ + V C ET H A R + L++V+
Sbjct: 162 TPLHLAVMNGKVSILEDFVSSCAASFHYLTREEETIFHLAVRYGCYDAFVFLVQVS 329
>AV769205
Length = 600
Score = 26.6 bits (57), Expect = 2.5
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +1
Query: 86 LMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVN 121
L+LLL + PTA+ S+ A SH H +LV+
Sbjct: 190 LLLLLPMRPTASHATTVASASSTRAASSHHHCELVS 297
>TC16400 similar to UP|O48783 (O48783) At2g26680 protein, partial (58%)
Length = 1053
Score = 26.6 bits (57), Expect = 2.5
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = -3
Query: 96 AACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEI 129
A+C L C + CSH HL ++L+ + SEI
Sbjct: 808 ASC*LCHVCSCQPVLICSHQHL*SIHLIFHHSEI 707
>TC7802 homologue to UP|Q32904 (Q32904) Light harvesting protein, complete
Length = 1268
Score = 26.2 bits (56), Expect = 3.3
Identities = 13/23 (56%), Positives = 15/23 (64%)
Frame = +2
Query: 150 HTGENKEFSLLCLHVFLTLLSTL 172
H E + FS+LCLH FLT S L
Sbjct: 1187 HVKELELFSILCLHRFLTS*SIL 1255
>AV411945
Length = 405
Score = 25.8 bits (55), Expect = 4.3
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 7/54 (12%)
Frame = -2
Query: 86 LMLLLEVNPTAACKLNPTCKSAFF----VACSHGHLD---LVNLLLNLSEIVEP 132
L+L ++ + CKL+P C F +A H D L+NLLL L +++ P
Sbjct: 365 LLLKSKIMSSFLCKLSPQCLKLMFHIHDLAWLMQHFDSSSLLNLLL*LFQVLLP 204
>TC9688
Length = 715
Score = 25.4 bits (54), Expect = 5.6
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = +1
Query: 48 CIEMVSEIVRLCPDMVSAENENMETPIHEACRQEN 82
C+EM S++ L D S E + ETP +E + E+
Sbjct: 205 CLEMKSQLFNLEVDSGSIEILDHETPENEIAKSES 309
>TC9035 similar to UP|Q9FLZ8 (Q9FLZ8) Emb|CAB87751.1 (AT5g39360/MUL8_40),
partial (34%)
Length = 593
Score = 25.4 bits (54), Expect = 5.6
Identities = 20/77 (25%), Positives = 32/77 (40%), Gaps = 2/77 (2%)
Frame = +3
Query: 76 EACRQENVKVLMLLLE--VNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPG 133
E C K+ +LL + +A+C+L + + C +GH+ + LL LS+ E
Sbjct: 75 EVCPYCKAKLWSMLLANMIPKSASCRLGSYEECVEYYVCLNGHMLGICTLLPLSDSEEAS 254
Query: 134 LAGFDQACFHIAASRGH 150
D F I H
Sbjct: 255 EKE*DIHNFQIGIQSSH 305
>TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome 5, BAC
clone:F24B18, partial (24%)
Length = 691
Score = 25.4 bits (54), Expect = 5.6
Identities = 22/64 (34%), Positives = 27/64 (41%), Gaps = 2/64 (3%)
Frame = -2
Query: 11 NNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENE-- 68
NN IS S I +R N R D NT + ASK P SAE++
Sbjct: 213 NNSISNNSIIPVLRIANNNNRADTVVNTVIADASKLPVATTADGAKPAAPHDDSAESKTL 34
Query: 69 NMET 72
N+ET
Sbjct: 33 NLET 22
>BP053310
Length = 473
Score = 25.0 bits (53), Expect = 7.3
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +2
Query: 114 HGHLDLVNLLLNLSEIVEPGL 134
H H + L L + EI+EPGL
Sbjct: 173 HSHTNFNKLCLVIKEIIEPGL 235
>AV776018
Length = 476
Score = 25.0 bits (53), Expect = 7.3
Identities = 21/68 (30%), Positives = 32/68 (46%), Gaps = 12/68 (17%)
Frame = -1
Query: 119 LVNL-----LLNLSEIVEPGL-------AGFDQACFHIAASRGHTGENKEFSLLCLHVFL 166
L+NL LL SE+ EP + A ++ C + AAS + + F+ L +L
Sbjct: 407 LINLSRSKSLLEQSELEEPSISVV*AFGASEEEFCSYEAASDLKLSDFRRFNSLETFSWL 228
Query: 167 TLLSTLPF 174
LS+ PF
Sbjct: 227 LFLSSFPF 204
>TC14269 weakly similar to UP|Q13484 (Q13484) Ankyrin G119, partial (3%)
Length = 656
Score = 25.0 bits (53), Expect = 7.3
Identities = 14/55 (25%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Frame = +1
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDM-VSAENENMETPIHEACRQENVKVLMLLLE 91
TPLHLA++ G + ++ E++ ++ + TP+H A ++ + L+E
Sbjct: 76 TPLHLAAEGGHLGVMDELLERGANIDARTKGACGWTPLHIAAKERRKDAVKFLIE 240
>AU089476
Length = 548
Score = 25.0 bits (53), Expect = 7.3
Identities = 13/26 (50%), Positives = 15/26 (57%), Gaps = 4/26 (15%)
Frame = +1
Query: 153 ENKEFSLL----CLHVFLTLLSTLPF 174
EN +S+ CL VFL L TLPF
Sbjct: 253 ENNNYSMSDTIPCLFVFLRLCETLPF 330
>TC13038
Length = 419
Score = 25.0 bits (53), Expect = 7.3
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = +3
Query: 35 TFNTPLHLASKYGCIEMVSEIVRLCPDMVSA 65
T+NT L SKYG + V + + + DM A
Sbjct: 99 TYNTLLKARSKYGSVLEVQQCLAIYQDMQKA 191
>BP042422
Length = 526
Score = 24.6 bits (52), Expect = 9.6
Identities = 19/69 (27%), Positives = 30/69 (42%)
Frame = -2
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTGENKEF 157
CK+ P+ F +A H HL L +D+AC H ++ + N++
Sbjct: 363 CKVCPSHH*PFELAFHHQHL-------------RNHLYYYDKACSHRVEAKR*SFPNRDT 223
Query: 158 SLLCLHVFL 166
L LH+FL
Sbjct: 222 CPLPLHLFL 196
>AI967607
Length = 378
Score = 24.6 bits (52), Expect = 9.6
Identities = 16/61 (26%), Positives = 31/61 (50%)
Frame = +1
Query: 37 NTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTA 96
+TPLHLA++ G ++ + E++ D + + P A + ++ + LL NP +
Sbjct: 142 STPLHLAARGGSLDCIRELLAWGADRLQRDASG-RIPYMVALKHKHGAACVSLL--NPAS 312
Query: 97 A 97
A
Sbjct: 313 A 315
>BP041664
Length = 487
Score = 24.6 bits (52), Expect = 9.6
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = -3
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTA 96
E+ME + E + + VL+LLL NP+A
Sbjct: 206 EDMEGAVDEGRGRVEIVVLLLLLHRNPSA 120
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,278,739
Number of Sequences: 28460
Number of extensions: 48051
Number of successful extensions: 240
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 236
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 238
length of query: 174
length of database: 4,897,600
effective HSP length: 84
effective length of query: 90
effective length of database: 2,506,960
effective search space: 225626400
effective search space used: 225626400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 52 (24.6 bits)
Medicago: description of AC138130.15


