

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
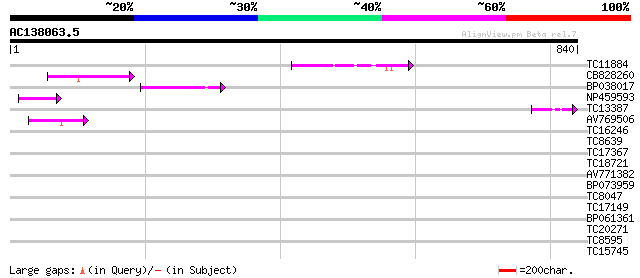
Query= AC138063.5 + phase: 0 /pseudo
(840 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11884 85 4e-17
CB828260 79 2e-15
BP038017 59 4e-09
NP459593 Pge1 protein [Lotus japonicus] 50 2e-06
TC13387 47 1e-05
AV769506 44 1e-04
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 40 0.002
TC8639 similar to UP|FRI4_SOYBN (Q948P5) Ferritin 4, chloroplast... 29 2.7
TC17367 similar to GB|BAB03049.1|9294683|AP001305 syringomycin b... 29 3.5
TC18721 weakly similar to UP|BAD10141 (BAD10141) Zinc finger pro... 28 4.6
AV771382 28 4.6
BP073959 28 6.0
TC8047 similar to UP|Q9ASV9 (Q9ASV9) At1g53210/F12M16_12, partia... 28 6.0
TC17149 similar to GB|AAP37818.1|30725592|BT008459 At1g43860 {Ar... 28 6.0
BP061361 28 7.9
TC20271 weakly similar to UP|Q84XH9 (Q84XH9) RCa4 (Fragment), pa... 28 7.9
TC8595 similar to PIR|T09390|T09390 21K protein precursor - alfa... 28 7.9
TC15745 28 7.9
>TC11884
Length = 557
Score = 85.1 bits (209), Expect = 4e-17
Identities = 58/191 (30%), Positives = 95/191 (49%), Gaps = 10/191 (5%)
Frame = +1
Query: 418 HFGCTTTNGVESAHGTLKGYLEDSKGDLAKGWKAINAMLTVQFTEIQATFGRNSTVLEHK 477
HFG TT+N E AH +LK L D KGDLA W A +++ + TEI A+F R+ ++H
Sbjct: 1 HFGTTTSNRAEGAHASLKKMLRDCKGDLATSWDASHSLTCNRHTEILASFERSIHRIDHI 180
Query: 478 FKGNKMYKSLEYNVSRTAMNYIYKQASRVKDCGSDSKKCGCVIRWTHALPCTCLI--YKK 535
F Y ++ VS + + + R+K G +C C++R TH LPC C + Y++
Sbjct: 181 FM-CPFYTNIRGFVSNKCLQLLDAEHIRMKSYG----RCDCLLRETHELPCGCELAGYER 345
Query: 536 LNLGSCIRLDEINIHWKKLCFE-----VDDDVEE---SNRHVSIQNELNAIQERFNKADY 587
I + I WK+L +E V D + H +Q E+ A+ F+ D
Sbjct: 346 ------ISYEAIYPFWKRLSWEHVPEPVADTISNHICGMNHGDMQPEVEALTHYFSSLDT 507
Query: 588 NMKLIIKEQMR 598
+ +++ +++
Sbjct: 508 GGQSMVRRKLQ 540
>CB828260
Length = 557
Score = 79.3 bits (194), Expect = 2e-15
Identities = 47/132 (35%), Positives = 71/132 (53%), Gaps = 4/132 (3%)
Frame = +3
Query: 57 GSGRRKPKLVLAFERSGEYKGTKKSKR-EGTGSRKCGCPFRLRGY---FSATKLWSLNIV 112
G+G RK ++L E+ G+Y + EGT ++K CPFRL+G + W L ++
Sbjct: 162 GNGSRKAYVMLGCEKHGKYVPYRDPDLVEGTRTQKTECPFRLKGRPMKNGTDRDWRLKVM 341
Query: 113 SGLHNHKMEPKLEGHMLAGRLTAKESKIVGDMTRNLIKPKNILLDLKGRWKDNRTVAKQI 172
G HNH+ L GH GRL ++E + V M+++ + P+ +LL LK N T QI
Sbjct: 342 EGAHNHEPARSLLGHNFVGRLNSEEKEQVEKMSKSWVLPRKMLLTLKENNPLNLTTISQI 521
Query: 173 YNARHRYKLSIR 184
Y A R + S+R
Sbjct: 522 YGACKRLRKSLR 557
>BP038017
Length = 570
Score = 58.5 bits (140), Expect = 4e-09
Identities = 35/127 (27%), Positives = 60/127 (46%), Gaps = 1/127 (0%)
Frame = +2
Query: 194 FQKFEENNYVFNYRT-VSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLF 252
F+K + N F Y + + ++F+A S +N F ++ D+ Y+ NQY++P
Sbjct: 191 FKKMQGKNPGFYYAIQLDDENRMINVFWADARSRSAYNYFGDAVIFDTMYRPNQYQVPFA 370
Query: 253 EIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAV 312
G+ G A L E E +FTW + + + + P I TD+D + AV
Sbjct: 371 PFTGVNHHGQNVLFGCALLLDESESSFTWLFRTWLSAM--NDRPPVSITTDQDRAIQAAV 544
Query: 313 ANIFPKS 319
A +FP++
Sbjct: 545 AQVFPET 565
>NP459593 Pge1 protein [Lotus japonicus]
Length = 633
Score = 49.7 bits (117), Expect = 2e-06
Identities = 24/63 (38%), Positives = 37/63 (58%)
Frame = +1
Query: 14 IDVAEFFVTKEKKKYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSG 73
I +AE TK + + ++I+WAR E G+ +I+ +S GS +RKP + L ER G
Sbjct: 427 IKIAETTATKIAEAFASHTDLIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGG 606
Query: 74 EYK 76
+YK
Sbjct: 607 KYK 615
>TC13387
Length = 511
Score = 47.0 bits (110), Expect = 1e-05
Identities = 23/68 (33%), Positives = 41/68 (59%)
Frame = +1
Query: 773 EEKWLTVPDMGHVIASLYNKVVVVLKYGNNFSETCFPLRGRPPTNPSARIMCLGLIPNHF 832
E+KW+ +P+MG+++A+ + V + + + +S PLRG P P R++ +G + NHF
Sbjct: 4 EDKWMQLPEMGYLVATRFQVVFISISFQGCWS--YLPLRGEGPP-PVHRVIAIGHVINHF 174
Query: 833 VHVFLRPG 840
V + L G
Sbjct: 175 VQLHLTLG 198
>AV769506
Length = 345
Score = 43.9 bits (102), Expect = 1e-04
Identities = 29/100 (29%), Positives = 46/100 (46%), Gaps = 11/100 (11%)
Frame = +1
Query: 28 YEDREEMINWARGQAIEAGFTLIIDKS-YSGSGRRKPKLVLAFERSGEY--------KGT 78
+ R+++INW G AIE G+ +I KS Y G+ RK ++L E+ G+Y
Sbjct: 43 FPTRDDLINWVHGIAIENGYVTLITKSDYGGNESRKAYVMLGCEKHGKYVPYRDPDLVAD 222
Query: 79 KKSKREGTGSRKCGC--PFRLRGYFSATKLWSLNIVSGLH 116
+ + T C C +G + +LW + V G H
Sbjct: 223 EDPSAQSTLEPACDCLNHHHKQGSKPSPQLWPPSTVDGPH 342
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/54 (37%), Positives = 33/54 (61%), Gaps = 1/54 (1%)
Frame = +2
Query: 199 ENNYVFNYR-TVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPL 251
+N+ +F Y+ T +G ++++LF+ S +N F V+ DSTYK N+Y PL
Sbjct: 188 DNDPMFFYKFTKTGDESLENLFWCDGVSRMDYNVFGDVIAFDSTYKKNKYNKPL 349
>TC8639 similar to UP|FRI4_SOYBN (Q948P5) Ferritin 4, chloroplast precursor
(SFerH-4), partial (43%)
Length = 515
Score = 29.3 bits (64), Expect = 2.7
Identities = 22/62 (35%), Positives = 29/62 (46%), Gaps = 5/62 (8%)
Frame = +3
Query: 509 CGSDSKKCG--CVIRWTHALPCTCLIYK---KLNLGSCIRLDEINIHWKKLCFEVDDDVE 563
CG + +CG C+IR LPC I + GSC EI K+ C EVD E
Sbjct: 336 CG*RTNQCGVQCLIR----LPCDVCILR*GQCCAQGSC*LFSEIKCRGKRAC*EVDGISE 503
Query: 564 ES 565
++
Sbjct: 504 QT 509
>TC17367 similar to GB|BAB03049.1|9294683|AP001305 syringomycin biosynthesis
enzyme-like protein {Arabidopsis thaliana;}, partial
(17%)
Length = 341
Score = 28.9 bits (63), Expect = 3.5
Identities = 23/77 (29%), Positives = 32/77 (40%)
Frame = +3
Query: 603 PHTTSLTPPIQKLKTKGAQKGAQKRKRSTLDDSSTTREPSFWEHVDAQFPDSQASQTKPS 662
P L PP +L+ Q K T + SST PSF E + P + + ++PS
Sbjct: 120 PQYHLLRPP--QLRFLSPQHSQSKPTNPTSNHSSTNPAPSFSETSHSTRPTTSTTSSRPS 293
Query: 663 LQKKKSVCVVNPSSTPT 679
K + PS PT
Sbjct: 294 -GMKSFPTLAAPSRAPT 341
>TC18721 weakly similar to UP|BAD10141 (BAD10141) Zinc finger protein-like,
partial (16%)
Length = 378
Score = 28.5 bits (62), Expect = 4.6
Identities = 11/31 (35%), Positives = 16/31 (51%)
Frame = +3
Query: 807 CFPLRGRPPTNPSARIMCLGLIPNHFVHVFL 837
C R PP P AR++C+ + +H V L
Sbjct: 126 CHSARATPPCTPGARVVCIFYV*DHMASVRL 218
>AV771382
Length = 432
Score = 28.5 bits (62), Expect = 4.6
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +3
Query: 37 WARGQAIEAGFTLIIDKSYSGSGRRKPKLVL 67
W RG + + F+ I +KS S S + PKL L
Sbjct: 336 WPRGGGLASSFSFIYNKSSSVSKKNSPKLEL 428
>BP073959
Length = 478
Score = 28.1 bits (61), Expect = 6.0
Identities = 16/30 (53%), Positives = 20/30 (66%), Gaps = 3/30 (10%)
Frame = -1
Query: 680 PTQYIPCIDQMPNVMKP---YIEKIVNVDG 706
P + I C++Q NV KP YIE IV+VDG
Sbjct: 307 PLEKIKCVNQSQNVKKPSEKYIE-IVSVDG 221
>TC8047 similar to UP|Q9ASV9 (Q9ASV9) At1g53210/F12M16_12, partial (31%)
Length = 1144
Score = 28.1 bits (61), Expect = 6.0
Identities = 17/72 (23%), Positives = 33/72 (45%), Gaps = 7/72 (9%)
Frame = -2
Query: 524 HALPCTCLIYKKLNLGSCIRLDEINIHWKKLCF-------EVDDDVEESNRHVSIQNELN 576
H + C C IY + C+ L + ++ + CF + ++ EES S+ ++
Sbjct: 501 HIIHCHCTIYLRKC*RGCLSLITTSKYYCRHCFT*IGC*RKKNEGNEESRNSCSL*KVID 322
Query: 577 AIQERFNKADYN 588
I +R +K+ N
Sbjct: 321 CIDQRISKSSSN 286
>TC17149 similar to GB|AAP37818.1|30725592|BT008459 At1g43860 {Arabidopsis
thaliana;}, partial (48%)
Length = 597
Score = 28.1 bits (61), Expect = 6.0
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = -3
Query: 674 PSSTPTPTQYIPCIDQMPN 692
P STP P + C+DQ PN
Sbjct: 517 PESTPKPASWKCCVDQQPN 461
>BP061361
Length = 349
Score = 27.7 bits (60), Expect = 7.9
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -3
Query: 512 DSKKCGCVIRWTHALPCTCLIYKKLNLGSCIRLDEINIH 550
D +KCG L C L + L+L C++++E+ +H
Sbjct: 269 DHRKCG*TDTALGCLSCRELTLESLDLHLCLKVNEVPVH 153
>TC20271 weakly similar to UP|Q84XH9 (Q84XH9) RCa4 (Fragment), partial (10%)
Length = 692
Score = 27.7 bits (60), Expect = 7.9
Identities = 28/109 (25%), Positives = 46/109 (41%), Gaps = 7/109 (6%)
Frame = +3
Query: 550 HWKKLCFEVDDDVEESNRHVSIQNELN-----AIQERFNKADYNMKLIIKEQMREMTFPH 604
H+ +L + V SN + I NEL+ + R + + + E +RE H
Sbjct: 9 HFSELQRRLQISVSPSNSRLPIVNELDDKELLELASRLAGDEVSANSLAIEGLRE----H 176
Query: 605 TTSLTPPIQKLKTKGAQKGAQKR--KRSTLDDSSTTREPSFWEHVDAQF 651
+ KLK G +R R ++ S++ ++P F EHV QF
Sbjct: 177 WKKTESNMVKLKELGPLLRDYQRLIHRHHIERSASDQQPGFGEHVMHQF 323
>TC8595 similar to PIR|T09390|T09390 21K protein precursor - alfalfa
{Medicago sativa;}, partial (93%)
Length = 811
Score = 27.7 bits (60), Expect = 7.9
Identities = 21/72 (29%), Positives = 25/72 (34%)
Frame = +2
Query: 610 PPIQKLKTKGAQKGAQKRKRSTLDDSSTTREPSFWEHVDAQFPDSQASQTKPSLQKKKSV 669
PP +L T SS PS P ++ S KPS Q K+
Sbjct: 134 PPNTQLSASNPSPPTPPPSNKTHTSSSKLHSPS---------PSTEPSPPKPSSQDAKTS 286
Query: 670 CVVNPSSTPTPT 681
NP STP T
Sbjct: 287 GDSNPRSTPPST 322
>TC15745
Length = 517
Score = 27.7 bits (60), Expect = 7.9
Identities = 15/51 (29%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Frame = +1
Query: 338 LDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEELYADAVLQ-FRKVCE 387
L+C S G H K +V+ M WE +V + L+ + +LQ + ++C+
Sbjct: 28 LECFCLPSTGCHWKDECVVHRAMLVWEFVVECHTFWLFKEEMLQHYIQICK 180
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,671,381
Number of Sequences: 28460
Number of extensions: 206528
Number of successful extensions: 1156
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1143
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1151
length of query: 840
length of database: 4,897,600
effective HSP length: 98
effective length of query: 742
effective length of database: 2,108,520
effective search space: 1564521840
effective search space used: 1564521840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC138063.5


