

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.9 - phase: 0
(672 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
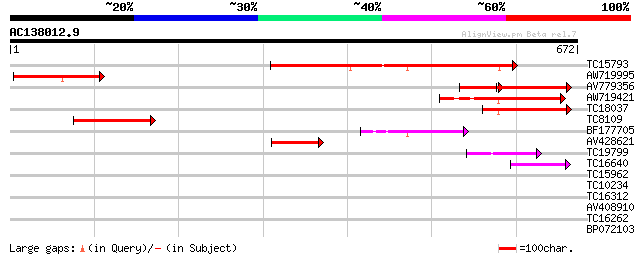
Score E
Sequences producing significant alignments: (bits) Value
TC15793 similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper protein (AT... 233 1e-61
AW719995 148 2e-36
AV779356 103 2e-31
AW719421 122 2e-28
TC18037 weakly similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper prot... 121 4e-28
TC8109 similar to UP|Q37899 (Q37899) Major tail protein gp25, pa... 87 1e-17
BF177705 80 1e-15
AV428621 66 2e-11
TC19799 weakly similar to PIR|PQ0476|PQ0476 pistil extensin-like... 54 8e-08
TC16640 similar to UP|Q9FK34 (Q9FK34) Gb|AAD25781.1, partial (13%) 52 2e-07
TC15962 similar to UP|Q9FHC6 (Q9FHC6) Emb|CAB83315.1, partial (13%) 38 0.006
TC10234 35 0.030
TC16312 similar to GB|AAM65771.1|21593804|AY088230 ribosomal pro... 29 2.8
AV408910 28 4.8
TC16262 similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase ho... 28 6.3
BP072103 28 6.3
>TC15793 similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper protein
(AT5g58430/mqj2_20), partial (43%)
Length = 969
Score = 233 bits (593), Expect = 1e-61
Identities = 129/306 (42%), Positives = 192/306 (62%), Gaps = 14/306 (4%)
Frame = +2
Query: 310 LDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVI 369
L+ +RWM AS +A +LFP E++ CD VF GFSSA F+E+C+ QLLNFAD +
Sbjct: 56 LEDEIERWMKASKVALMILFPSERRLCDRVFFGFSSAADLSFMEVCRGTAIQLLNFADAV 235
Query: 370 AYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLVNEAIAVRNRLGDASRVLFMK 425
A G+ S RLF+++D++ + +L+P+FESLF + SL NEAI + RLG+A R +FM+
Sbjct: 236 AIGTRSPERLFRVLDVYETMRDLIPEFESLFCDQYSVSLRNEAITIWKRLGEAIRGVFME 415
Query: 426 MHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQI-------LEEYPEVH 478
+ N I R PA +V G H +T VM+Y+ +ACR + LEQ+ L+EYP +
Sbjct: 416 LENLIRRDPAKAEVPG--GGLHPITRYVMNYLRAACRSLQTLEQVFEDYGHPLKEYPRLD 589
Query: 479 NEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETI 538
+ V +SS QM+ IM +L+ L KS+ KD AL ++F++NN +I + S L T+
Sbjct: 590 DRVPSSSSMSVQMDWIMELLESNLEAKSKIYKDPALCYVFLMNNGRYIVQKTRDSELGTL 769
Query: 539 FGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESIT---KELLKEKIHLFNNRFE 595
G+DW + + AK++Q LY+RS+W++V+ FLK+D + + +KEK+ FN FE
Sbjct: 770 LGDDWIRKHTAKVRQYHVLYQRSSWNKVLGFLKMDTGSMPSGGVAKSMKEKLRSFNILFE 949
Query: 596 AICRVQ 601
ICRVQ
Sbjct: 950 DICRVQ 967
>AW719995
Length = 364
Score = 148 bits (374), Expect = 2e-36
Identities = 74/111 (66%), Positives = 88/111 (78%), Gaps = 3/111 (2%)
Frame = +1
Query: 5 IIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFIM--- 61
+I+I L+Q VWRFV FASA VGLLC ALS+SFNHLFG+W KI LY++FS IM
Sbjct: 31 LIQILSWLMQPNVWRFVSFASAVVGLLCNALSSSFNHLFGDWTFFKISLYSVFSVIMSLM 210
Query: 62 ILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCA 112
+L+A W++SRSL+FKAH AFLVLTITS+YSF DK +NGKPD YSLISCA
Sbjct: 211 MLFAKKWRHSRSLQFKAHLAFLVLTITSLYSFISDKXMNGKPDVYSLISCA 363
>AV779356
Length = 518
Score = 103 bits (256), Expect(2) = 2e-31
Identities = 48/89 (53%), Positives = 66/89 (73%)
Frame = -3
Query: 578 ITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI 637
+ + L+KEK+ LFN E + RVQS W +Y ++LR EI +S+ N+LLPAYGIF+GR +
Sbjct: 357 LKQNLMKEKLDLFNFCLEQMSRVQSTWRVYDNKLREEIRTSLENVLLPAYGIFIGRFPDV 178
Query: 638 LGNQAYKYIKYGMIEIQDLLNHLFLGNKM 666
LG A+KYI+YGM +IQ+ LNHLF G +M
Sbjct: 177 LGKHAHKYIEYGMFDIQERLNHLFNGGQM 91
Score = 50.1 bits (118), Expect(2) = 2e-31
Identities = 20/52 (38%), Positives = 35/52 (66%)
Frame = -2
Query: 534 RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKE 585
++ I G DWF +++ + NL+LY RS+WD+V D LKL+++E + ++ E
Sbjct: 502 KMVVISGQDWFPSSRELVHPNLELYLRSSWDKVFDLLKLNHHELMVSDVEAE 347
>AW719421
Length = 575
Score = 122 bits (306), Expect = 2e-28
Identities = 68/153 (44%), Positives = 97/153 (62%), Gaps = 4/153 (2%)
Frame = +2
Query: 510 KDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDF 569
KD AL + FM+NN +I+ S+L T G+D FQ K QQN +LY RS+W V++F
Sbjct: 2 KDPALCYFFMMNNMRYID-----SQLXTTLGDDRFQK---KAQQNFELYCRSSWSNVLEF 157
Query: 570 LKLDNNES----ITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLP 625
LKLD +ES + + + EK++LF F+ +C VQS W + QLR +II SV N+LLP
Sbjct: 158 LKLDIDESWDSNVAAKSMIEKLNLFYQHFKEVCGVQSTWRVCDKQLRKQIIESVENMLLP 337
Query: 626 AYGIFVGRLHGILGNQAYKYIKYGMIEIQDLLN 658
AY F+ + +LG Q+ ++IKY M +IQ +LN
Sbjct: 338 AYRNFIEQFKDVLGRQSDEFIKYEMCDIQGMLN 436
>TC18037 weakly similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper protein
(AT5g58430/mqj2_20), partial (8%)
Length = 495
Score = 121 bits (303), Expect = 4e-28
Identities = 56/109 (51%), Positives = 81/109 (73%), Gaps = 4/109 (3%)
Frame = +1
Query: 561 SAWDEVMDFLKLDNNES----ITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEII 616
S+W++++D LK+++NES + +L+K+K++LFN FE ICRVQS W + QLR EI
Sbjct: 4 SSWNKMLDILKVESNESEAPNVAADLMKDKLNLFNMHFEEICRVQSTWSVIDEQLREEIR 183
Query: 617 SSVGNILLPAYGIFVGRLHGILGNQAYKYIKYGMIEIQDLLNHLFLGNK 665
SV +ILLPAYG F+GR +LG AY+YI+YG+ +I+D LN+LF+G K
Sbjct: 184 ISVNDILLPAYGNFIGRFQDVLGKHAYEYIEYGLFDIEDSLNYLFVGAK 330
>TC8109 similar to UP|Q37899 (Q37899) Major tail protein gp25, partial (5%)
Length = 842
Score = 86.7 bits (213), Expect = 1e-17
Identities = 42/97 (43%), Positives = 62/97 (63%)
Frame = -1
Query: 76 FKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMSMSLSRQSQCGIEVDLLY 135
F+++ F VL I SVYSF +DK VNGKP+ SL++ A F +MS+SLS+ + G E+ +
Sbjct: 782 FRSYVIFAVLIIISVYSFLYDKAVNGKPEVTSLVANAEFTLMSLSLSKLIKSGYEIGMFA 603
Query: 136 FFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSS 172
+FLGC +QL+ I L++V F L ++ SS S
Sbjct: 602 YFLGCFTIQLVTINWALILVAIVFGCPLFVMHSSSDS 492
>BF177705
Length = 412
Score = 80.1 bits (196), Expect = 1e-15
Identities = 52/135 (38%), Positives = 74/135 (54%), Gaps = 7/135 (5%)
Frame = +2
Query: 416 GDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQI----- 470
G A +FM++ N I +AK VS G H +T VM+Y+ +ACR R LE +
Sbjct: 14 GKAITWIFMELENLIR--DSAKTEVSDGGVH-PITCDVMNYLRAACRSRETLEHVFKEYG 184
Query: 471 --LEEYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEA 528
L+EYPE ++V +SS Q+ IM +L+ L KS+ KD AL +F++NN +I
Sbjct: 185 HPLKEYPEFDDKVPSSSSMPVQVNLIMELLESSLEAKSKIYKDPALGDVFLMNNVRYIVQ 364
Query: 529 MNKFSRLETIFGNDW 543
K L T+ GNDW
Sbjct: 365 EAKDGELGTLLGNDW 409
>AV428621
Length = 230
Score = 66.2 bits (160), Expect = 2e-11
Identities = 31/62 (50%), Positives = 41/62 (66%)
Frame = +2
Query: 311 DTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIA 370
+T F RW+ AS++A +LFP E+ CD + GFSSA F E C+ + QLLNFADV+A
Sbjct: 38 ETGFGRWVKASNVALKILFPNER*LCDRILLGFSSAADFSFTEDCRGSVIQLLNFADVVA 217
Query: 371 YG 372
G
Sbjct: 218 IG 223
>TC19799 weakly similar to PIR|PQ0476|PQ0476 pistil extensin-like protein
(clone pMG08) - common tobacco
(fragment) {Nicotiana tabacum;} , partial (12%)
Length = 601
Score = 53.9 bits (128), Expect = 8e-08
Identities = 29/89 (32%), Positives = 49/89 (54%)
Frame = +2
Query: 542 DWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQ 601
DW N+K K+++ + Y+R AW++V+ L +N + + + FN F CR Q
Sbjct: 2 DWLANHKLKVKEYVSKYERVAWNKVVAALP-ENPTAAGEVTARACFVAFNAAFR*ECRKQ 178
Query: 602 SAWFIYGSQLRGEIISSVGNILLPAYGIF 630
S+W + +LR EI S+G++L+ Y F
Sbjct: 179 SSWVVSDPKLREEIKGSIGSMLVLKYSDF 265
>TC16640 similar to UP|Q9FK34 (Q9FK34) Gb|AAD25781.1, partial (13%)
Length = 576
Score = 52.4 bits (124), Expect = 2e-07
Identities = 31/72 (43%), Positives = 42/72 (58%), Gaps = 1/72 (1%)
Frame = +1
Query: 594 FEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL-GNQAYKYIKYGMIE 652
FE I RVQ+AW + QLR E+ S+ ++PAY FVGR L G + KYIKY +
Sbjct: 7 FEEIYRVQTAWKVPDEQLREEMRISISEKVIPAYRSFVGRFKSQLEGRHSGKYIKYTPED 186
Query: 653 IQDLLNHLFLGN 664
++ L LF G+
Sbjct: 187 LETYLLDLFEGS 222
>TC15962 similar to UP|Q9FHC6 (Q9FHC6) Emb|CAB83315.1, partial (13%)
Length = 602
Score = 37.7 bits (86), Expect = 0.006
Identities = 19/49 (38%), Positives = 28/49 (56%)
Frame = +3
Query: 590 FNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL 638
FN FE + + QS W + S+LR + +V +LLPAY FV R ++
Sbjct: 12 FNVMFEELHQKQSQWTVPDSELRESLRLAVAEVLLPAYRSFVKRFGALV 158
>TC10234
Length = 902
Score = 35.4 bits (80), Expect = 0.030
Identities = 26/106 (24%), Positives = 47/106 (43%), Gaps = 6/106 (5%)
Frame = +1
Query: 230 DRLMEE---YNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQK 286
D + EE + G E + + + +L I + M+ GY KEC VY RK ++ +
Sbjct: 580 DEISEEEFTFTAGNSISETERVSMLAMEDLKAIAESMIACGYGKECVKVYTIMRKSIVDE 759
Query: 287 GLLNKIFVLPEAKINTERERE---RYLDTMFQRWMTASDIATTVLF 329
L + L ++N + ++ L+ + W+ A +A LF
Sbjct: 760 AL----YHLGVERLNLPQVQKMDWEVLELKIRSWLKAVKVAVGTLF 885
>TC16312 similar to GB|AAM65771.1|21593804|AY088230 ribosomal protein S1
{Arabidopsis thaliana;}, partial (5%)
Length = 691
Score = 28.9 bits (63), Expect = 2.8
Identities = 18/49 (36%), Positives = 27/49 (54%)
Frame = +2
Query: 124 QSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSS 172
QSQ I LY FL CL++ +M I+ A FS+S + ++ +SS
Sbjct: 230 QSQTSISKTTLYLFLICLLIAVMAIR-------ATFSFSGYVAQNLVSS 355
>AV408910
Length = 417
Score = 28.1 bits (61), Expect = 4.8
Identities = 14/49 (28%), Positives = 24/49 (48%)
Frame = +2
Query: 52 FLYTLFSFIMILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVN 100
FL+ + S L N+W+ S+ LRF L LT+ + F +++
Sbjct: 170 FLFLIHSQYQYLQHNLWRLSKRLRFPIKHRPLTLTLLGLLFSFLSHLIS 316
>TC16262 similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase homolog
COK-4, partial (6%)
Length = 623
Score = 27.7 bits (60), Expect = 6.3
Identities = 22/72 (30%), Positives = 36/72 (49%), Gaps = 2/72 (2%)
Frame = +1
Query: 585 EKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISS--VGNILLPAYGIFVGRLHGILGNQA 642
E I + NN E+ R+ ++I S+LR E I VGNI Y +++ + L ++
Sbjct: 34 EGIEMCNNGSESSDRITMHFYIM-SRLRTEHIDPALVGNIAPECYAVYIDIIDRCLKHEP 210
Query: 643 YKYIKYGMIEIQ 654
+ G +EIQ
Sbjct: 211 DERPTMGEVEIQ 246
>BP072103
Length = 455
Score = 27.7 bits (60), Expect = 6.3
Identities = 13/45 (28%), Positives = 24/45 (52%), Gaps = 1/45 (2%)
Frame = -2
Query: 424 MKMHNFI-FRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKL 467
+K H + F++ A Q+V G HHQ S +S+ C ++++
Sbjct: 289 LKPHYLLCFQMAALHQLV*EVGHHHQQDCFFTSVISTLCSSKKQM 155
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,484,000
Number of Sequences: 28460
Number of extensions: 157323
Number of successful extensions: 1077
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1060
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1067
length of query: 672
length of database: 4,897,600
effective HSP length: 96
effective length of query: 576
effective length of database: 2,165,440
effective search space: 1247293440
effective search space used: 1247293440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC138012.9


