

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137994.2 + phase: 0 /pseudo
(379 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
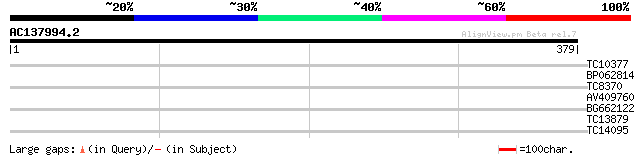
Score E
Sequences producing significant alignments: (bits) Value
TC10377 similar to UP|THIC_SALTI (Q8Z326) Thiamine biosynthesis ... 30 0.51
BP062814 29 1.5
TC8370 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger-... 28 2.5
AV409760 26 9.6
BG662122 26 9.6
TC13879 similar to GB|AAL49941.1|17979249|AY070475 AT3g04610/F7O... 26 9.6
TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein... 26 9.6
>TC10377 similar to UP|THIC_SALTI (Q8Z326) Thiamine biosynthesis protein
thiC, partial (40%)
Length = 756
Score = 30.4 bits (67), Expect = 0.51
Identities = 35/105 (33%), Positives = 48/105 (45%), Gaps = 12/105 (11%)
Frame = -2
Query: 267 CCSKSRLSTPISTIS-------TTG*TQIDLIP-MKYADLFPILLERNL-VHTKAPLPVP 317
C S+LS I I T G*+ I LI +KY + P+L+ + L V + AP +
Sbjct: 470 CQELSKLSCIICIIDGSRS*PITNG*SHIILIADVKY--VIPVLICKILFVIS*APFCMN 297
Query: 318 ARLSARYRPNLFCA---FHQGASCHDIEHCFSLKKIVQNLIPENL 359
S Y + FC Q SC D E ++L + IPENL
Sbjct: 296 GSSSGDYSCHAFCC*RNISQKNSCMDSEVIYTLFSLFNQGIPENL 162
>BP062814
Length = 509
Score = 28.9 bits (63), Expect = 1.5
Identities = 19/59 (32%), Positives = 33/59 (55%)
Frame = +3
Query: 270 KSRLSTPISTISTTG*TQIDLIPMKYADLFPILLERNLVHTKAPLPVPARLSARYRPNL 328
K +STP T+S+T ++ + +++A IL + ++ K P+P +L A YRP L
Sbjct: 3 KKNISTPS*TVSSTD-QKVPFLRLQWA--MDILTKIVILLKKIPVP*DEQLDALYRPTL 170
>TC8370 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger-like
protein, partial (20%)
Length = 776
Score = 28.1 bits (61), Expect = 2.5
Identities = 15/34 (44%), Positives = 23/34 (67%), Gaps = 1/34 (2%)
Frame = -3
Query: 16 SVKSYVRVDNLQEKYNEIQREMKA-LHRKELFGI 48
S K+ R +NL++KYNE+ R +K L+R+ GI
Sbjct: 564 SGKTKSRSENLEKKYNELLRSLKQ*LNRRRAGGI 463
>AV409760
Length = 424
Score = 26.2 bits (56), Expect = 9.6
Identities = 12/38 (31%), Positives = 18/38 (46%)
Frame = -2
Query: 296 ADLFPILLERNLVHTKAPLPVPARLSARYRPNLFCAFH 333
+ LFP+ ER + + L P L +L+C FH
Sbjct: 333 SSLFPLYQERKIDQPEMNLTDPTTLETNLVHSLYCNFH 220
>BG662122
Length = 386
Score = 26.2 bits (56), Expect = 9.6
Identities = 9/17 (52%), Positives = 14/17 (81%)
Frame = +2
Query: 339 HDIEHCFSLKKIVQNLI 355
HDI+ CF+LK+ ++ LI
Sbjct: 236 HDIDDCFTLKREIEKLI 286
>TC13879 similar to GB|AAL49941.1|17979249|AY070475 AT3g04610/F7O18_9
{Arabidopsis thaliana;}, partial (12%)
Length = 684
Score = 26.2 bits (56), Expect = 9.6
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -1
Query: 130 RLSSSNTTTTRTWPQIE 146
R S +TTT+RTWP+ E
Sbjct: 54 RWKSRSTTTSRTWPRAE 4
>TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein S2,
partial (88%)
Length = 1134
Score = 26.2 bits (56), Expect = 9.6
Identities = 29/104 (27%), Positives = 45/104 (42%), Gaps = 21/104 (20%)
Frame = +1
Query: 246 KSTSTTVSGLSPCCRCYA*RQCCSKSRL-------STPISTISTTG*TQID-------LI 291
+ST ++ L P R +*R C S+SR S P S+ +TT T L+
Sbjct: 274 RSTRSSTCWLDPLSRTRS*RSCRSRSRPVPVSEPGSRPSSSSATTMVTSGSA*SAARKLL 453
Query: 292 P-------MKYADLFPILLERNLVHTKAPLPVPARLSARYRPNL 328
P + + L P + + +++P P PARL P+L
Sbjct: 454 PPFAVGLSLLSSLLSPSGGDTGVTRSESPTPFPARLPESVVPSL 585
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.147 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,711,040
Number of Sequences: 28460
Number of extensions: 120539
Number of successful extensions: 888
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 884
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 888
length of query: 379
length of database: 4,897,600
effective HSP length: 92
effective length of query: 287
effective length of database: 2,279,280
effective search space: 654153360
effective search space used: 654153360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC137994.2


