

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137826.3 - phase: 0 /pseudo
(1546 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
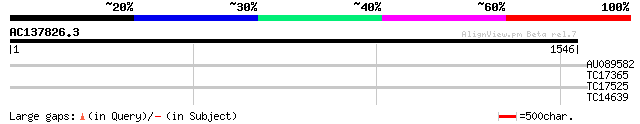
AU089582 33 0.004
TC17365 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (19%) 31 1.7
TC17525 similar to UP|O82624 (O82624) T9A4.14 protein, partial (... 29 5.0
TC14639 weakly similar to GB|AAM91068.1|22136564|AY129482 AT5g11... 29 5.0
>AU089582
Length = 383
Score = 33.1 bits (74), Expect(2) = 0.004
Identities = 12/21 (57%), Positives = 17/21 (80%)
Frame = +3
Query: 1359 GHWDKFLTLCEFSYNNNYTLS 1379
G WD++L+L EF+YNN+Y S
Sbjct: 249 GSWDQYLSLMEFAYNNSYRSS 311
Score = 25.4 bits (54), Expect(2) = 0.004
Identities = 9/22 (40%), Positives = 15/22 (67%)
Frame = +1
Query: 1332 SSNGWTIRENNPSVRGHIEGMC 1353
SSN W++RE+ +RG+ +C
Sbjct: 169 SSN*WSVREDYSDLRGYASCLC 234
>TC17365 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (19%)
Length = 364
Score = 30.8 bits (68), Expect = 1.7
Identities = 16/35 (45%), Positives = 18/35 (50%)
Frame = -2
Query: 435 PASNDKNSRGHPRGW*GGNHRGCEGRENGNVGRGA 469
PA + RG RGW G+HRG G V RGA
Sbjct: 210 PARKGRGGRGGARGWEVGSHRG----GGGGVARGA 118
>TC17525 similar to UP|O82624 (O82624) T9A4.14 protein, partial (22%)
Length = 586
Score = 29.3 bits (64), Expect = 5.0
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = -3
Query: 553 THVYQLYLVIFMGFQSWVI*LCWI*LL 579
THV+ +Y+ + + SWV +CW LL
Sbjct: 575 THVFVIYIYVTTQY*SWVYIICWYNLL 495
>TC14639 weakly similar to GB|AAM91068.1|22136564|AY129482
AT5g11740/T22P22_130 {Arabidopsis thaliana;}, partial
(61%)
Length = 507
Score = 29.3 bits (64), Expect = 5.0
Identities = 17/52 (32%), Positives = 21/52 (39%)
Frame = -1
Query: 430 TRAVVPASNDKNSRGHPRGW*GGNHRGCEGRENGNVGRGALQPRKKVVCQDG 481
T + PAS+D R GG G EGR + RG + R DG
Sbjct: 276 TEIIHPASSDPERRSKGESSDGGKETGGEGRRDSGRRRGGARSRSLSGSSDG 121
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.359 0.161 0.583
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,594,176
Number of Sequences: 28460
Number of extensions: 426347
Number of successful extensions: 3945
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2869
Number of HSP's successfully gapped in prelim test: 97
Number of HSP's that attempted gapping in prelim test: 1002
Number of HSP's gapped (non-prelim): 3081
length of query: 1546
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1444
effective length of database: 1,994,680
effective search space: 2880317920
effective search space used: 2880317920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137826.3


