

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
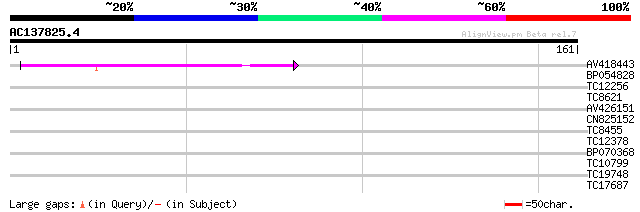
Query= AC137825.4 - phase: 0
(161 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV418443 54 1e-08
BP054828 26 2.9
TC12256 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_12... 26 3.8
TC8621 similar to UP|RS8_PRUAR (O81361) 40S ribosomal protein S8... 26 3.8
AV426151 26 3.8
CN825152 25 4.9
TC8455 homologue to UP|Q9FUB9 (Q9FUB9) Beta-carotene hydroxylase... 25 4.9
TC12378 25 8.4
BP070368 25 8.4
TC10799 weakly similar to UP|O23678 (O23678) CER1-like protein, ... 25 8.4
TC19748 weakly similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoen... 25 8.4
TC17687 weakly similar to GB|AAH05628.1|13542865|BC005628 acidic... 25 8.4
>AV418443
Length = 351
Score = 53.9 bits (128), Expect = 1e-08
Identities = 27/81 (33%), Positives = 48/81 (58%), Gaps = 2/81 (2%)
Frame = +1
Query: 4 LLGVLVMIVLLGNSIYAYGA--INQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSF 61
+L + ++ + N+ YA+ + + R+I+ N +GP +G+++PN +EL+PLL +S +
Sbjct: 115 ILSMFLVAFMWINAQYAFVSCTLTPQVQRKIARANQKGPYLGLIIPNTFELDPLLQNSDY 294
Query: 62 VPHNKFPYFDFAGRHFRIGEL 82
P DFAGR FR G +
Sbjct: 295 TPSKL--TIDFAGRRFRFGAI 351
>BP054828
Length = 476
Score = 26.2 bits (56), Expect = 2.9
Identities = 17/40 (42%), Positives = 21/40 (52%), Gaps = 3/40 (7%)
Frame = +1
Query: 34 NINNEGPSIGIVVPNAYELN---PLLHSSSFVPHNKFPYF 70
NIN S P+A L+ PLL ++SF PH F YF
Sbjct: 292 NINRWAGSPRRRFPSAISLSDRSPLLPAASFSPHPGFGYF 411
>TC12256 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_120, partial
(33%)
Length = 640
Score = 25.8 bits (55), Expect = 3.8
Identities = 11/36 (30%), Positives = 21/36 (57%)
Frame = +2
Query: 31 EISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNK 66
++ + ++ P I I++ L P+L+ SF+P NK
Sbjct: 302 DLRSEDSSHPLILIIIFRGQHLQPILNQLSFLPSNK 409
>TC8621 similar to UP|RS8_PRUAR (O81361) 40S ribosomal protein S8, partial
(98%)
Length = 857
Score = 25.8 bits (55), Expect = 3.8
Identities = 10/30 (33%), Positives = 18/30 (59%)
Frame = -2
Query: 62 VPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
+PH KFP + + HF + L + +IV++
Sbjct: 265 LPHEKFPVSNLSALHFTLPPLTRILLIVLL 176
>AV426151
Length = 331
Score = 25.8 bits (55), Expect = 3.8
Identities = 13/44 (29%), Positives = 21/44 (47%)
Frame = +2
Query: 27 LSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYF 70
L W S + +E + V + + P LH S + PH +F +F
Sbjct: 134 LDWA*RSKVGSESNA*SDRVSRSTAIPPSLHCSLYFPHFRFLWF 265
>CN825152
Length = 697
Score = 25.4 bits (54), Expect = 4.9
Identities = 9/17 (52%), Positives = 10/17 (57%), Gaps = 3/17 (17%)
Frame = +2
Query: 139 WAHTGLWH---WQVHTS 152
WAH LWH W +H S
Sbjct: 536 WAHKALWHASLWHMHES 586
>TC8455 homologue to UP|Q9FUB9 (Q9FUB9) Beta-carotene hydroxylase, partial
(49%)
Length = 566
Score = 25.4 bits (54), Expect = 4.9
Identities = 9/17 (52%), Positives = 10/17 (57%), Gaps = 3/17 (17%)
Frame = +3
Query: 139 WAHTGLWH---WQVHTS 152
WAH LWH W +H S
Sbjct: 27 WAHKALWHASLWHMHES 77
>TC12378
Length = 464
Score = 24.6 bits (52), Expect = 8.4
Identities = 8/14 (57%), Positives = 9/14 (64%)
Frame = +3
Query: 139 WAHTGLWHWQVHTS 152
WA T W W VHT+
Sbjct: 147 WARTLQWDWAVHTT 188
>BP070368
Length = 300
Score = 24.6 bits (52), Expect = 8.4
Identities = 11/35 (31%), Positives = 18/35 (51%), Gaps = 2/35 (5%)
Frame = +1
Query: 42 IGIVVPNAYELNPLLHSSSFVPHNK--FPYFDFAG 74
+GI PN Y + ++ + HNK F D++G
Sbjct: 34 VGIYTPNMYNYSHIIDLKALKTHNKRQFTTHDYSG 138
>TC10799 weakly similar to UP|O23678 (O23678) CER1-like protein, partial
(13%)
Length = 605
Score = 24.6 bits (52), Expect = 8.4
Identities = 27/104 (25%), Positives = 44/104 (41%), Gaps = 5/104 (4%)
Frame = +2
Query: 48 NAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGE---LEKKKVIVVMSGEGMLNAGLATQ 104
N E H S F+P ++FP F + + ++ V S E L +
Sbjct: 17 NEAERQKAAHGSLFIPFSRFPPKKLRKDCFYLSTPAMVTPPSLVNVHSCENWL-----PR 181
Query: 105 LLLTLFNIEGVLHYGIAGNLNSRFQIGDV--TIPKYWAHTGLWH 146
+++ + I G+LH N+N + GDV +I K W L+H
Sbjct: 182 RVMSAWRIAGILHALEDWNIN---ECGDVMFSIEKVW-QASLYH 301
>TC19748 weakly similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyruvate
translocator protein-like, partial (22%)
Length = 427
Score = 24.6 bits (52), Expect = 8.4
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +3
Query: 8 LVMIVLLGNSIYAYGAINQLSWREISNINNE 38
L M+ ++ S Y+YGAIN L + +I+++
Sbjct: 309 LTMLHMISCSAYSYGAINFLDLVPLQHIHSK 401
>TC17687 weakly similar to GB|AAH05628.1|13542865|BC005628 acidic nuclear
phosphoprotein 32 family, member B {Mus musculus;},
partial (9%)
Length = 702
Score = 24.6 bits (52), Expect = 8.4
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = -1
Query: 52 LNPLLHSSSFVPHNKFP 68
LNP+LH SFV N P
Sbjct: 318 LNPILHDRSFVSQNSPP 268
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.142 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,207,973
Number of Sequences: 28460
Number of extensions: 44217
Number of successful extensions: 348
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 346
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 347
length of query: 161
length of database: 4,897,600
effective HSP length: 83
effective length of query: 78
effective length of database: 2,535,420
effective search space: 197762760
effective search space used: 197762760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 51 (24.3 bits)
Medicago: description of AC137825.4


