

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137823.16 + phase: 0
(621 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
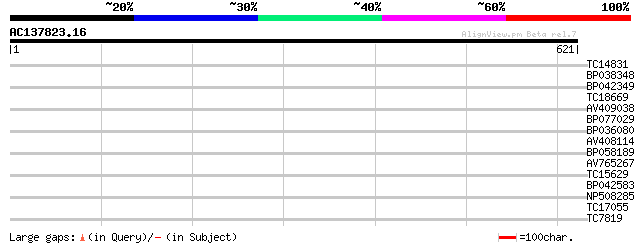
Score E
Sequences producing significant alignments: (bits) Value
TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragm... 31 0.67
BP038348 31 0.67
BP042349 30 1.5
TC18669 similar to UP|Q93Y39 (Q93Y39) ATP-dependent RNA helicase... 29 2.6
AV409038 28 3.3
BP077029 28 3.3
BP036080 28 4.4
AV408114 28 4.4
BP058189 27 7.5
AV765267 27 7.5
TC15629 similar to UP|Q40540 (Q40540) Pit2 protein, partial (21%) 27 9.7
BP042583 27 9.7
NP508285 myc-like regulatory protein [Lotus japonicus] 27 9.7
TC17055 similar to UP|Q9FG04 (Q9FG04) Dbj|BAA86974.1 (AT5g06660/... 27 9.7
TC7819 similar to UP|Q43817 (Q43817) Lipoxygenase , partial (97%) 27 9.7
>TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragment),
partial (63%)
Length = 851
Score = 30.8 bits (68), Expect = 0.67
Identities = 17/56 (30%), Positives = 26/56 (46%)
Frame = +3
Query: 345 TALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYR 400
TA + DA S W++QP + Y +I CS++ E +E+ A YR
Sbjct: 153 TATATSTADARSTWTEQPRPSSTY----LISPCSSLYEILGIPAGASSQEIKAAYR 308
>BP038348
Length = 526
Score = 30.8 bits (68), Expect = 0.67
Identities = 24/90 (26%), Positives = 36/90 (39%), Gaps = 6/90 (6%)
Frame = -3
Query: 289 EREVSGLKASLNTLMSEIQRLNKLC-AERKEAEDSLKK-----KWKKIEEFDGRRSELES 342
+RE++ + L ++QRL C A + + E ++K KWKK G + LE
Sbjct: 422 KREINAKHVEIRKLKEDVQRLQGQCSAMQVQMEKMVEKKKGFFKWKKFPFSXGESAVLE- 246
Query: 343 IYTALLKANTDAASFWSQQPSTAREYALST 372
K D F Q P A + T
Sbjct: 245 ------KTEQDGVEFERQTPGAATNVDMKT 174
>BP042349
Length = 402
Score = 29.6 bits (65), Expect = 1.5
Identities = 17/43 (39%), Positives = 23/43 (52%)
Frame = -2
Query: 384 SNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSS 426
+N KDL+ + +SPD LY PS QA+ +GS SS
Sbjct: 392 TNQKKDLVHCHIVLVQKSPDRRLY-FPSLQQAIPFIMGSETSS 267
>TC18669 similar to UP|Q93Y39 (Q93Y39) ATP-dependent RNA helicase
(Fragment), partial (3%)
Length = 461
Score = 28.9 bits (63), Expect = 2.6
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = -2
Query: 32 DAQRSGVDSSVEMVNSFSAKNEKEAVYSTVKGSKSSDDVIVIETTRE 78
D + G D + + V K K+ + V G++SSDDV +ET E
Sbjct: 202 DIHKPGGDDNKKQVVPKQGKKLKKQKHDGVDGARSSDDVPEVETDGE 62
>AV409038
Length = 349
Score = 28.5 bits (62), Expect = 3.3
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +2
Query: 38 VDSSVEMVNSFSAKNEKEAVYSTVKGSKSSDDVIVIE 74
+ +SVE V SA +EK+ V + S +S+D+I IE
Sbjct: 65 ISNSVEPVGQNSADSEKQVVANGKDSSLASNDIIEIE 175
>BP077029
Length = 486
Score = 28.5 bits (62), Expect = 3.3
Identities = 18/72 (25%), Positives = 35/72 (48%), Gaps = 5/72 (6%)
Frame = -3
Query: 268 IGATSQNVGSLRQLQLDVWAKEREV----SGLKASLNTLMSEIQRLNKLCAERKEAEDSL 323
I A N+ + ++ ++REV LKA + L ++ L K+ +D++
Sbjct: 457 INAIKANLSVFQTAKVTAIKRKREVFEEGKALKAQRDELREQVPHLRDEREVAKKIQDNI 278
Query: 324 KKKWKKI-EEFD 334
+ +W K+ E+FD
Sbjct: 277 RAEWSKLGEKFD 242
>BP036080
Length = 409
Score = 28.1 bits (61), Expect = 4.4
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = -1
Query: 255 LHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGL 295
L+GG D+T+ G T +N SLR + VW + + +GL
Sbjct: 409 LNGGADITN-----GLTHRNHRSLRLQRQQVWIRSKLTTGL 302
>AV408114
Length = 424
Score = 28.1 bits (61), Expect = 4.4
Identities = 19/42 (45%), Positives = 23/42 (54%)
Frame = -3
Query: 323 LKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPST 364
L KK K IEE RS+L LK+ TD ASF + P+T
Sbjct: 284 LPKKCKAIEELI*LRSKL----WIALKSTTDVASFTTPSPNT 171
>BP058189
Length = 537
Score = 27.3 bits (59), Expect = 7.5
Identities = 13/35 (37%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Frame = +3
Query: 180 YENNIVMDVSSSDGSSPLQYP-LYGNGKLGADVPP 213
++ + MDVS+ +GSSP +P ++ N +L + V P
Sbjct: 159 FKRKLPMDVSTMNGSSPAPFPIMHPNLRLKSHVEP 263
>AV765267
Length = 493
Score = 27.3 bits (59), Expect = 7.5
Identities = 23/91 (25%), Positives = 40/91 (43%)
Frame = +1
Query: 49 SAKNEKEAVYSTVKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGV 108
S + E E +Y+T + + +++ +E E +RK+C+ A + + +F S
Sbjct: 160 SCEQETEWIYNTCRVMELTNNKRRLELEEEAVLRKSCKLSDAAEEQETRKNFLELSLSLN 339
Query: 109 LSNPQAEAAKLGFDFDGQIPDEVRTVIVNCL 139
SN Q A D I R + +NCL
Sbjct: 340 QSNDQNRAGDQS-DSSSLIHQLGRDITINCL 429
>TC15629 similar to UP|Q40540 (Q40540) Pit2 protein, partial (21%)
Length = 616
Score = 26.9 bits (58), Expect = 9.7
Identities = 18/64 (28%), Positives = 29/64 (45%), Gaps = 5/64 (7%)
Frame = +3
Query: 87 SLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLGFDFDGQIPDEVRTVI-----VNCLKS 141
S + + +S+P S VL + +A + +DF+ +I D VI V LK
Sbjct: 210 SHIVFTKASADNSYPIRTLSAVLGSEEAAKEAVVYDFEDEINDNFSAVITPENAVQLLKQ 389
Query: 142 PPLL 145
P +L
Sbjct: 390 PGVL 401
>BP042583
Length = 459
Score = 26.9 bits (58), Expect = 9.7
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = -1
Query: 551 MEKWLPELKTGVLNAQQSLEACKYVWGLLDEW 582
+E+WL L + +L E C Y W LL W
Sbjct: 393 LEQWLVLLGSCMLKMPLD*EMCLYFWVLLTSW 298
>NP508285 myc-like regulatory protein [Lotus japonicus]
Length = 1529
Score = 26.9 bits (58), Expect = 9.7
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = -2
Query: 77 REKNIRKACESLVAYMVDKIQSSFP 101
REKN+ AC +L ++ V + S+FP
Sbjct: 310 REKNLLSAC*ALHSHTVSPVSSAFP 236
>TC17055 similar to UP|Q9FG04 (Q9FG04) Dbj|BAA86974.1 (AT5g06660/F15M7_19),
partial (26%)
Length = 458
Score = 26.9 bits (58), Expect = 9.7
Identities = 24/68 (35%), Positives = 31/68 (45%), Gaps = 7/68 (10%)
Frame = -3
Query: 453 IPSICRVSAALQ----YAAGGLEGSDAGLASILESLEFCLK---RRGSEASVLEDLLKAI 505
IP+ R S L+ YA G +E S S LES CL + GS S L A+
Sbjct: 339 IPTTVRPSEYLKSEKGYAMGSIEKSTRKRVSKLESGSNCLNCRAKNGSFPSSLRFKYNAL 160
Query: 506 NLVHIRRD 513
N+ I R+
Sbjct: 159 NINQINRN 136
>TC7819 similar to UP|Q43817 (Q43817) Lipoxygenase , partial (97%)
Length = 2842
Score = 26.9 bits (58), Expect = 9.7
Identities = 26/84 (30%), Positives = 37/84 (43%)
Frame = +2
Query: 436 ISAAILTARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASILESLEFCLKRRGSEA 495
+ A +L DPSA I + YAA GLE DA + + E + K S+A
Sbjct: 1895 LPADLLKRGVAVEDPSAPHGIRLLIEDYPYAADGLEIWDAIKSWVGEYVNLYYK---SDA 2065
Query: 496 SVLEDLLKAINLVHIRRDLVQSGH 519
+++D L +LVQ GH
Sbjct: 2066 EIVQD----AELQAFWTELVQVGH 2125
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,085,722
Number of Sequences: 28460
Number of extensions: 106565
Number of successful extensions: 416
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 416
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 416
length of query: 621
length of database: 4,897,600
effective HSP length: 96
effective length of query: 525
effective length of database: 2,165,440
effective search space: 1136856000
effective search space used: 1136856000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC137823.16


