

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.15 + phase: 0
(279 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
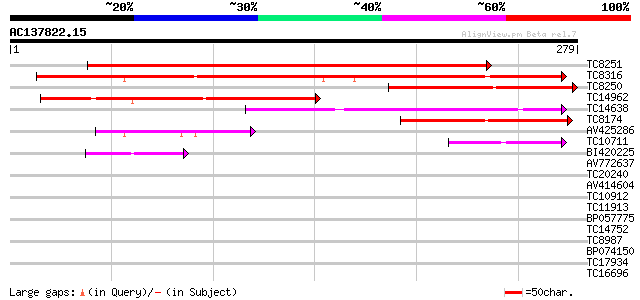
TC8251 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Ara... 365 e-102
TC8316 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Ara... 237 2e-63
TC8250 weakly similar to GB|AAO24559.1|27808558|BT003127 At5g398... 170 2e-43
TC14962 similar to GB|AAP37696.1|30725348|BT008337 At5g15120 {Ar... 137 2e-33
TC14638 similar to UP|Q9SJI9 (Q9SJI9) Expressed protein, partial... 134 2e-32
TC8174 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Ara... 81 2e-16
AV425286 53 5e-08
TC10711 50 4e-07
BI420225 39 7e-04
AV772637 35 0.018
TC20240 weakly similar to UP|Q9SJI9 (Q9SJI9) Expressed protein, ... 32 0.12
AV414604 27 2.9
TC10912 27 3.9
TC11913 weakly similar to UP|ABPA_PRUPE (Q9ZRA4) Auxin-binding p... 27 3.9
BP057775 27 5.0
TC14752 similar to UP|Q9LDV4 (Q9LDV4) Alanine aminotransferase ... 27 5.0
TC8987 similar to UP|ABPA_PRUPE (Q9ZRA4) Auxin-binding protein A... 27 5.0
BP074150 27 5.0
TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase ... 27 5.0
TC16696 similar to UP|Q9AR81 (Q9AR81) Germin-like protein precur... 27 5.0
>TC8251 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;}, partial (41%)
Length = 909
Score = 365 bits (936), Expect = e-102
Identities = 169/199 (84%), Positives = 186/199 (92%)
Frame = +3
Query: 39 VPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKE 98
VPKAL++LFDSC++TFKGP TVPSP+DV K+CHILD+++P D GLSRDLQFFKPGNIIKE
Sbjct: 9 VPKALKDLFDSCRETFKGPDTVPSPQDVQKMCHILDDIRPGDFGLSRDLQFFKPGNIIKE 188
Query: 99 NQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEAS 158
NQRVTYTTVYKC NFSLCIFF+PE+GVIPLHNHPGMTVFSKLLLGQMHIKSYDWV EAS
Sbjct: 189 NQRVTYTTVYKCANFSLCIFFIPEKGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVVPEAS 368
Query: 159 HNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKE 218
NLL ++KLRLAKLKA+ FTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKE
Sbjct: 369 DNLLDQTTKLRLAKLKADNVFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKE 548
Query: 219 DGRDCSYYKDYPYNAFPNE 237
DGRDCSYY+D+PY AFP+E
Sbjct: 549 DGRDCSYYRDHPYAAFPSE 605
>TC8316 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;}, partial (63%)
Length = 1796
Score = 237 bits (604), Expect = 2e-63
Identities = 126/269 (46%), Positives = 170/269 (62%), Gaps = 8/269 (2%)
Frame = +2
Query: 14 KVCYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFK--GPGTVPSPRDVHKLCH 71
++C + +K +RR ++ +Q LF +CKQ F GPG VPSP+ + L
Sbjct: 272 ELCELSAETKSTNSKSRKNRRHRQRNVLPVQRLFQTCKQVFASTGPGVVPSPQHIEMLHS 451
Query: 72 ILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNH 131
IL +KPEDVGL D+ +F N ++ ++TY +Y+C+NFS+ IF LP GVIPLHNH
Sbjct: 452 ILAGIKPEDVGLKPDMPYFM-NNSARKMPKITYLHIYECENFSMGIFCLPPGGVIPLHNH 628
Query: 132 PGMTVFSKLLLGQMHIKSYDWV-DHEASHNLLQPSSKL-----RLAKLKANKTFTAPCDT 185
P MTVFSKLL G MHIKSYDWV D + + SS++ RLAK+K + FTAPCD
Sbjct: 629 PEMTVFSKLLFGTMHIKSYDWVVDLPSEMPTILKSSEIQTQEKRLAKVKVDADFTAPCDP 808
Query: 186 SVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKD 245
S+LYPT GGN+H FTA+T CAVL V+GPPY+ DGR CSYY ++P++ F + I ++
Sbjct: 809 SILYPTDGGNMHCFTAVTACAVLYVLGPPYNDPDGRHCSYYHNFPFSNF--SDGISMPEE 982
Query: 246 KDDSYGLLEEIDMPENCQMDGIEYLGPPI 274
+ Y L+E + PE Q+ G Y GP I
Sbjct: 983 ERTGYEWLQEREKPEKFQVAGKMYSGPKI 1069
>TC8250 weakly similar to GB|AAO24559.1|27808558|BT003127 At5g39890
{Arabidopsis thaliana;}, partial (16%)
Length = 661
Score = 170 bits (431), Expect = 2e-43
Identities = 80/93 (86%), Positives = 85/93 (91%)
Frame = +3
Query: 187 VLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDK 246
VLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYY+D+PY AFPNEE I EVKDK
Sbjct: 3 VLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYRDHPYAAFPNEE-ISEVKDK 179
Query: 247 DDSYGLLEEIDMPENCQMDGIEYLGPPIDDTMF 279
+DSY LEEI+MPEN QMDGIEYLGPPI +T F
Sbjct: 180 NDSYAWLEEIEMPENSQMDGIEYLGPPITETAF 278
>TC14962 similar to GB|AAP37696.1|30725348|BT008337 At5g15120 {Arabidopsis
thaliana;}, partial (36%)
Length = 632
Score = 137 bits (345), Expect = 2e-33
Identities = 69/140 (49%), Positives = 91/140 (64%), Gaps = 2/140 (1%)
Frame = +1
Query: 16 CYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGT--VPSPRDVHKLCHIL 73
C + R N RR KK P +Q+LFD+CK+ F GT +P P+D+ +L +L
Sbjct: 190 CELPRETITSSGSKRNRRRQKKMPP--VQKLFDTCKEVFASGGTGFIPPPQDIQRLQAVL 363
Query: 74 DNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPG 133
D ++PEDVGL D+ F+ + + ++TY +Y+C+ FS+ IF LP GVIPLHNHPG
Sbjct: 364 DAIRPEDVGLKPDMPHFRSSSA-QRIPKITYLHIYECEKFSMGIFCLPPSGVIPLHNHPG 540
Query: 134 MTVFSKLLLGQMHIKSYDWV 153
MTVFSKLL G MHIKSYDWV
Sbjct: 541 MTVFSKLLFGTMHIKSYDWV 600
>TC14638 similar to UP|Q9SJI9 (Q9SJI9) Expressed protein, partial (61%)
Length = 826
Score = 134 bits (336), Expect = 2e-32
Identities = 70/158 (44%), Positives = 95/158 (59%)
Frame = +3
Query: 117 IFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKAN 176
IF +P +IPLHNHPGMTV SKLL G ++ KSYDW+D S + S+ R AKL +
Sbjct: 3 IFCMPPSSIIPLHNHPGMTVLSKLLYGSLYXKSYDWIDVPGSAD----PSQARPAKLVKD 170
Query: 177 KTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPN 236
TAP T+VLYP++GGNIH F AITPCA+ D++ PPYS E R C+Y++ P
Sbjct: 171 TEMTAPSPTTVLYPSSGGNIHSFQAITPCAIFDILSPPYSSEHERHCTYFRRSQKKDLPV 350
Query: 237 EEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPPI 274
++ V + ++ LEE P++ + Y GP I
Sbjct: 351 NLQLDGVTASEVTW--LEEFQPPDDFVIRRGLYRGPVI 458
>TC8174 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;}, partial (24%)
Length = 1091
Score = 81.3 bits (199), Expect = 2e-16
Identities = 38/85 (44%), Positives = 58/85 (67%)
Frame = +2
Query: 193 GGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGL 252
GGN+H FTA+T CAVLDV+GPPYS +GR C+YY +YP++ + E I +++ + Y
Sbjct: 8 GGNMHCFTAVTACAVLDVLGPPYSDYEGRHCTYYNNYPFSDI-SVEGISIPEEERNGYEW 184
Query: 253 LEEIDMPENCQMDGIEYLGPPIDDT 277
L+E + E+ ++DG Y GP I ++
Sbjct: 185 LQEKEQLEDLEVDGKMYSGPKIHES 259
>AV425286
Length = 320
Score = 53.1 bits (126), Expect = 5e-08
Identities = 31/91 (34%), Positives = 49/91 (53%), Gaps = 12/91 (13%)
Frame = +2
Query: 43 LQELFDSCKQTFK--GPGTVPSPRDVHKLCHILDNMKPEDVGL-------SRDLQFF--- 90
+Q L+D CK F G + PS + + KL ILD ++P DVGL R FF
Sbjct: 38 VQTLYDHCKSIFSLSGKPSPPSSQALLKLSSILDTVQPTDVGLRVETADDDRGHGFFGVN 217
Query: 91 KPGNIIKENQRVTYTTVYKCDNFSLCIFFLP 121
+ + + Q +TY +++CD+F++C+F P
Sbjct: 218 QLNRVARWAQPITYVDIHECDSFTMCMFCFP 310
>TC10711
Length = 620
Score = 50.1 bits (118), Expect = 4e-07
Identities = 23/58 (39%), Positives = 34/58 (57%)
Frame = +3
Query: 217 KEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPPI 274
+++GR C+YY DYPY+ F + IG+ ++D Y L EI+ P + M Y GP I
Sbjct: 3 EDEGRKCTYYHDYPYSTFSGKGSIGD--GEEDEYAWLAEIETPGDLYMRSGVYAGPAI 170
>BI420225
Length = 319
Score = 39.3 bits (90), Expect = 7e-04
Identities = 19/51 (37%), Positives = 30/51 (58%)
Frame = +2
Query: 38 HVPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQ 88
++P +Q L+ CK +F G V S + K+C L+ +KP DVGL ++ Q
Sbjct: 161 NMPYYVQRLYRLCKASFSPDGPV-SQEAIEKVCEKLEKIKPSDVGLEQEAQ 310
>AV772637
Length = 551
Score = 34.7 bits (78), Expect = 0.018
Identities = 20/56 (35%), Positives = 31/56 (54%)
Frame = +2
Query: 39 VPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGN 94
+P LQ L+ CK +F G V S + K+ L+ +KP DVGL ++ Q + G+
Sbjct: 320 MPFYLQRLYRLCKASFAPDGPV-SEEAIAKVQEKLEKIKPIDVGLEQEAQVVRNGS 484
>TC20240 weakly similar to UP|Q9SJI9 (Q9SJI9) Expressed protein, partial
(25%)
Length = 495
Score = 32.0 bits (71), Expect = 0.12
Identities = 19/61 (31%), Positives = 29/61 (47%)
Frame = +2
Query: 214 PYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPP 273
PYS ++GR CSY++ P ++ V D ++ LEE P+ + Y GP
Sbjct: 2 PYSSDNGRHCSYFQKSSRKDLPETLEMNGVTVSDVTW--LEEFQPPDEFAIRRGLYRGPV 175
Query: 274 I 274
I
Sbjct: 176 I 178
>AV414604
Length = 427
Score = 27.3 bits (59), Expect = 2.9
Identities = 15/44 (34%), Positives = 21/44 (47%)
Frame = +2
Query: 159 HNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAI 202
H +P +L ++ + F CD SVL +T GN E AI
Sbjct: 155 HVSSRPELPAKLLRMHFHDCFVRGCDGSVLLNSTAGNTAEKDAI 286
>TC10912
Length = 781
Score = 26.9 bits (58), Expect = 3.9
Identities = 13/48 (27%), Positives = 24/48 (49%)
Frame = -1
Query: 60 VPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTV 107
VPS H L H +DN++ ++ + DL P + ++ ++ TV
Sbjct: 655 VPSSNQTHLLKHQIDNIQNKERIVQEDLSTLAPIRL*RDKVQIVIITV 512
>TC11913 weakly similar to UP|ABPA_PRUPE (Q9ZRA4) Auxin-binding protein
ABP19a precursor, partial (61%)
Length = 530
Score = 26.9 bits (58), Expect = 3.9
Identities = 9/24 (37%), Positives = 17/24 (70%)
Frame = +1
Query: 124 GVIPLHNHPGMTVFSKLLLGQMHI 147
GV+P+H+HPG + +L G++ +
Sbjct: 64 GVVPIHSHPGASELVIILQGRITV 135
>BP057775
Length = 452
Score = 26.6 bits (57), Expect = 5.0
Identities = 12/35 (34%), Positives = 20/35 (56%)
Frame = +3
Query: 12 VNKVCYVKRVIAKKRNKPYNHRRVKKHVPKALQEL 46
++K C +K A +KP HRR+ H+ +L +L
Sbjct: 333 ISKFCTLKV*HAPSVHKPSLHRRISTHILSSLPQL 437
>TC14752 similar to UP|Q9LDV4 (Q9LDV4) Alanine aminotransferase , partial
(49%)
Length = 1054
Score = 26.6 bits (57), Expect = 5.0
Identities = 19/52 (36%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Frame = -3
Query: 155 HEASHNLLQPSSKL--RLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITP 204
H NLL+PSS++ RLA+ +F+A D+ PT GG + + I P
Sbjct: 524 HVTPSNLLKPSSRIFARLAREDKIFSFSAMNDS*DSSPTLGGLMIKLARI*P 369
>TC8987 similar to UP|ABPA_PRUPE (Q9ZRA4) Auxin-binding protein ABP19a
precursor, partial (97%)
Length = 985
Score = 26.6 bits (57), Expect = 5.0
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = +1
Query: 124 GVIPLHNHPGMTVFSKLLLGQMHI 147
GVIPLH HPG L++ Q HI
Sbjct: 310 GVIPLHTHPGAN--ELLIVTQGHI 375
>BP074150
Length = 393
Score = 26.6 bits (57), Expect = 5.0
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -2
Query: 157 ASHNLLQPSSKLRLAKLKANKTFTAPC 183
A+H LLQPS L +L+ K F + C
Sbjct: 326 ANHRLLQPSFAGGLTELRTRKLFLSSC 246
>TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 564
Score = 26.6 bits (57), Expect = 5.0
Identities = 10/25 (40%), Positives = 20/25 (80%), Gaps = 1/25 (4%)
Frame = -1
Query: 31 NHRRVK-KHVPKALQELFDSCKQTF 54
NH+RV+ +H+ A++ +FD+ +QT+
Sbjct: 399 NHKRVRGRHLQMAMETMFDAERQTY 325
>TC16696 similar to UP|Q9AR81 (Q9AR81) Germin-like protein precursor,
partial (96%)
Length = 834
Score = 26.6 bits (57), Expect = 5.0
Identities = 12/20 (60%), Positives = 12/20 (60%)
Frame = +1
Query: 114 SLCIFFLPERGVIPLHNHPG 133
SL L GVIPLH HPG
Sbjct: 247 SLARLDLASGGVIPLHTHPG 306
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,426,325
Number of Sequences: 28460
Number of extensions: 84642
Number of successful extensions: 378
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 371
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 371
length of query: 279
length of database: 4,897,600
effective HSP length: 89
effective length of query: 190
effective length of database: 2,364,660
effective search space: 449285400
effective search space used: 449285400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC137822.15


