

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137703.1 + phase: 0
(233 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
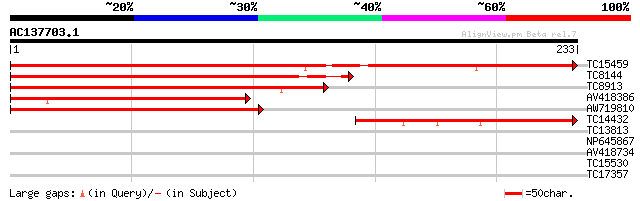
TC15459 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37,... 323 1e-89
TC8144 similar to UP|LB38_ARATH (Q9SN23) LOB domain protein 38, ... 245 4e-66
TC8913 similar to UP|Q7XYW2 (Q7XYW2) Seed specific protein Bn15D... 174 9e-45
AV418386 158 9e-40
AW719810 147 2e-36
TC14432 132 4e-32
TC13813 similar to UP|Q8BTG6 (Q8BTG6) CDNA: FLJ21415 FIS, partia... 26 6.8
NP645867 proliferating floral organs protein [Lotus japonicus] 26 6.8
AV418734 25 8.8
TC15530 similar to UP|AAQ84318 (AAQ84318) Fiber protein Fb27, pa... 25 8.8
TC17357 similar to UP|Q9FFC3 (Q9FFC3) Protease-like protein, par... 25 8.8
>TC15459 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37, partial
(46%)
Length = 1097
Score = 323 bits (829), Expect = 1e-89
Identities = 171/242 (70%), Positives = 187/242 (76%), Gaps = 9/242 (3%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWI+T EAQGHATVFVAKFFGRA LMSFISNVP+PQR
Sbjct: 126 MSCNGCRVLRKGCSESCILRPCLQWIDTAEAQGHATVFVAKFFGRADLMSFISNVPQPQR 305
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
P+LFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTL+PLPEL LDAP
Sbjct: 306 PSLFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLKPLPELLGLDAPS 485
Query: 121 ---TATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINL 177
TA D+ASE E W+IR+PN ++R +N S KR+R EE K+ +L
Sbjct: 486 HVPTAADDASEGE--VTYDTWRIRDPN---PNARLTNSRSSAGKRQRPEELAKLQTTTDL 650
Query: 178 DLRLTPIFQQKAV------EERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNL 231
+LRLTP F Q E RR GSPSM SEESVTTTACL +GIGD S DRKVL L
Sbjct: 651 NLRLTPGFLQNVSSYGCKREARRPGSPSMNSEESVTTTACLGSGIGDHCSRDEDRKVLKL 830
Query: 232 FI 233
F+
Sbjct: 831 FV 836
>TC8144 similar to UP|LB38_ARATH (Q9SN23) LOB domain protein 38, partial
(46%)
Length = 742
Score = 245 bits (626), Expect = 4e-66
Identities = 120/141 (85%), Positives = 127/141 (89%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPE QR
Sbjct: 345 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPETQR 524
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PALFQSLLFEACGRTVNPV GAVGLLWTGNWHVCQAAVETVLRGGTLRPLPEL+ DAP
Sbjct: 525 PALFQSLLFEACGRTVNPVKGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELSGFDAP- 701
Query: 121 TATDEASEAEDGGADAMWKIR 141
+DEAS+ + +W+I+
Sbjct: 702 --SDEASDTD------VWRIQ 740
>TC8913 similar to UP|Q7XYW2 (Q7XYW2) Seed specific protein Bn15D17A,
partial (53%)
Length = 615
Score = 174 bits (442), Expect = 9e-45
Identities = 81/141 (57%), Positives = 106/141 (74%), Gaps = 10/141 (7%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+C +RPCLQWI++PE+Q +ATVF+AKF+GRAGLM+ +++ PE R
Sbjct: 113 MSCNGCRVLRKGCSENCSIRPCLQWIKSPESQANATVFLAKFYGRAGLMNLVNSGPEHLR 292
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPL---------- 110
PA+F+SLL+EACGR VNP+ G+VGLLW+G+W +CQAAVE VL+G + P+
Sbjct: 293 PAIFRSLLYEACGRIVNPIYGSVGLLWSGSWQLCQAAVEAVLKGAPITPITSEAAASGRG 472
Query: 111 PELTLLDAPLTATDEASEAED 131
P L D + DE S A +
Sbjct: 473 PPLKAYDIRHVSRDENSAASN 535
>AV418386
Length = 352
Score = 158 bits (399), Expect = 9e-40
Identities = 73/100 (73%), Positives = 88/100 (88%), Gaps = 1/100 (1%)
Frame = +1
Query: 1 MSCNGCRVLRKGCS-ESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQ 59
MSCNGCRVLRKGCS + C+LR CL+WIE+P+AQ +ATVFVAKFFGRA LMSF+S+VP Q
Sbjct: 49 MSCNGCRVLRKGCSGDDCMLRHCLRWIESPQAQANATVFVAKFFGRATLMSFLSSVPTNQ 228
Query: 60 RPALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVE 99
R ALF+SLL+EA GR +NPV+GAVGLLW+GNW +CQ+ VE
Sbjct: 229 RSALFESLLYEAVGRALNPVSGAVGLLWSGNWQLCQSGVE 348
>AW719810
Length = 564
Score = 147 bits (370), Expect = 2e-36
Identities = 62/104 (59%), Positives = 85/104 (81%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
+SCNGCRVLRKGCS C +RPCL WI++P +Q +AT F+AKF+GRAGL++ ++ P+ +
Sbjct: 67 LSCNGCRVLRKGCSPDCPIRPCLNWIDSPASQANATFFLAKFYGRAGLLNLLTAAPQQLQ 246
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRG 104
P++F+SLL+EACGR VNP+ G+ GLL TG+WH+C+AAV VL G
Sbjct: 247 PSVFKSLLYEACGRIVNPIFGSTGLLSTGSWHLCEAAVNAVLIG 378
>TC14432
Length = 623
Score = 132 bits (333), Expect = 4e-32
Identities = 73/96 (76%), Positives = 79/96 (82%), Gaps = 5/96 (5%)
Frame = +2
Query: 143 PNFGLASSRSSNKVCSGS-KRRRSEEFVKVPAA---INLDLRLTPIFQQKAVEE-RRHGS 197
PNF SSRS+NKV +G KRRRSE+ VK+PAA INLDLRLTPIF QKA E RR GS
Sbjct: 5 PNFRFTSSRSNNKVSAGGGKRRRSEDHVKLPAAAAAINLDLRLTPIFLQKAPESNRRAGS 184
Query: 198 PSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI 233
PSM SEESVTTTACLE GIGD+W+HGGDRKVLNLFI
Sbjct: 185 PSMNSEESVTTTACLEGGIGDQWAHGGDRKVLNLFI 292
>TC13813 similar to UP|Q8BTG6 (Q8BTG6) CDNA: FLJ21415 FIS, partial (7%)
Length = 591
Score = 25.8 bits (55), Expect = 6.8
Identities = 12/27 (44%), Positives = 15/27 (55%), Gaps = 5/27 (18%)
Frame = +3
Query: 2 SCNGCRVLRKGCSE-----SCILRPCL 23
SC+GC VL + C+ SC L P L
Sbjct: 303 SCHGCNVLSQCCNSYEYCVSCCLNPAL 383
>NP645867 proliferating floral organs protein [Lotus japonicus]
Length = 1350
Score = 25.8 bits (55), Expect = 6.8
Identities = 25/114 (21%), Positives = 42/114 (35%)
Frame = +1
Query: 48 LMSFISNVPEPQRPALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTL 107
++ ++ +P RP LF S+ G T+ P + + G+ + AV+ +
Sbjct: 580 ILGSLTQLPPTLRPRLFPSI-----GLTITPT--CIDVTVAGDDMISPYAVKNL------ 720
Query: 108 RPLPELTLLDAPLTATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSK 161
T E+ + GG +MW P L S S VC+ K
Sbjct: 721 ---------------TSESFHIDGGGFYSMWGTTSPLPRLCSLESGRMVCAEGK 837
>AV418734
Length = 287
Score = 25.4 bits (54), Expect = 8.8
Identities = 13/45 (28%), Positives = 19/45 (41%), Gaps = 1/45 (2%)
Frame = -3
Query: 117 DAPLTATDEASEAEDGGADAMWKI-REPNFGLASSRSSNKVCSGS 160
D P+ DGG D+ W+ R +F + S + C GS
Sbjct: 213 DVPVVLIQNCCSCHDGGFDSRWRP*RMDSFD*GCNSSYMRTCHGS 79
>TC15530 similar to UP|AAQ84318 (AAQ84318) Fiber protein Fb27, partial (59%)
Length = 692
Score = 25.4 bits (54), Expect = 8.8
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = -1
Query: 75 TVNPVNGAVGLLWTGNWHVCQAAVETV 101
T+NP+N + GL W H+ AV+ +
Sbjct: 365 TLNPINWSSGLAWGRGQHLHPLAVQNL 285
>TC17357 similar to UP|Q9FFC3 (Q9FFC3) Protease-like protein, partial
(24%)
Length = 577
Score = 25.4 bits (54), Expect = 8.8
Identities = 11/24 (45%), Positives = 12/24 (49%)
Frame = -2
Query: 38 FVAKFFGRAGLMSFISNVPEPQRP 61
FV FFG I +PEP RP
Sbjct: 72 FVTSFFGAITYTCIIMVIPEPTRP 1
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,585,701
Number of Sequences: 28460
Number of extensions: 61800
Number of successful extensions: 318
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 313
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 313
length of query: 233
length of database: 4,897,600
effective HSP length: 87
effective length of query: 146
effective length of database: 2,421,580
effective search space: 353550680
effective search space used: 353550680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC137703.1


