

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
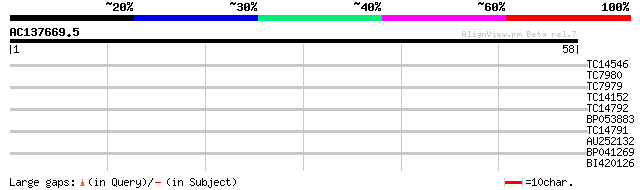
Query= AC137669.5 + phase: 0
(58 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxida... 28 0.28
TC7980 similar to UP|Q9SJM6 (Q9SJM6) At2g36320 protein (Zinc fin... 28 0.47
TC7979 similar to UP|Q9SJM6 (Q9SJM6) At2g36320 protein (Zinc fin... 28 0.47
TC14152 weakly similar to UP|AB18_PEA (Q06930) ABA-responsive pr... 25 3.1
TC14792 similar to UP|O73620 (O73620) Nuclear protein SDK2, part... 25 3.1
BP053883 25 3.1
TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), par... 25 3.1
AU252132 25 4.0
BP041269 24 6.8
BI420126 23 8.9
>TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (66%)
Length = 738
Score = 28.5 bits (62), Expect = 0.28
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = -2
Query: 2 ARRERQRKSKKQKWIWGSLTTV 23
ARRER+ +S +KW WG++ V
Sbjct: 80 ARRERRMESAAKKWEWGAIWIV 15
>TC7980 similar to UP|Q9SJM6 (Q9SJM6) At2g36320 protein (Zinc finger-like
protein), partial (36%)
Length = 670
Score = 27.7 bits (60), Expect = 0.47
Identities = 12/37 (32%), Positives = 19/37 (50%)
Frame = -3
Query: 1 MARRERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLP 37
++RR R + + + WIW +L V + G WS P
Sbjct: 608 VSRRRRSWRERFRLWIWSTLLAVPSNGCLCSIWSRDP 498
>TC7979 similar to UP|Q9SJM6 (Q9SJM6) At2g36320 protein (Zinc finger-like
protein), partial (73%)
Length = 1262
Score = 27.7 bits (60), Expect = 0.47
Identities = 12/37 (32%), Positives = 19/37 (50%)
Frame = -1
Query: 1 MARRERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLP 37
++RR R + + + WIW +L V + G WS P
Sbjct: 422 VSRRRRSWRERFRLWIWSTLLAVPSNGCLCSIWSRDP 312
>TC14152 weakly similar to UP|AB18_PEA (Q06930) ABA-responsive protein
ABR18, partial (92%)
Length = 827
Score = 25.0 bits (53), Expect = 3.1
Identities = 6/14 (42%), Positives = 13/14 (92%)
Frame = -2
Query: 4 RERQRKSKKQKWIW 17
+++QRK +K++W+W
Sbjct: 688 QQKQRKKRKREWLW 647
>TC14792 similar to UP|O73620 (O73620) Nuclear protein SDK2, partial (5%)
Length = 492
Score = 25.0 bits (53), Expect = 3.1
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -1
Query: 10 SKKQKWIWGSLTTVIALGTAA 30
S + W WGS +T +A G+ A
Sbjct: 165 SSTESWSWGSTSTGLAAGSTA 103
>BP053883
Length = 230
Score = 25.0 bits (53), Expect = 3.1
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = +1
Query: 3 RRERQRKSKKQKWIWGSLTTVI 24
RRE ++ + Q+W W SL +I
Sbjct: 148 RRENRKAASIQQWFW*SLVRII 213
>TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), partial (37%)
Length = 1779
Score = 25.0 bits (53), Expect = 3.1
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -1
Query: 10 SKKQKWIWGSLTTVIALGTAA 30
S + W WGS +T +A G+ A
Sbjct: 150 SSTESWSWGSTSTGLAAGSTA 88
>AU252132
Length = 350
Score = 24.6 bits (52), Expect = 4.0
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +1
Query: 20 LTTVIALGTAAVAWSYLPAGGESY 43
++ +A+ T V+W YLPA +SY
Sbjct: 184 ISAPMAVMTKRVSWEYLPATFKSY 255
>BP041269
Length = 417
Score = 23.9 bits (50), Expect = 6.8
Identities = 10/29 (34%), Positives = 15/29 (51%)
Frame = -3
Query: 10 SKKQKWIWGSLTTVIALGTAAVAWSYLPA 38
SK+ W WG+ ++ + AW LPA
Sbjct: 409 SKELVWGWGTCLELLVVHQ*TQAWDPLPA 323
>BI420126
Length = 441
Score = 23.5 bits (49), Expect = 8.9
Identities = 14/40 (35%), Positives = 23/40 (57%)
Frame = +2
Query: 16 IWGSLTTVIALGTAAVAWSYLPAGGESYSAEDHPVPKHDD 55
I +L T +AL TAA+++ + + HPVPKH++
Sbjct: 74 ILNNLWTWLALLTAALSFWRI-------RSSSHPVPKHNN 172
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.126 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,100,957
Number of Sequences: 28460
Number of extensions: 10634
Number of successful extensions: 57
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 57
length of query: 58
length of database: 4,897,600
effective HSP length: 34
effective length of query: 24
effective length of database: 3,929,960
effective search space: 94319040
effective search space used: 94319040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 49 (23.5 bits)
Medicago: description of AC137669.5


