

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
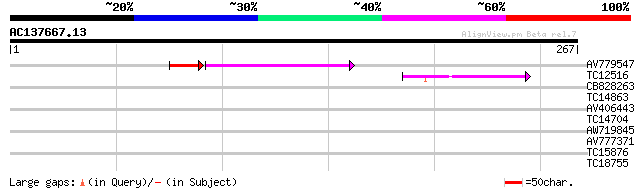
Query= AC137667.13 - phase: 0 /pseudo
(267 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV779547 58 3e-11
TC12516 similar to UP|Q9M5V7 (Q9M5V7) Disease resistance-like pr... 46 6e-06
CB828263 28 2.1
TC14863 28 2.1
AV406443 27 3.6
TC14704 27 3.6
AW719845 27 4.7
AV777371 27 4.7
TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 27 4.7
TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fr... 26 8.0
>AV779547
Length = 556
Score = 57.8 bits (138), Expect(2) = 3e-11
Identities = 35/70 (50%), Positives = 40/70 (57%)
Frame = +2
Query: 93 GACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMNKDSNKTLRWCKDG 152
GACKN AL FCL S+M + +R K EIM K F +M KDS K +W KDG
Sbjct: 308 GACKNWQKLLMAL*ERGRRFCLLSVM*LHLRCGNKAEIMEKLF*SMKKDSKKI*KWFKDG 487
Query: 153 GKLKHKSPIS 162
GKL K PIS
Sbjct: 488 GKL*QKWPIS 517
Score = 25.8 bits (55), Expect(2) = 3e-11
Identities = 13/16 (81%), Positives = 13/16 (81%)
Frame = +1
Query: 76 SLIFIVVFSKNYASST 91
S I IVVFSK YASST
Sbjct: 259 SQILIVVFSKYYASST 306
>TC12516 similar to UP|Q9M5V7 (Q9M5V7) Disease resistance-like protein
(Fragment), partial (36%)
Length = 859
Score = 46.2 bits (108), Expect = 6e-06
Identities = 27/62 (43%), Positives = 36/62 (57%), Gaps = 2/62 (3%)
Frame = +3
Query: 186 YKSTSLPNY--LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
Y +T +PN L G+D LE LL DVRV+G+ GMGGIGK IA ++N+
Sbjct: 513 YIATLVPNTKELIGVDKSIAHLES-LLCQESKDVRVIGIWGMGGIGKTTIAEEIFNRRHS 689
Query: 244 QF 245
Q+
Sbjct: 690 QY 695
>CB828263
Length = 570
Score = 27.7 bits (60), Expect = 2.1
Identities = 22/63 (34%), Positives = 30/63 (46%), Gaps = 8/63 (12%)
Frame = +3
Query: 113 CLFSMMLILMR*DIKKEIMLKPFSNMNKDSNKTLRWCKDGGKLKHKSPIS--------GW 164
C FSM+LI + +MLK S+M K CK GG+L K IS GW
Sbjct: 168 CPFSMILIPPMSATRPGLMLKHLSSMVK*IR-----CKSGGRL*EKQLISLVGIALSTGW 332
Query: 165 DVR 167
+++
Sbjct: 333 NLK 341
>TC14863
Length = 1803
Score = 27.7 bits (60), Expect = 2.1
Identities = 9/20 (45%), Positives = 16/20 (80%)
Frame = +1
Query: 238 YNKIFHQFPVLFLIDDLRKI 257
Y K++HQFP++ L+ ++ KI
Sbjct: 1657 YQKLYHQFPLMILLRNVMKI 1716
>AV406443
Length = 417
Score = 26.9 bits (58), Expect = 3.6
Identities = 15/49 (30%), Positives = 24/49 (48%), Gaps = 5/49 (10%)
Frame = +3
Query: 195 LAGMDSLTEELEKHL-----LLDSVDDVRVVGVCGMGGIGKKAIATALY 238
+A ++ L++ + H+ L D V+GV G G+GK I LY
Sbjct: 117 VASLNLLSDSWDFHIDRFLPFLTENTDFTVIGVIGPPGVGKSTIMNELY 263
>TC14704
Length = 587
Score = 26.9 bits (58), Expect = 3.6
Identities = 12/24 (50%), Positives = 17/24 (70%), Gaps = 1/24 (4%)
Frame = -3
Query: 224 GMGGIGKKAIATALYNKI-FHQFP 246
GMGGIGK+ I T+L ++ H+ P
Sbjct: 276 GMGGIGKEEIFTSLKQRVTLHELP 205
>AW719845
Length = 209
Score = 26.6 bits (57), Expect = 4.7
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = +3
Query: 215 DDVRVVGVCGMGGIGKKAIA 234
D V VV V G+GG+GK +A
Sbjct: 33 DGVSVVAVTGLGGLGKTTLA 92
>AV777371
Length = 515
Score = 26.6 bits (57), Expect = 4.7
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = -2
Query: 172 QIEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLD 212
QIE++ E+ MN GYK P A + L+K LLL+
Sbjct: 484 QIERL-EKFMNAPGYKKKVSPEIGAKNQEKLDSLKKRLLLE 365
>TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (13%)
Length = 558
Score = 26.6 bits (57), Expect = 4.7
Identities = 16/60 (26%), Positives = 28/60 (46%), Gaps = 4/60 (6%)
Frame = +1
Query: 128 KEIMLKPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHND----AQIEKIVEEIMNI 183
++ M+KP + KDS K +W L + + + N A I I++++MNI
Sbjct: 109 EKAMIKPETRYGKDSEKVKKW---RSSLSQVCDLKAFHYKENSGYERAFIPDIIKDVMNI 279
>TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fragment),
complete
Length = 880
Score = 25.8 bits (55), Expect = 8.0
Identities = 9/22 (40%), Positives = 15/22 (67%)
Frame = +3
Query: 218 RVVGVCGMGGIGKKAIATALYN 239
+V+ V GMGG+GK + +Y+
Sbjct: 261 KVISVTGMGGMGKTTLVKQVYD 326
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.339 0.149 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,473,475
Number of Sequences: 28460
Number of extensions: 55626
Number of successful extensions: 427
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 427
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 427
length of query: 267
length of database: 4,897,600
effective HSP length: 89
effective length of query: 178
effective length of database: 2,364,660
effective search space: 420909480
effective search space used: 420909480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC137667.13


