

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137666.7 - phase: 0
(1079 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
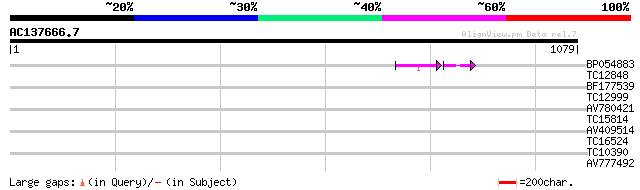
Score E
Sequences producing significant alignments: (bits) Value
BP054883 47 6e-09
TC12848 GB|AAA37921.1|553942|MUSIGF2RA insulin-like growth facto... 37 0.017
BF177539 33 0.18
TC12999 30 1.6
AV780421 29 3.5
TC15814 similar to UP|AAR08414 (AAR08414) Metal transport protei... 29 4.5
AV409514 28 5.9
TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC... 28 5.9
TC10390 similar to UP|Q9LHJ7 (Q9LHJ7) RING zinc finger protein-l... 28 5.9
AV777492 28 5.9
>BP054883
Length = 545
Score = 46.6 bits (109), Expect(2) = 6e-09
Identities = 30/96 (31%), Positives = 50/96 (51%), Gaps = 7/96 (7%)
Frame = -3
Query: 734 KNQSRPEGCIVERYIVEEAVEFFTDFLSNVESIGLPMSRHSG----RTTGEGIGASK--- 786
+N+SRPE I E Y+VEE + + +L L R +G + G IG K
Sbjct: 543 RNKSRPEVSIAEAYLVEECLTLCSRYLHGGVETRLNRMRRNGDRSHPSLGHPIGGKKKGE 364
Query: 787 LVTISDIELEQAHLYVLHNADEVEPYVKKHKEILQS 822
+++++ QAH Y+L N DEV+ +++ K +L +
Sbjct: 363 VISLNYKSKNQAHRYILFNHDEVQNFIR*IKILLST 256
Score = 31.6 bits (70), Expect(2) = 6e-09
Identities = 18/61 (29%), Positives = 26/61 (42%), Gaps = 1/61 (1%)
Frame = -1
Query: 826 NRSENWL-TREHNRRFISWLKEHIFSEIAKNPDSISERLRWLANGPNANVLSYSSYVINK 884
N+ + W + + FI W K D + L+ L+ GPN +S YVIN
Sbjct: 176 NKRKRWSKAKSQGQDFIEWFKTRALM------DDVPGHLKDLSRGPNIIARRFSGYVING 15
Query: 885 Y 885
Y
Sbjct: 14 Y 12
>TC12848 GB|AAA37921.1|553942|MUSIGF2RA insulin-like growth factor 2
receptor {Mus musculus;}, partial (23%)
Length = 664
Score = 37.0 bits (84), Expect = 0.017
Identities = 19/64 (29%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +3
Query: 7 SANRASKEYEDGVQEFVRFAIANAEDTSKIICPCLECCYTN-VSANDLQDHLICNGIDKS 65
S+ R +++E+ FVR+ + IICPC +C + + +++ DHL+C
Sbjct: 264 SSFRLRRKFEEDSIGFVRYWLEGKRGL--IICPCDKCGFNKRFTRDEVLDHLLCKPFPDG 437
Query: 66 YTCW 69
YT W
Sbjct: 438 YTFW 449
>BF177539
Length = 327
Score = 33.5 bits (75), Expect = 0.18
Identities = 19/55 (34%), Positives = 27/55 (48%), Gaps = 1/55 (1%)
Frame = +3
Query: 21 EFVRFAIANAEDTSKIICPCLECCYTNVSANDL-QDHLICNGIDKSYTCWTMHGE 74
EF R ++ A I+CPC C + D+ +HLI ++YT W HGE
Sbjct: 12 EFERSSVDGA-----ILCPCGLCNFRKWLTRDVVYEHLIIKQFPRNYTFWFDHGE 161
>TC12999
Length = 528
Score = 30.4 bits (67), Expect = 1.6
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = +3
Query: 499 GWKKKSIFFELPYWKSLYVRHFLDVMHIEKNVFDSV 534
GW S+FF LP+ SLY H L ++ V + V
Sbjct: 123 GWAASSLFFFLPHILSLYFLHLLPQEKVQSTVSNCV 230
>AV780421
Length = 582
Score = 29.3 bits (64), Expect = 3.5
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = +1
Query: 492 SGELVTGGWKKKSIFFELPYWKSL 515
+GELV GW +S ELPY K+L
Sbjct: 4 NGELVGSGWPTRSRITELPYIKTL 75
>TC15814 similar to UP|AAR08414 (AAR08414) Metal transport protein, partial
(14%)
Length = 655
Score = 28.9 bits (63), Expect = 4.5
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -1
Query: 670 KLPTLQNEIVVTLCELEMYFPPSFFDIMVHL 700
KL T QNE+ L L YF ++ +I++HL
Sbjct: 415 KLKTYQNELKTYLVHLNAYFQLAYSNILLHL 323
>AV409514
Length = 412
Score = 28.5 bits (62), Expect = 5.9
Identities = 12/35 (34%), Positives = 23/35 (65%)
Frame = +1
Query: 474 DQLYEKVKLLSSKFGKPFSGELVTGGWKKKSIFFE 508
++L++K++L++ G L TGGW+K ++FE
Sbjct: 325 EKLFKKLELVN--------GVLFTGGWEKSGLYFE 405
>TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC19_9
{Arabidopsis thaliana;}, partial (41%)
Length = 997
Score = 28.5 bits (62), Expect = 5.9
Identities = 20/81 (24%), Positives = 38/81 (46%), Gaps = 1/81 (1%)
Frame = +1
Query: 75 KKTKSA-KRRNERDPSTDFEKDTNYEFDQVEEILNVIEDDLRDCPKMFERVVIDAETPLY 133
KK K+A KR + E++TN+ + +LN ++ +D P +E+V +
Sbjct: 562 KKRKAADKRGGGEEEEGKVEEETNWSNGEDIALLNALKAFPKDAPMRWEKVAVAVPGKSK 741
Query: 134 EGCTKFTRLSTILKLYKLKAG 154
C K R++ + K ++ G
Sbjct: 742 AACVK--RVAELKKGFRSSKG 798
>TC10390 similar to UP|Q9LHJ7 (Q9LHJ7) RING zinc finger protein-like,
partial (44%)
Length = 722
Score = 28.5 bits (62), Expect = 5.9
Identities = 20/75 (26%), Positives = 28/75 (36%)
Frame = +1
Query: 62 IDKSYTCWTMHGEKKTKSAKRRNERDPSTDFEKDTNYEFDQVEEILNVIEDDLRDCPKMF 121
I S+T W+ T K+R +PS T+ E E N D+ +CP +
Sbjct: 262 IFSSFTMWSFASRAFTSITKKRRSAEPS-----HTSAECSDDEVCSNASRDEGLECPICW 426
Query: 122 ERVVIDAETPLYEGC 136
E I P C
Sbjct: 427 ESFNIVENVPYVLWC 471
>AV777492
Length = 472
Score = 28.5 bits (62), Expect = 5.9
Identities = 14/39 (35%), Positives = 23/39 (58%), Gaps = 4/39 (10%)
Frame = -2
Query: 707 ETQYCGPAYMRWMYPIERYMKILKGYVK----NQSRPEG 741
+ Q C P+Y +PIE +M+ L+G +K + RP+G
Sbjct: 339 QQQVC*PSYHHHNHPIEFFMRWLEG*IKMVGQKKQRPQG 223
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,036,933
Number of Sequences: 28460
Number of extensions: 318240
Number of successful extensions: 1496
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1484
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1495
length of query: 1079
length of database: 4,897,600
effective HSP length: 100
effective length of query: 979
effective length of database: 2,051,600
effective search space: 2008516400
effective search space used: 2008516400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137666.7


