

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.6 - phase: 0
(456 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
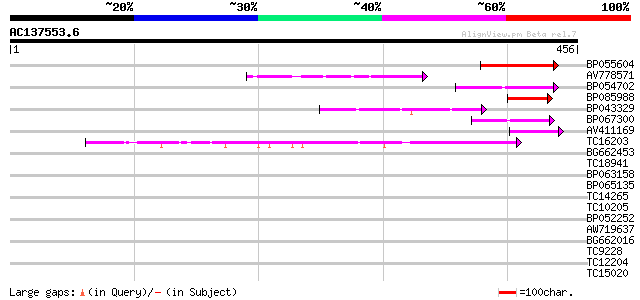
Score E
Sequences producing significant alignments: (bits) Value
BP055604 59 1e-09
AV778571 54 7e-08
BP054702 50 1e-06
BP085988 44 7e-05
BP043329 43 9e-05
BP067300 42 2e-04
AV411169 41 4e-04
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 40 8e-04
BG662453 39 0.001
TC18941 weakly similar to UP|Q8GZF0 (Q8GZF0) Resistance protein ... 39 0.002
BP063158 38 0.003
BP065135 37 0.009
TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resista... 37 0.009
TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%) 37 0.009
BP052252 35 0.026
AW719637 34 0.057
BG662016 34 0.057
TC9228 weakly similar to UP|Q8YPY5 (Q8YPY5) Glycine cleavage T-p... 33 0.075
TC12204 weakly similar to UP|Q9ZSN3 (Q9ZSN3) NBS-LRR-like protei... 33 0.13
TC15020 homologue to UP|Q9QX99 (Q9QX99) Transcription factor (T-... 31 0.37
>BP055604
Length = 464
Score = 59.3 bits (142), Expect = 1e-09
Identities = 31/63 (49%), Positives = 38/63 (60%)
Frame = -3
Query: 379 SLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSL 438
SL+ L I P L SLPD F L L++L I+ CP L R TGEDW KIAHVP + +
Sbjct: 447 SLRYLRIWGCPGLRSLPDGFQHLSALERLIIEGCPELKERCKEGTGEDWDKIAHVPDVYI 268
Query: 439 RLE 441
L+
Sbjct: 267 LLK 259
Score = 38.5 bits (88), Expect = 0.002
Identities = 21/40 (52%), Positives = 24/40 (59%)
Frame = -3
Query: 281 GLISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL 320
GL SLR L C L SLP L++L+RLII CPEL
Sbjct: 456 GLHSLRYLRIWGCPGLRSLPDGFQHLSALERLIIEGCPEL 337
>AV778571
Length = 452
Score = 53.5 bits (127), Expect = 7e-08
Identities = 47/147 (31%), Positives = 72/147 (48%), Gaps = 1/147 (0%)
Frame = +3
Query: 191 SIQSIPQFELPSLPSVKEV-YVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTV 249
SI IP +L LP+++++ +V E I FL L+ L I+ ++
Sbjct: 51 SITQIP--DLSGLPNLEKLSFVYSENLIKIHESVGFLN-------KLRILDIEGCCKIRT 203
Query: 250 LPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSL 309
P L SLEELY+ C+ LES P + + ++ ++ V + LP S+ LT L
Sbjct: 204 FPP--MKLASLEELYLSHCSSLESFP-EILAKMENITLIG-VYHTPIKELPSSIGNLTRL 371
Query: 310 QRLIIHYCPELILPANMNMLNSLREVS 336
+RL +H C L LP+ M M L E+S
Sbjct: 372 RRLELHGCGMLQLPSGMAM*TELEELS 452
>BP054702
Length = 528
Score = 49.7 bits (117), Expect = 1e-06
Identities = 32/85 (37%), Positives = 48/85 (55%), Gaps = 2/85 (2%)
Frame = -3
Query: 359 LSLRDFPSLRSLP-DWLGDTLSLQELEISKFPKLTSLPDNFDQL-ENLQKLCIDRCPRLV 416
L L +F SL+ L + L SL +L I K P L SLP+ DQL +L L +++CP L
Sbjct: 379 LGLYNFSSLKFLEGNGLQHLSSLHQLHIDKCPSLVSLPE--DQLPHSLAVLNLEKCPLLE 206
Query: 417 NRLARRTGEDWYKIAHVPILSLRLE 441
R R G+ W K++ +P++ + E
Sbjct: 205 ARYRIRKGKHWSKVSRIPVIKINEE 131
>BP085988
Length = 474
Score = 43.5 bits (101), Expect = 7e-05
Identities = 19/36 (52%), Positives = 24/36 (65%)
Frame = -2
Query: 401 LENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPIL 436
L L++L I+ CP L R TGEDW KIAHVP++
Sbjct: 473 LSALERLIIEGCPELKERCKEGTGEDWDKIAHVPLV 366
>BP043329
Length = 484
Score = 43.1 bits (100), Expect = 9e-05
Identities = 40/136 (29%), Positives = 68/136 (49%), Gaps = 2/136 (1%)
Frame = +3
Query: 250 LPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSL 309
+P E+ +L +LE +++ CN + IP+++ L L+ L + S+P S+T LTSL
Sbjct: 81 IPPEIGNLTNLEVVWLTQCNLVGVIPDSIG-NLKKLKDLDLALNDLYGSIPSSLTGLTSL 257
Query: 310 QRLIIHYCPELI--LPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSL 367
+++ + Y L LP M L LR + I L +P L++L+L +
Sbjct: 258 RQIEL-YNNSLSGELPRGMGNLTELRLLDASMNHLTGRIPEELCSLP-LESLNLYENRFE 431
Query: 368 RSLPDWLGDTLSLQEL 383
LP + D+ +L EL
Sbjct: 432 GELPASIADSPNLYEL 479
Score = 32.7 bits (73), Expect = 0.13
Identities = 22/82 (26%), Positives = 44/82 (52%)
Frame = +3
Query: 231 GKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSF 290
G + NL+ + + + + V+P+ + +L+ L++L + + SIP+++ GL SLR +
Sbjct: 96 GNLTNLEVVWLTQCNLVGVIPDSIGNLKKLKDLDLALNDLYGSIPSSL-TGLTSLRQIEL 272
Query: 291 VICHSLNSLPQSVTTLTSLQRL 312
LP+ + LT L+ L
Sbjct: 273 YNNSLSGELPRGMGNLTELRLL 338
>BP067300
Length = 523
Score = 42.4 bits (98), Expect = 2e-04
Identities = 26/67 (38%), Positives = 35/67 (51%)
Frame = -2
Query: 372 DWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIA 431
D L SL+ L I+ PKL +P ++ L I R PRL R R EDW KIA
Sbjct: 507 DSLQKLTSLETLGIACCPKLQCMPAKLPC--SISTLHIVRSPRLEERCRGRKSEDWPKIA 334
Query: 432 HVPILSL 438
H+P++ +
Sbjct: 333 HIPMIRI 313
>AV411169
Length = 424
Score = 41.2 bits (95), Expect = 4e-04
Identities = 21/43 (48%), Positives = 25/43 (57%)
Frame = +3
Query: 403 NLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSLRLESDVV 445
+LQ L I CP L R TGEDW KIAHVP + + + D V
Sbjct: 3 SLQLLSIALCPALAERCKDGTGEDWDKIAHVPKVQIDFD*DSV 131
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 40.0 bits (92), Expect = 8e-04
Identities = 85/379 (22%), Positives = 158/379 (41%), Gaps = 29/379 (7%)
Frame = +2
Query: 62 NTNS-TNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCS 120
N NS T E L L+ ++L + Y G IP G + +++L+ NC+
Sbjct: 716 NANSLTGRVPESLAKLKTLKELHLGYSNAYEG-GIP------PAFGSMENLRLLEMANCN 874
Query: 121 ---QLPP-LGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNL--ER 174
++PP LG L L++L++ QM N+ P E+S+ + SL ++++ DL E
Sbjct: 875 LTGEIPPSLGNLTKLHSLFV-QMNNLT--GTIPPELSSMMSLMSL-DLSINDLTGEIPES 1042
Query: 175 VLRIEGVEMLSQLSKLSIQSIPQF--ELPSLPSVK--EVYVGGETEEDIDHEASFL---- 226
+++ + +++ S+P F +LP+L +++ E ++ FL
Sbjct: 1043 FSKLKNLTLMNFFQNKFRGSLPSFIGDLPNLETLQVWENNFSFVLPHNLGGNGRFLYFDV 1222
Query: 227 --RDIAGKMP-------NLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNN 277
+ G +P LK +I +P + RSL ++ + + +P
Sbjct: 1223 TKNHLTGLIPPDLCKSGRLKTFIITDNFFRGPIPKGIGECRSLTKIRVANNFLDGPVPPG 1402
Query: 278 VFYGLISLRILSFVICHSLNSLP-----QSVTTLTSLQRLIIHYCPELILPANMNMLNSL 332
VF L S+ I LP +S+ TLT L +PA M L +L
Sbjct: 1403 VFQ-LPSVTITELSNNRLNGELPSVISGESLGTLTLSNNLFTGK-----IPAAMKNLRAL 1564
Query: 333 REVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLT 392
+ +S+ + I G+ +IP+L +++ +P + SL +++S+
Sbjct: 1565 QSLSLDANEFIGEIPGGVFEIPMLTKVNISGNNLTGPIPTTITHRASLTAVDLSRNNLAG 1744
Query: 393 SLPDNFDQLENLQKLCIDR 411
+P L +L L + R
Sbjct: 1745 EVPKGMKNLMDLSILNLSR 1801
>BG662453
Length = 460
Score = 39.3 bits (90), Expect = 0.001
Identities = 47/171 (27%), Positives = 75/171 (43%), Gaps = 1/171 (0%)
Frame = +2
Query: 226 LRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISL 285
L + G + LK L + + + LP + + R+LEEL + NKL +P+ + + L++L
Sbjct: 83 LPNSVGCLSKLKVLNVSG-NLIEYLPKSIENCRALEELNA-NFNKLSQLPDTMGFELLNL 256
Query: 286 RILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRG 345
+ LS V + L LP+S + LTSL+ IL A +N L
Sbjct: 257 KKLS-VNSNKLVFLPRSTSHLTSLK----------ILDARLNCL---------------- 355
Query: 346 IYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISK-FPKLTSLP 395
RSLPD L + ++L+ L +S+ F L +LP
Sbjct: 356 ----------------------RSLPDDLENLINLETLNVSQNFQYLDTLP 442
>TC18941 weakly similar to UP|Q8GZF0 (Q8GZF0) Resistance protein KR4,
partial (3%)
Length = 569
Score = 38.9 bits (89), Expect = 0.002
Identities = 24/64 (37%), Positives = 33/64 (51%)
Frame = +2
Query: 373 WLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAH 432
WL SL++LEI P+L SLP+ +L L I CP L G++ KIAH
Sbjct: 56 WLQHLTSLEKLEICYSPQLESLPEE-GLPSSLSVLIIXPCPLLEASCHSHGGKECPKIAH 232
Query: 433 VPIL 436
+P +
Sbjct: 233 IPCI 244
>BP063158
Length = 431
Score = 38.1 bits (87), Expect = 0.003
Identities = 30/84 (35%), Positives = 43/84 (50%), Gaps = 3/84 (3%)
Frame = -1
Query: 356 LQNLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDR-C 412
L L + FP+L+ L D+ G SL EL +S L LP+ +++ L I R C
Sbjct: 401 LHYLDISHFPNLKKL-DYRGLCHLPSLTELRLSDCSGLPCLPEE-GLPKSISTLSIRRMC 228
Query: 413 PRLVNRLARRTGEDWYKIAHVPIL 436
P L R + GEDW KI+H ++
Sbjct: 227 PLLKQRCQKPHGEDWGKISHSKLI 156
>BP065135
Length = 492
Score = 36.6 bits (83), Expect = 0.009
Identities = 17/62 (27%), Positives = 32/62 (51%)
Frame = -3
Query: 380 LQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSLR 439
L +L I + P L +LP++ L +++CP L R G+ W K++ +P++ +
Sbjct: 244 LHQLHIDQCPSLVALPEDATGHTALAV*SVEKCP*LEARYQIGKGKHWSKVSRIPVIKIN 65
Query: 440 LE 441
E
Sbjct: 64 EE 59
>TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resistance
protein-like protein MsR1, partial (5%)
Length = 766
Score = 36.6 bits (83), Expect = 0.009
Identities = 24/65 (36%), Positives = 34/65 (51%), Gaps = 3/65 (4%)
Frame = +1
Query: 351 EDIPLLQNLSLRDFPS---LRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKL 407
++I L+NL L S L+ LPD +G L+ L+IS L SLP+ L NL+ L
Sbjct: 1 QEIGKLENLELLRLSSCTDLKGLPDSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSL 180
Query: 408 CIDRC 412
+ C
Sbjct: 181 YMTSC 195
>TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%)
Length = 586
Score = 36.6 bits (83), Expect = 0.009
Identities = 19/61 (31%), Positives = 35/61 (57%)
Frame = +3
Query: 350 LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
+ ++ +Q L + D SL LP+ +G+ L++L + +LT LP + LENL+++
Sbjct: 21 ITNLEQVQFLDISDCISLSHLPEDMGELKKLEKLNVRGCSRLTELPTSIMDLENLEEVEC 200
Query: 410 D 410
D
Sbjct: 201 D 203
Score = 30.4 bits (67), Expect = 0.63
Identities = 16/46 (34%), Positives = 27/46 (57%)
Frame = +3
Query: 370 LPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
LP + + +Q L+IS L+ LP++ +L+ L+KL + C RL
Sbjct: 9 LPKSITNLEQVQFLDISDCISLSHLPEDMGELKKLEKLNVRGCSRL 146
>BP052252
Length = 472
Score = 35.0 bits (79), Expect = 0.026
Identities = 18/62 (29%), Positives = 33/62 (53%)
Frame = -2
Query: 103 SILGRLVDVKLMNCINCSQLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLT 162
S L +V++ L+ C +PPL LPFL +L++ +++I ++ FPSL
Sbjct: 453 SWLANIVEISLIKCNTXRTIPPLEHLPFLKSLFICYHGELEFI---YFQDGRRTFFPSLE 283
Query: 163 EM 164
++
Sbjct: 282 KL 277
>AW719637
Length = 571
Score = 33.9 bits (76), Expect = 0.057
Identities = 40/168 (23%), Positives = 78/168 (45%), Gaps = 8/168 (4%)
Frame = +2
Query: 250 LPNELSSLRSLEELYIIDCNKLE-SIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTS 308
+P E+ SL +L+ L + N LE +IP+N+ L SL L+ H +P+S+ +L +
Sbjct: 14 IPEEICSLSNLQSLSL-HTNFLEGNIPSNIG-NLSSLVNLTLFDNHLSGEIPKSIGSLRN 187
Query: 309 LQ-------RLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSL 361
LQ + + P I ++ L E SI G + + ++ + ++ +++
Sbjct: 188 LQVFRAGGNKNLKGEIPWEIGNCTSLVMLGLAETSISGS-----LPSSIQLLKRIKTIAI 352
Query: 362 RDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
S+P+ +G+ LQ L + + S+P +L L+ L +
Sbjct: 353 YTTLLSGSIPEEIGNCSELQNLYLYQNSISGSIPSQIGELSKLKSLLL 496
>BG662016
Length = 459
Score = 33.9 bits (76), Expect = 0.057
Identities = 42/158 (26%), Positives = 69/158 (43%), Gaps = 23/158 (14%)
Frame = +1
Query: 245 HQLTVLPNELSSLRSLEELYI----------------------IDCNKLESIPNNVFYGL 282
+QLT LPN + L L+ L + + NKL +P+ + + L
Sbjct: 19 NQLTSLPNSIGCLSKLKLLNVSGNFIESLPKTIENCSALEELNANFNKLSKLPDTIGFEL 198
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDR 342
I+L+ L+ V + L LP S + LT+L +L A +N L +L E
Sbjct: 199 INLKKLA-VNSNKLVLLPSSTSHLTAL----------TVLDARLNCLRALPE-------- 321
Query: 343 RRGIYNGLEDIPLLQNLSL-RDFPSLRSLPDWLGDTLS 379
LE++ L+ L++ ++F L +LP +G LS
Sbjct: 322 ------NLENLINLETLNVSQNFRYLDTLPYSIGLLLS 417
>TC9228 weakly similar to UP|Q8YPY5 (Q8YPY5) Glycine cleavage T-protein,
aminomethyltransferase, partial (6%)
Length = 576
Score = 33.5 bits (75), Expect = 0.075
Identities = 19/54 (35%), Positives = 27/54 (49%)
Frame = +1
Query: 383 LEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPIL 436
L IS KL ++PD + L++L I P GED++K+ HVP L
Sbjct: 4 LSISDCNKLKTVPDGLKFVTGLRELEIRWMPTSFKTRLGTAGEDYHKVQHVPSL 165
>TC12204 weakly similar to UP|Q9ZSN3 (Q9ZSN3) NBS-LRR-like protein cD7
(Fragment), partial (3%)
Length = 328
Score = 32.7 bits (73), Expect = 0.13
Identities = 20/62 (32%), Positives = 32/62 (51%)
Frame = +1
Query: 259 SLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCP 318
++ L++ C KL+S P + L+SL L+ C L S+P+ SL L I +C
Sbjct: 124 NMTNLHLEKCKKLKSFPQQMNKMLLSLMTLNIKECPELESIPEGGFP-DSLNLLEIFHCA 300
Query: 319 EL 320
+L
Sbjct: 301 KL 306
Score = 29.6 bits (65), Expect = 1.1
Identities = 26/97 (26%), Positives = 40/97 (40%), Gaps = 1/97 (1%)
Frame = +1
Query: 320 LILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTL- 378
++ + L SL + I G P + NL L L+S P + L
Sbjct: 19 VVTGVQLQYLQSLNSLRICNCPNFESFPEGGLRAPNMTNLHLEKCKKLKSFPQQMNKMLL 198
Query: 379 SLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
SL L I + P+L S+P+ ++L L I C +L
Sbjct: 199 SLMTLNIKECPELESIPEG-GFPDSLNLLEIFHCAKL 306
>TC15020 homologue to UP|Q9QX99 (Q9QX99) Transcription factor (T-cell
leukemia, homeobox 1), partial (5%)
Length = 1304
Score = 31.2 bits (69), Expect = 0.37
Identities = 29/104 (27%), Positives = 49/104 (46%), Gaps = 1/104 (0%)
Frame = +1
Query: 235 NLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICH 294
N+ ++ +D L +S L L L + + + IP+++ L +L+ L+
Sbjct: 232 NINQITLDPAGYSGSLTPLISQLTQLVTLDLAENSFSGPIPSSLS-SLSNLKTLTLRSNS 408
Query: 295 SLNSLPQSVTTLTSLQRL-IIHYCPELILPANMNMLNSLREVSI 337
S+P S+TTL SL L H LP +MN L +LR + +
Sbjct: 409 FSGSIPPSITTLKSLYSLDFSHNSLSGSLPNSMNSLINLRRIDL 540
Score = 26.9 bits (58), Expect = 7.0
Identities = 18/62 (29%), Positives = 31/62 (49%)
Frame = +1
Query: 346 IYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQ 405
I + L + L+ L+LR S+P + SL L+ S SLP++ + L NL+
Sbjct: 349 IPSSLSSLSNLKTLTLRSNSFSGSIPPSITTLKSLYSLDFSHNSLSGSLPNSMNSLINLR 528
Query: 406 KL 407
++
Sbjct: 529 RI 534
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.139 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,642,327
Number of Sequences: 28460
Number of extensions: 104545
Number of successful extensions: 546
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 510
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 540
length of query: 456
length of database: 4,897,600
effective HSP length: 93
effective length of query: 363
effective length of database: 2,250,820
effective search space: 817047660
effective search space used: 817047660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC137553.6


