

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.4 - phase: 0 /pseudo
(921 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
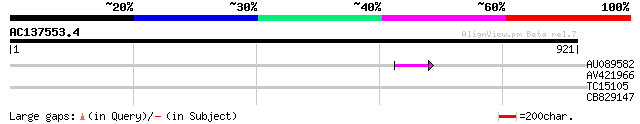
AU089582 44 1e-04
AV421966 31 0.78
TC15105 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein... 30 1.7
CB829147 28 8.6
>AU089582
Length = 383
Score = 43.9 bits (102), Expect = 1e-04
Identities = 23/64 (35%), Positives = 36/64 (55%)
Frame = +1
Query: 625 DCEVAWYSFEHCIK*RPEIYF*ILEKFARGFRYQVEVEFGVSSADRWSVGEDDSVVRGFV 684
DC +AW + + + R I+ LE F+ F ++ E+ SS++ WSV ED S +RG+
Sbjct: 43 DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WSVREDYSDLRGYA 222
Query: 685 EDLC 688
LC
Sbjct: 223 SCLC 234
>AV421966
Length = 469
Score = 31.2 bits (69), Expect = 0.78
Identities = 20/59 (33%), Positives = 34/59 (56%)
Frame = +2
Query: 512 EYSPGSYEDVSRSKEEILVVWIEARCGTVCVLLFDLLEVEN*ASETCWYDGVLRCARVE 570
+Y G Y+D+++ +EE++ + + G +L D L+V ASE YD VL+ AR +
Sbjct: 200 QYLAGRYKDITKFQEEVMGLPLP---GIEAILSSDELQV---ASEDAVYDFVLKWARTQ 358
>TC15105 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein AtPOB1,
partial (31%)
Length = 831
Score = 30.0 bits (66), Expect = 1.7
Identities = 20/57 (35%), Positives = 32/57 (56%)
Frame = +1
Query: 512 EYSPGSYEDVSRSKEEILVVWIEARCGTVCVLLFDLLEVEN*ASETCWYDGVLRCAR 568
+Y G Y+D+++ KEE++ + + G +L D L V+ SE YD VL+ AR
Sbjct: 487 QYLAGQYKDLTKFKEEVMALPLS---GIEVILASDHLRVK---SEDDVYDFVLKWAR 639
>CB829147
Length = 564
Score = 27.7 bits (60), Expect = 8.6
Identities = 12/29 (41%), Positives = 18/29 (61%), Gaps = 3/29 (10%)
Frame = +3
Query: 611 QFSCCIVG---RGLY*RDCEVAWYSFEHC 636
Q +CC+VG +G Y DC W++F +C
Sbjct: 285 QNACCLVGEK*KG*Y--DCNATWHTFIYC 365
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.366 0.163 0.618
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,196,082
Number of Sequences: 28460
Number of extensions: 205965
Number of successful extensions: 2713
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1461
Number of HSP's successfully gapped in prelim test: 73
Number of HSP's that attempted gapping in prelim test: 1204
Number of HSP's gapped (non-prelim): 1610
length of query: 921
length of database: 4,897,600
effective HSP length: 99
effective length of query: 822
effective length of database: 2,080,060
effective search space: 1709809320
effective search space used: 1709809320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137553.4


