

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.15 + phase: 0 /pseudo
(355 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
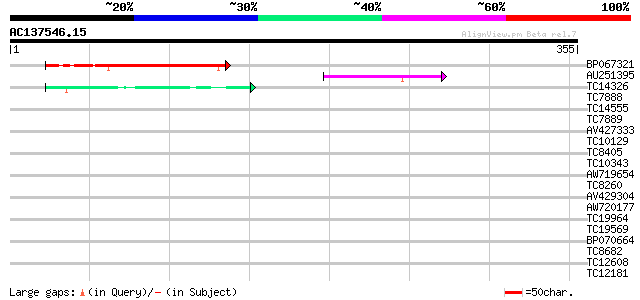
Score E
Sequences producing significant alignments: (bits) Value
BP067321 107 2e-24
AU251395 44 5e-05
TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific... 40 8e-04
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 39 0.002
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 38 0.003
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 37 0.004
AV427333 37 0.007
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 34 0.043
TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase is... 34 0.043
TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (... 33 0.056
AW719654 32 0.12
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 32 0.16
AV429304 32 0.21
AW720177 31 0.28
TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%) 31 0.36
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 31 0.36
BP070664 31 0.36
TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate... 30 0.47
TC12608 30 0.47
TC12181 similar to AAQ89616 (AAQ89616) At3g01370, partial (14%) 30 0.47
>BP067321
Length = 513
Score = 107 bits (268), Expect = 2e-24
Identities = 60/128 (46%), Positives = 79/128 (60%), Gaps = 12/128 (9%)
Frame = +3
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKT-------PENVKISEDGVSYVI 75
P K ++NP P + P +P LK + K KPV + +V I E GVSY++
Sbjct: 141 PPSKKSQNP--PPLPP--NPGLKHHHNSKY-FKPVERGGVISSDGDRSVIIGESGVSYLL 305
Query: 76 EGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKE-----F 130
GAPFEF++SY+ETPK+KP+ +REP F+PF P TMPRPWTG+ PL +K+K K F
Sbjct: 306 PGAPFEFQFSYSETPKAKPLAIREPAFLPFAPATMPRPWTGKAPLKSAKEKKKRRKVPLF 485
Query: 131 DSFVLPPP 138
DS PPP
Sbjct: 486 DSLKPPPP 509
>AU251395
Length = 306
Score = 43.5 bits (101), Expect = 5e-05
Identities = 24/81 (29%), Positives = 40/81 (48%), Gaps = 4/81 (4%)
Frame = +1
Query: 197 GFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVI----YRRGGVIY 252
G ++N+H HWK R + K+ C + V Q LE ++GG ++ +G I
Sbjct: 4 GVFDGTVENMHLHWKYRELVKVICKEGSLEAVHQVAQTLEAESGGILVAVERVSKGYAII 183
Query: 253 LFRGRNYNHKTRPRFPLMLWK 273
++RG+NY+ R +L K
Sbjct: 184 VYRGKNYSRPASLRPQTLLNK 246
>TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific
extensin-like protein precursor (PELP), partial (11%)
Length = 630
Score = 39.7 bits (91), Expect = 8e-04
Identities = 45/135 (33%), Positives = 50/135 (36%), Gaps = 3/135 (2%)
Frame = +2
Query: 23 PTGKPNKNPSKP--KVDPQSHPALKFSNIPKQKLKPVNK-TPENVKISEDGVSYVIEGAP 79
P P K PS P K P P +K + P K PV TP VK SY P
Sbjct: 230 PPQPPVKAPSPPIVKSPPYFPPPVKAPSPPIVKSPPVKAPTPPIVKSPP---SY-----P 385
Query: 80 FEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPH 139
K KS PV+ PP V P P + P PP K PPPH
Sbjct: 386 PPVKAPSPPIVKSPPVKAPTPPIVKSPPYYPPPV---KAPTPPQVKT---------PPPH 529
Query: 140 KKGVKPVQSPGPFLP 154
VKP +P P P
Sbjct: 530 PPVVKPPVAPIPSTP 574
Score = 26.2 bits (56), Expect = 8.9
Identities = 18/66 (27%), Positives = 26/66 (39%), Gaps = 6/66 (9%)
Frame = +2
Query: 99 EPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVL------PPPHKKGVKPVQSPGPF 152
+PP VT+P P + P PP + +K ++ PPP K P+ P
Sbjct: 161 KPPTPSTPAVTVPTPPLVKAPTPPPQPPVKAPSPPIVKSPPYFPPPVKAPSPPIVKSPPV 340
Query: 153 LPGTSP 158
T P
Sbjct: 341 KAPTPP 358
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 38.5 bits (88), Expect = 0.002
Identities = 43/144 (29%), Positives = 57/144 (38%), Gaps = 5/144 (3%)
Frame = +3
Query: 23 PTGKPNKNPSK--PKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPF 80
P KP K P P+ P P+ K + PK PV K P K +V + P
Sbjct: 267 PVHKPPKYPPSHGPEPCPPPKPSPKPPHYPKP---PVVKPPHVPKPPVVKPPHVPK--PP 431
Query: 81 EFKYSYTETPKSKPVQ-MREPPFVPFGPVTMPRPWTGRPPL--PPSKKKLKEFDSFVLPP 137
K Y P P + +PP+VP PV P P+ +PP+ PP +P
Sbjct: 432 IVKPPYVPKPPHVPKPPIVKPPYVPKPPVVKPPPYVPKPPVVSPP-----------YVPK 578
Query: 138 PHKKGVKPVQSPGPFLPGTSPRYV 161
P V P P P + +P YV
Sbjct: 579 PPVVPVTPPYVPKPPVVPVTPPYV 650
Score = 37.7 bits (86), Expect = 0.003
Identities = 40/144 (27%), Positives = 59/144 (40%), Gaps = 11/144 (7%)
Frame = +3
Query: 22 RPTGKPNKNPSKPKVDPQ---SHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGA 78
+P+ KP P P V P P +K ++PK P+ K P +
Sbjct: 327 KPSPKPPHYPKPPVVKPPHVPKPPVVKPPHVPKP---PIVKPP------------YVPKP 461
Query: 79 PFEFKYSYTETPK-SKPVQMREPPFVPFGPVTMPRPWTGRPPL----PPSKKK---LKEF 130
P K + P KP ++ PP+VP PV P P+ +PP+ PP K +
Sbjct: 462 PHVPKPPIVKPPYVPKPPVVKPPPYVPKPPVVSP-PYVPKPPVVPVTPPYVPKPPVVPVT 638
Query: 131 DSFVLPPPHKKGVKPVQSPGPFLP 154
+V PP K P+ P P++P
Sbjct: 639 PPYVPKPPIVK--PPIVYPPPYVP 704
Score = 31.2 bits (69), Expect = 0.28
Identities = 40/151 (26%), Positives = 54/151 (35%), Gaps = 15/151 (9%)
Frame = +3
Query: 23 PTGKPNKNPSKPKVDPQSH----PALKFSNIPKQKLKPVNKT----PENVKISEDGVSYV 74
P KP P P V P + P + +PK + PV P V ++ V
Sbjct: 480 PIVKPPYVPKPPVVKPPPYVPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKP 659
Query: 75 -IEGAPFEFKYSYTETPKS-KPVQMREPPFVPF-----GPVTMPRPWTGRPPLPPSKKKL 127
I P + Y P KP + PP+VP P+ P P+ PP+ PS
Sbjct: 660 PIVKPPIVYPPPYVPKPPIVKPPIVYPPPYVPTPPIVKPPIVYPPPYVPLPPVVPS---- 827
Query: 128 KEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
V PP + V P P P +P
Sbjct: 828 ---PPIVTPPTPTPPI--VTPPTPTPPVVTP 905
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 37.7 bits (86), Expect = 0.003
Identities = 44/165 (26%), Positives = 58/165 (34%), Gaps = 7/165 (4%)
Frame = +2
Query: 1 MALKLATTFPISASNADQSSRRP----TGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKP 56
+ + +A+ S SN+ +S P T +P S P S P ++ S P P
Sbjct: 155 ICIVIASVGAQSPSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPP 334
Query: 57 VNKTPENVKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTG 116
+P V S V AP + P S P PP P P P T
Sbjct: 335 AQSSPPPVS-SPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATPPPALTPVPATS 511
Query: 117 RPPLPPSKKKLKEFDSFVLPPPHKKGV---KPVQSPGPFLPGTSP 158
P P K+K P P V P ++PGP L SP
Sbjct: 512 PAPAP---AKVKSKSPAPAPAPALAPVVSLSPSEAPGPSLSSLSP 637
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 37.4 bits (85), Expect = 0.004
Identities = 40/136 (29%), Positives = 55/136 (40%), Gaps = 7/136 (5%)
Frame = +1
Query: 26 KPNKNPSKPKVDPQSHPALKFSNIPKQKL-KPVNKTPENVKISEDGVSYVIEGAPFEFKY 84
KP P P V P +K +PK + KP P+ +S YV +
Sbjct: 7 KPPYVPKPPHVPKP--PIVKPPYVPKPPVVKPPPYVPKPPVVSPP---YVPKPPVVPVTP 171
Query: 85 SYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPL--PPSKKKLKEFDSFVLPPPHKKG 142
Y P PV PP+VP P+ P P+ +PP+ PP V PPP+
Sbjct: 172 PYVPKPPVVPVT---PPYVPKPPIVYPPPYVPKPPIVKPP----------IVYPPPYVPT 312
Query: 143 ---VK-PVQSPGPFLP 154
VK P+ P P++P
Sbjct: 313 PPIVKPPIVYPPPYVP 360
Score = 33.9 bits (76), Expect = 0.043
Identities = 44/159 (27%), Positives = 56/159 (34%), Gaps = 23/159 (14%)
Frame = +1
Query: 23 PTGKPNKNPSKPKVDPQSH----PALKFSNIPKQKLKPVNKT----PENVKISEDGVS-- 72
P KP P P V P + P + +PK + PV P V ++ V
Sbjct: 49 PIVKPPYVPKPPVVKPPPYVPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKP 228
Query: 73 -------YV----IEGAPFEFKYSYTETPKS-KPVQMREPPFVPFGPVTMPRPWTGRPPL 120
YV I P + Y TP KP + PP+VP PV +P P PP
Sbjct: 229 PIVYPPPYVPKPPIVKPPIVYPPPYVPTPPIVKPPIVYPPPYVPLPPV-VPSPPIVTPPT 405
Query: 121 P-PSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
P P V+ PP P P P +P P
Sbjct: 406 PTPPIVTPPTPTPPVVTPPSPPTETPCPPPPPVVPSPPP 522
>AV427333
Length = 387
Score = 36.6 bits (83), Expect = 0.007
Identities = 29/95 (30%), Positives = 33/95 (34%), Gaps = 15/95 (15%)
Frame = +3
Query: 82 FKYSYTETPKSKPVQMREPPFVPFGPVTMPR---------------PWTGRPPLPPSKKK 126
F + + P S P PP PF P P P G P PPS
Sbjct: 63 FYFPHFPPPPSHPYNPPTPPSHPFNPPPTPSHYFKSPPPPSHSFAPPPRGHPSPPPSSPP 242
Query: 127 LKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYV 161
F PPPH V+P P P P T P +V
Sbjct: 243 PPSSHPFTPPPPH---VRP--PPSPHHPITPPPHV 332
Score = 30.8 bits (68), Expect = 0.36
Identities = 31/126 (24%), Positives = 44/126 (34%), Gaps = 2/126 (1%)
Frame = +3
Query: 28 NKNPSKPKVDPQSHPALKFSNIPKQKLKPVNK-TPENVKISEDGVSYVIEGAPFEFKYSY 86
+K+ SK K+ + P F + P P N TP + + +P +S+
Sbjct: 18 DKSRSKSKM-ASTQPNFYFPHFPPPPSHPYNPPTPPSHPFNPPPTPSHYFKSPPPPSHSF 194
Query: 87 TETPKSKPVQMREPPFVPFG-PVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKP 145
P+ P P P P T P P PP P + PPPH + P
Sbjct: 195 APPPRGHPSPPPSSPPPPSSHPFTPPPPHVRPPPSPHHP---------ITPPPHVRPPPP 347
Query: 146 VQSPGP 151
P P
Sbjct: 348 PLPPSP 365
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 33.9 bits (76), Expect = 0.043
Identities = 27/96 (28%), Positives = 35/96 (36%), Gaps = 2/96 (2%)
Frame = +1
Query: 86 YTETPKSKPVQMREPPFVPFGPVTMPRPWTGRP--PLPPSKKKLKEFDSFVLPPPHKKGV 143
YT TP P +PP++P P T P P + P P PPS + P P K
Sbjct: 169 YTPTPPKTPSPTSQPPYIPLPPKT-PSPISQPPHVPTPPSTPSPISQPPYT-PTPPKTPS 342
Query: 144 KPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINE 179
Q P P +P + P T N+
Sbjct: 343 PTSQPPHTPTPPNTPSPISQPPYTPTPPKTPSPTNQ 450
>TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase isolog,
25331-24357 (At1g11090), partial (18%)
Length = 552
Score = 33.9 bits (76), Expect = 0.043
Identities = 29/126 (23%), Positives = 40/126 (31%), Gaps = 10/126 (7%)
Frame = +3
Query: 4 KLATTFPISASNADQSSRRPTGKPNKNPSKPKVD----PQSHPALKFSNIPKQKLKPVNK 59
K TT P ++ + +S PT + + +PS P P S P + +
Sbjct: 90 KRNTTPPKESATPNPTSTLPTARSSPSPSSPSTPKSKPPSS*PTATAPTPAGSSRRSAST 269
Query: 60 TPENVKISEDGVSYVIEGAPFEFKYSYTETPKSKP------VQMREPPFVPFGPVTMPRP 113
+P S S S T T P P PF P + P
Sbjct: 270 SPPGATPSSPPTSSATAAPTASAATSATSTASPPPRSPSSSTSAAASPTPPFRPSSSASP 449
Query: 114 WTGRPP 119
W G PP
Sbjct: 450 WAGSPP 467
>TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (Fibrillarin
2) (AtFib2), partial (85%)
Length = 1116
Score = 33.5 bits (75), Expect = 0.056
Identities = 26/59 (44%), Positives = 29/59 (49%)
Frame = -2
Query: 100 PPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
PPF+P P PRP RPPLPP LPPP + +K V P P LP SP
Sbjct: 233 PPFIP--PPLPPRP-PPRPPLPPPPP---------LPPP--RALKGVPLPPP-LPPPSP 102
>AW719654
Length = 491
Score = 32.3 bits (72), Expect = 0.12
Identities = 17/42 (40%), Positives = 18/42 (42%)
Frame = +3
Query: 98 REPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPH 139
R PP P P P P T PP P K L F F +P H
Sbjct: 210 RHPPLPPPPPPLPPLPQTATPPPLPQTKTLPPFHFFNVPTLH 335
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 32.0 bits (71), Expect = 0.16
Identities = 39/134 (29%), Positives = 48/134 (35%), Gaps = 3/134 (2%)
Frame = +3
Query: 8 TFP--ISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVK 65
+FP ISA++ SS P KP S P P + P +K PV +P
Sbjct: 78 SFPSEISANHYAYSSPPPPPKPYSYHSPP-------PPVHSPPPPYEKPHPVYHSPPPPY 236
Query: 66 ISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFG-PVTMPRPWTGRPPLPPSK 124
P +S P KP + PP P P P P PP PP K
Sbjct: 237 EK-----------PHPVYHSPPPPPPHKPYKYPSPPPPPHKYPHPHPHPVYHSPPPPPPK 383
Query: 125 KKLKEFDSFVLPPP 138
K K + PPP
Sbjct: 384 KHYK----YSSPPP 413
Score = 28.1 bits (61), Expect = 2.3
Identities = 20/62 (32%), Positives = 23/62 (36%)
Frame = +3
Query: 90 PKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSP 149
P PV PP P P P PP PP +K ++ S PP H SP
Sbjct: 429 PHPHPVYHSPPP--PVHTYPHPHPVYHSPPPPPPHEKPYKYASPPPPPVHTYPHPVYHSP 602
Query: 150 GP 151
P
Sbjct: 603 PP 608
>AV429304
Length = 355
Score = 31.6 bits (70), Expect = 0.21
Identities = 28/89 (31%), Positives = 34/89 (37%), Gaps = 3/89 (3%)
Frame = +3
Query: 86 YTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKP 145
Y P P + PP VP P T P+ PP PS V PP+ P
Sbjct: 66 YVPIPPKTPSPVYSPPNVPSPPQTPHAPYVPTPPKTPSP---------VYSPPNVP--SP 212
Query: 146 VQSP-GPFLPGTS--PRYVMSREEVLGEP 171
Q+P P++P S P V S V P
Sbjct: 213 PQTPHAPYVPTPSKTPSXVYSPPNVPTPP 299
>AW720177
Length = 505
Score = 31.2 bits (69), Expect = 0.28
Identities = 34/164 (20%), Positives = 60/164 (35%), Gaps = 11/164 (6%)
Frame = +2
Query: 12 SASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGV 71
S S++ + ++ K K K + H SN P ++ P P+N ++
Sbjct: 17 SLSHSQKKKKKKKKKKKKLLKKHEGIGSYHRLRSPSNFPPRRRNPHRPKPQNQVVTRPSQ 196
Query: 72 SYVIEGAPFEFKYSYTETPKSKPVQMREPPF---------VPFGPVTMPRPWTGRPPLPP 122
++ P F + + +P R P +P V +P P R P
Sbjct: 197 THQPPNPPHRFLRRVSLLLRHQPTLSRPLPLPLRHSPLRRLPRATVPLPHPPPSRXAPNP 376
Query: 123 SKKKLKEFDSFVLPPPHKK-GVKPVQSPGP-FLPGTSPRYVMSR 164
+ + LPP H + P++ P + P PRY+ +R
Sbjct: 377 KRPR--------LPPRHPRLPPLPLRHTTP*WTPPRRPRYLQAR 484
>TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%)
Length = 562
Score = 30.8 bits (68), Expect = 0.36
Identities = 19/51 (37%), Positives = 22/51 (42%), Gaps = 2/51 (3%)
Frame = -2
Query: 90 PKSKPVQMREPPFVPFG--PVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
P P R PP P P+ PRP + RPP PP +LPPP
Sbjct: 276 PPLPPRPPRSPPRPPLPNRPLPPPRPLSNRPPRPPP----------LLPPP 154
Score = 28.9 bits (63), Expect = 1.4
Identities = 19/56 (33%), Positives = 25/56 (43%), Gaps = 3/56 (5%)
Frame = -2
Query: 102 FVPFGPVTMPRPWTGRPPLPPSKKKLKE---FDSFVLPPPHKKGVKPVQSPGPFLP 154
F+P +PRP PPLPP + + LPPP +P + P P LP
Sbjct: 324 FLPAALKGVPRPPLPPPPLPPRPPRSPPRPPLPNRPLPPPRPLSNRPPRPP-PLLP 160
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 30.8 bits (68), Expect = 0.36
Identities = 23/78 (29%), Positives = 27/78 (34%), Gaps = 1/78 (1%)
Frame = +1
Query: 87 TETPKSKPVQMRE-PPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKP 145
T TP P ++ E PP P P P PP+ S P H P
Sbjct: 130 TTTPPPSPTKVTENPPTAPSTSTKSPTSSASEEPSPPNPS-----TSTTQQPKHSP-TSP 291
Query: 146 VQSPGPFLPGTSPRYVMS 163
SP P P T P M+
Sbjct: 292 PASPSPSPPSTKPPTAMA 345
>BP070664
Length = 392
Score = 30.8 bits (68), Expect = 0.36
Identities = 20/60 (33%), Positives = 28/60 (46%), Gaps = 2/60 (3%)
Frame = +2
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHP--ALKFSNIPKQKLKPVNKTPENVKIS 67
PI N+ R PTG P + + V S P A N+PK + KPVN +++S
Sbjct: 131 PIKYMNSPTHLRSPTGPPIRVGVRAAVKGVSSPIGAAYMHNLPKAQAKPVNLW*SRIQMS 310
>TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate
dioxygenase (4HPPD) (HPD) (HPPDase) , partial (29%)
Length = 576
Score = 30.4 bits (67), Expect = 0.47
Identities = 23/89 (25%), Positives = 33/89 (36%), Gaps = 4/89 (4%)
Frame = +3
Query: 105 FGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMSR 164
F P T R + PP+ P L+ S++ PPP P P + +M +
Sbjct: 6 FQPATSRRHFHHHPPITPRHVSLRRHFSYITPPP------------PHHPKPNTHSIMGK 149
Query: 165 EEVLGEPLTKE----EINELVRSTLKSSR 189
+ E + N VRS KS R
Sbjct: 150 QTTTTEEVQTGFKLLGFNRFVRSNPKSDR 236
>TC12608
Length = 735
Score = 30.4 bits (67), Expect = 0.47
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = -2
Query: 12 SASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNI 49
SAS PT +P +NPS PQ+HP FSN+
Sbjct: 209 SASPTQGHYPNPTSQPGQNPSH---SPQTHPFSPFSNL 105
>TC12181 similar to AAQ89616 (AAQ89616) At3g01370, partial (14%)
Length = 682
Score = 30.4 bits (67), Expect = 0.47
Identities = 19/66 (28%), Positives = 33/66 (49%), Gaps = 4/66 (6%)
Frame = +2
Query: 275 VPPVYPRLIQQVPEG----LTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEE 330
V P Y + + +P G LT +E T +R+ R + LG+N L + + +E
Sbjct: 191 VVPGYRQPFRLLPYGVKPKLTDDEMTTLRRLSRPIPSHFALGRNKKLQGLAAAIIKLWER 370
Query: 331 CELVRV 336
CE+V++
Sbjct: 371 CEIVKI 388
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,730,746
Number of Sequences: 28460
Number of extensions: 119000
Number of successful extensions: 1054
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 965
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1011
length of query: 355
length of database: 4,897,600
effective HSP length: 91
effective length of query: 264
effective length of database: 2,307,740
effective search space: 609243360
effective search space used: 609243360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC137546.15


