

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.13 - phase: 0
(363 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
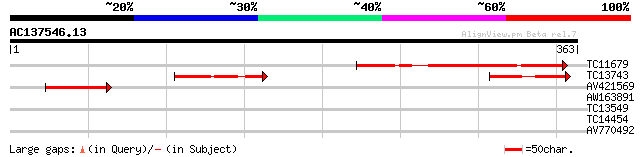
TC11679 similar to UP|Q8I738 (Q8I738) TcC31.4, partial (8%) 140 4e-34
TC13743 76 8e-15
AV421569 62 2e-10
AW163891 28 1.8
TC13549 similar to UP|Q9STG6 (Q9STG6) dUTP pyrophosphatase-like ... 28 3.2
TC14454 similar to UP|O04133 (O04133) SRC2, partial (88%) 28 3.2
AV770492 27 7.0
>TC11679 similar to UP|Q8I738 (Q8I738) TcC31.4, partial (8%)
Length = 652
Score = 140 bits (352), Expect = 4e-34
Identities = 75/135 (55%), Positives = 99/135 (72%)
Frame = +3
Query: 223 SFRSEASITPSVLATMTQIAHAELPRLILPKFGSKVSDQFVESGRQVPVMVPVSAAMAAF 282
SFR+EA I+P+VLAT++ +AHAELP ++ G ++PV++P+ AA+AA
Sbjct: 3 SFRTEAGISPAVLATLSHVAHAELP--LVASAGETT---------KLPVVMPLGAALAAC 149
Query: 283 ALHLQLRYGEKSDGVVTCRDAEVPGSVVVRPNMKLDHAWMVYSSNSKKKKSSEPDAREMC 342
A LQ+RYGEKSDG+VTCRDAEVPGSVVVRP KLDHAWMVYS S +E DA ++C
Sbjct: 150 AQLLQVRYGEKSDGLVTCRDAEVPGSVVVRPTRKLDHAWMVYS--SLNDDPAEGDASQVC 323
Query: 343 QAIFTLLVELGKTER 357
+A+ TLLVE+G+ +R
Sbjct: 324 EALLTLLVEVGQKKR 368
>TC13743
Length = 504
Score = 76.3 bits (186), Expect = 8e-15
Identities = 40/60 (66%), Positives = 45/60 (74%)
Frame = +3
Query: 106 EIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIA 165
EI +YIEE YWGS K V+LLGHSK G+ AAAALSLYW DL + + QSPYGGTPIA
Sbjct: 12 EIIEYIEETYWGSKKRVLLLGHSK-GVHAAAALSLYWYDLN----EIGIAQSPYGGTPIA 176
Score = 42.4 bits (98), Expect = 1e-04
Identities = 21/52 (40%), Positives = 32/52 (61%)
Frame = +3
Query: 308 SVVVRPNMKLDHAWMVYSSNSKKKKSSEPDAREMCQAIFTLLVELGKTEREV 359
SVVV P K+DHAW VY+ + DA ++C+A+ +LLVE+ + E+
Sbjct: 207 SVVVGPKQKMDHAWTVYTPLN--------DASQVCEALLSLLVEMAE*SHEL 338
>AV421569
Length = 262
Score = 61.6 bits (148), Expect = 2e-10
Identities = 27/42 (64%), Positives = 34/42 (80%)
Frame = +2
Query: 24 LQHAPGMPPVHDGSSRFLELLSDIRNGKDSIPSSFVYLLIPG 65
LQ APGMPPV DG+ RFLE+L +I++G +P+S VYLLIPG
Sbjct: 5 LQRAPGMPPVEDGTERFLEILGNIKHGVHKLPNSVVYLLIPG 130
>AW163891
Length = 335
Score = 28.5 bits (62), Expect = 1.8
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = +1
Query: 25 QHAPGMPPVHDGSSRFLELLSDIRNGKDSIPSSFVYLLIP 64
QH G P+ +GSS LE+ +D N S + + IP
Sbjct: 136 QHVYGYKPIPEGSSESLEISTDRENAASLAKSLYTAIGIP 255
>TC13549 similar to UP|Q9STG6 (Q9STG6) dUTP pyrophosphatase-like protein ,
partial (76%)
Length = 597
Score = 27.7 bits (60), Expect = 3.2
Identities = 21/57 (36%), Positives = 31/57 (53%), Gaps = 6/57 (10%)
Frame = +3
Query: 151 GLALVQSPYGGTPIA------SDILREGQIGDKETRRILELIICKIIKGDIRALEDL 201
G ++ + Y G P+ SD+ E +IGD RI +LII KI+ D+ +EDL
Sbjct: 183 GAGVIDADYRG-PVGVILFNHSDLDFEVKIGD----RIAQLIIEKIVTPDVAEVEDL 338
>TC14454 similar to UP|O04133 (O04133) SRC2, partial (88%)
Length = 1190
Score = 27.7 bits (60), Expect = 3.2
Identities = 25/88 (28%), Positives = 40/88 (45%)
Frame = +1
Query: 212 KHKLPLDIPLISFRSEASITPSVLATMTQIAHAELPRLILPKFGSKVSDQFVESGRQVPV 271
+++L L++ L+S R+ A T V+ T+ H + L+ G D F RQV
Sbjct: 394 QNRLSLEVKLLSDRTLAGDT--VIGTV----HIPIKELLDNPGGGGGGDSF----RQVSY 543
Query: 272 MVPVSAAMAAFALHLQLRYGEKSDGVVT 299
V S+ A + ++GEK D T
Sbjct: 544 QVKRSSGKAKGTFNFSYKFGEKVDAPAT 627
>AV770492
Length = 479
Score = 26.6 bits (57), Expect = 7.0
Identities = 17/38 (44%), Positives = 20/38 (51%)
Frame = -3
Query: 326 SNSKKKKSSEPDAREMCQAIFTLLVELGKTEREVEQVL 363
SN KKK+ S PD RE C +T GK EV +L
Sbjct: 390 SN*KKKEMSLPDQRECCYRHWT--C*KGKDTEEVSAIL 283
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,678,936
Number of Sequences: 28460
Number of extensions: 74064
Number of successful extensions: 345
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 337
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 340
length of query: 363
length of database: 4,897,600
effective HSP length: 91
effective length of query: 272
effective length of database: 2,307,740
effective search space: 627705280
effective search space used: 627705280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC137546.13


