

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137081.4 - phase: 0
(325 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
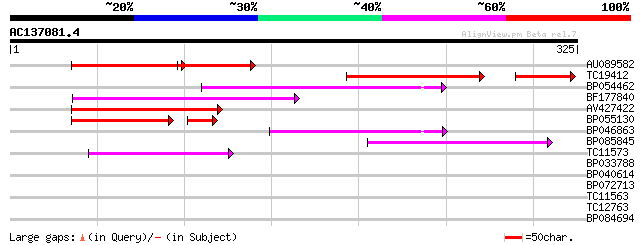
Score E
Sequences producing significant alignments: (bits) Value
AU089582 89 4e-33
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 97 3e-28
BP054462 92 9e-20
BF177840 89 1e-18
AV427422 86 1e-17
BP055130 62 9e-12
BP046863 60 7e-10
BP085845 58 2e-09
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 51 3e-07
BP033788 29 0.95
BP040614 28 2.1
BP072713 27 4.7
TC11563 27 4.7
TC12763 similar to UP|CADH_MEDSA (P31656) Cinnamyl-alcohol dehyd... 27 6.1
BP084694 26 8.0
>AU089582
Length = 383
Score = 89.4 bits (220), Expect(2) = 4e-33
Identities = 41/66 (62%), Positives = 52/66 (78%)
Frame = +2
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQ 95
P SI+SDR +FTS FW+S Q ALG++L++S+A+HPQTDGQSERTIQ LED+LR CV +
Sbjct: 65 PVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQILEDMLRACVXDL 244
Query: 96 GGASDS 101
G S
Sbjct: 245 RGXLGS 262
Score = 68.6 bits (166), Expect(2) = 4e-33
Identities = 31/45 (68%), Positives = 36/45 (79%)
Frame = +3
Query: 97 GASDSHLPLIEFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFESG 141
G+ D +L L+EF YNNSY SSI MAPFEALYG RCR+P+ WFE G
Sbjct: 249 GSWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPIGWFEVG 383
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 97.1 bits (240), Expect(2) = 3e-28
Identities = 46/79 (58%), Positives = 62/79 (78%)
Frame = +3
Query: 194 RVTPLTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYV 253
RV+P+ GV R K KL+P+FIGP+++ ERVG+V+YR+ LPP LS +H VFHVS LRKY+
Sbjct: 3 RVSPMKGVLRFGKKGKLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRKYL 182
Query: 254 ADPSHVIPRDDVQVRDNLT 272
DPSHVI +DVQ+ +L+
Sbjct: 183 YDPSHVIRHEDVQLDVHLS 239
Score = 44.3 bits (103), Expect(2) = 3e-28
Identities = 18/34 (52%), Positives = 23/34 (66%)
Frame = +2
Query: 291 KEIPLVRVVWGGATGESLTWELESKMRESYPELF 324
K++ V+V+W G +GE TWE E MRE YP LF
Sbjct: 296 KDVGSVKVLWRGPSGEEATWEAEDIMREKYPHLF 397
>BP054462
Length = 422
Score = 92.4 bits (228), Expect = 9e-20
Identities = 53/141 (37%), Positives = 81/141 (56%), Gaps = 1/141 (0%)
Frame = +1
Query: 111 NNSYHSSIGMAPFEALYGWRCRTPLCWFESGESVVLGPDLVHETTEKVRMIREKMKASQS 170
N++Y+ S M+PF+ALYG L + DL E + +R + SQ
Sbjct: 1 NSNYNRSAKMSPFQALYGREPPVLLQGTTIPSKIAAVNDLQVGRDELLSDLRANLLKSQD 180
Query: 171 RQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALK-SRKLTPKFIGPYQISERVGTVAY 229
++Y +K+R+D+++Q GD VFL++ P A K + KL+P++ GPY I ++G VAY
Sbjct: 181 MMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAKIGAVAY 360
Query: 230 RVGLPPHLSNLHDVFHVSQLR 250
R+ LP H S +H VFHVS L+
Sbjct: 361 RLELPAH-SRVHPVFHVSLLK 420
>BF177840
Length = 410
Score = 89.0 bits (219), Expect = 1e-18
Identities = 51/130 (39%), Positives = 73/130 (55%)
Frame = +2
Query: 37 SSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQG 96
+SIVSDRD +F S FW++L +G+KL S+ HPQTDGQ+E ++L LLR +
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 97 GASDSHLPLIEFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFESGESVVLGPDLVHETTE 156
++ LP IEF YN HS+ +PFE +YG+ TPL + +L E
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDLLPMPNTYLLKHKDGKAKAE 370
Query: 157 KVRMIREKMK 166
V+ + EK+K
Sbjct: 371 FVKRLHEKIK 400
>AV427422
Length = 417
Score = 85.5 bits (210), Expect = 1e-17
Identities = 42/87 (48%), Positives = 55/87 (62%)
Frame = +1
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQ 95
P SIVSDRDP F S FWK L G+KL++S+AYHP++DGQ+E + LE LR + +Q
Sbjct: 157 PLSIVSDRDPLFMSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQ 336
Query: 96 GGASDSHLPLIEFTYNNSYHSSIGMAP 122
+ +P E+ YN SYH S G P
Sbjct: 337 PKSWAHWVPWAEYWYNTSYHVSTGQTP 417
>BP055130
Length = 567
Score = 61.6 bits (148), Expect(2) = 9e-12
Identities = 32/59 (54%), Positives = 38/59 (64%)
Frame = +2
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLE 94
P +IVSDRD FTS FWK G+ L +SS+YHPQTDGQ+E + LE LR V E
Sbjct: 311 PRAIVSDRDKAFTSAFWKHFFKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHE 487
Score = 24.3 bits (51), Expect(2) = 9e-12
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = +3
Query: 103 LPLIEFTYNNSYHSSIG 119
LP E+ YN+S+ +SIG
Sbjct: 516 LPWAEYCYNSSFQTSIG 566
>BP046863
Length = 580
Score = 59.7 bits (143), Expect = 7e-10
Identities = 35/103 (33%), Positives = 61/103 (58%), Gaps = 1/103 (0%)
Frame = +1
Query: 150 LVHETTEKVRMIREKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRAL-KSR 208
L+ E + +R + +Q + ++ +K R+ +++Q G+ VFL++ P A K++
Sbjct: 274 LIEERDALLLELRGNLLKAQDQMRAQANKHRRYVDYQVGNWVFLKLQPYKLQNLAQRKNQ 453
Query: 209 KLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRK 251
KL+P+F GP+++ ERV VAY + L S +H VFH+S L K
Sbjct: 454 KLSPRFYGPFKVLERVVQVAY*LDLXSE-SRVHPVFHLSLLEK 579
>BP085845
Length = 464
Score = 58.2 bits (139), Expect = 2e-09
Identities = 35/106 (33%), Positives = 57/106 (53%)
Frame = -2
Query: 206 KSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYVADPSHVIPRDDV 265
K RK K IGP++I E V +AY + +L H V HV+ L + + D S I + V
Sbjct: 421 KKRKTESKIIGPFKILEMVCLIAY*LTHSLYLLAAHIVLHVTLLWRNLYDQSQNICHEGV 242
Query: 266 QVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWE 311
+ ++ + P+ + + KV+ +R K I V+V+W G +G +WE
Sbjct: 241 XLGEHWSHMEHPIVMVDMKVRCMRPKNIDDVKVIWRGLSG*EKSWE 104
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 50.8 bits (120), Expect = 3e-07
Identities = 28/83 (33%), Positives = 45/83 (53%)
Frame = +2
Query: 46 RFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPL 105
+FTS+ + D +G ++R SS HPQT+GQ+E + + ++ + E G LP+
Sbjct: 20 QFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWIDELPI 199
Query: 106 IEFTYNNSYHSSIGMAPFEALYG 128
+ ++YN SSI PF YG
Sbjct: 200 VLWSYNTMPQSSIKETPF*LTYG 268
>BP033788
Length = 518
Score = 29.3 bits (64), Expect = 0.95
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +3
Query: 53 KSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQ 95
KSLQ G++++L + P+ D ERT+Q D ++ V ++
Sbjct: 75 KSLQTKSGARIQLIPQHLPEGDDSKERTVQVTGDKRQIEVAQE 203
>BP040614
Length = 504
Score = 28.1 bits (61), Expect = 2.1
Identities = 22/65 (33%), Positives = 30/65 (45%), Gaps = 4/65 (6%)
Frame = +1
Query: 234 PPHLSNLHDVFHVSQLR----KYVADPSHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLR 289
PP+ L D+FH+S+ R KY +I + QVRD L R+D R L
Sbjct: 4 PPNY*GLRDLFHISKDRADSIKYEVF-VQMIEIYNEQVRDLLVSNGSNRRLDIRNNSQLN 180
Query: 290 GKEIP 294
G +P
Sbjct: 181 GLNVP 195
>BP072713
Length = 403
Score = 26.9 bits (58), Expect = 4.7
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +3
Query: 201 VGRALKSRKLTPKFIGPYQISERVGTV 227
V + LKS K P+ G Y SERVG V
Sbjct: 162 VVQTLKSLKWPPRIPGYYDYSERVGVV 242
>TC11563
Length = 470
Score = 26.9 bits (58), Expect = 4.7
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = +2
Query: 293 IPLVRVVWGGATGESLTWELESKMRESYPE 322
+P + + W GA + TWEL S +++S+P+
Sbjct: 8 VPQLLIQWEGAA--NCTWELLSYIQDSFPQ 91
>TC12763 similar to UP|CADH_MEDSA (P31656) Cinnamyl-alcohol dehydrogenase
(CAD) , partial (62%)
Length = 809
Score = 26.6 bits (57), Expect = 6.1
Identities = 10/46 (21%), Positives = 23/46 (49%)
Frame = -2
Query: 132 RTPLCWFESGESVVLGPDLVHETTEKVRMIREKMKASQSRQKSYHD 177
+ LCWF ++++ PD + +R+ A+ +++ S H+
Sbjct: 268 KATLCWFSIYINIIVIPDFLFAVLLNIRLACRAFFAASNKEASSHN 131
>BP084694
Length = 521
Score = 26.2 bits (56), Expect = 8.0
Identities = 16/43 (37%), Positives = 18/43 (41%), Gaps = 9/43 (20%)
Frame = +1
Query: 102 HLPLIEFTYNNSYHSSIG---------MAPFEALYGWRCRTPL 135
HL I F N H S+ +P EAL WRC T L
Sbjct: 250 HLKSISFYTINQQHVSLSP*SSVYSIYQSPNEALMNWRCFTKL 378
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,958,398
Number of Sequences: 28460
Number of extensions: 62927
Number of successful extensions: 333
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 327
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 329
length of query: 325
length of database: 4,897,600
effective HSP length: 90
effective length of query: 235
effective length of database: 2,336,200
effective search space: 549007000
effective search space used: 549007000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC137081.4


