

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.11 - phase: 2 /pseudo
(192 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
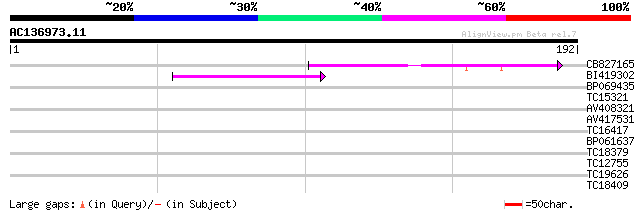
Score E
Sequences producing significant alignments: (bits) Value
CB827165 44 2e-05
BI419302 39 8e-04
BP069435 28 1.0
TC15321 similar to UP|BAD09664 (BAD09664) BHLH protein family-li... 27 2.3
AV408321 27 2.3
AV417531 26 3.9
TC16417 homologue to UP|AAR83884 (AAR83884) Ly200 protein, complete 26 3.9
BP061637 26 3.9
TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycop... 26 5.1
TC12755 similar to UP|Q9C587 (Q9C587) Replication factor C large... 25 6.6
TC19626 similar to UP|Q7XEQ1 (Q7XEQ1) Contains similarity to chr... 25 8.6
TC18409 weakly similar to UP|Q944A5 (Q944A5) AT3g01590/F4P13_13,... 25 8.6
>CB827165
Length = 550
Score = 43.9 bits (102), Expect = 2e-05
Identities = 29/99 (29%), Positives = 46/99 (46%), Gaps = 13/99 (13%)
Frame = +1
Query: 102 IFYYYTEWTFTLVMIYFALGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLE-------- 153
++ + WT T V IYF LG+ +S +GC++ K G+ + + D E
Sbjct: 166 LYGF*NRWTLTSVTIYFGLGSLLSMHGCYQHHKK----ARGDKVDNVDGDAEKGMHNAPA 333
Query: 154 -TKGSIFTFQSRY----AEEEFEQTAGFWGYLMQITFQV 187
GS + Q + ++ Q AG WGY+ QI FQ+
Sbjct: 334 IPPGSNASDQEKNLKAPEKDPLRQLAGTWGYIFQIIFQI 450
>BI419302
Length = 539
Score = 38.5 bits (88), Expect = 8e-04
Identities = 18/52 (34%), Positives = 29/52 (55%)
Frame = +2
Query: 56 GHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYT 107
G + +LW +C + +HP+ LL R+ SF+ L LL ++ I Y+YT
Sbjct: 383 GLLYEDELWNTCLKRIHPVWLLAYRIVSFIVLLGLLIGNMVADGVGILYFYT 538
>BP069435
Length = 423
Score = 28.1 bits (61), Expect = 1.0
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +2
Query: 89 MLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+L + +E DA I YYT +F L ++YF L
Sbjct: 101 LLRLVQAYEIDALIVKYYTFLSFELYLLYFLL 196
>TC15321 similar to UP|BAD09664 (BAD09664) BHLH protein family-like, partial
(23%)
Length = 1593
Score = 26.9 bits (58), Expect = 2.3
Identities = 22/62 (35%), Positives = 27/62 (43%), Gaps = 5/62 (8%)
Frame = +3
Query: 61 SQLWTSCWRGVHPLVLL--TTRLFSFVSLAMLLYLD---IHEYDASIFYYYTEWTFTLVM 115
S T C RG++PL LL RLF + LL L IH+Y I W L
Sbjct: 1296 STFLTCCRRGIYPLWLLYILPRLFVLIH**RLLKLAKP*IHDYFGDIEISAVGWACHLFC 1475
Query: 116 IY 117
+Y
Sbjct: 1476 LY 1481
>AV408321
Length = 348
Score = 26.9 bits (58), Expect = 2.3
Identities = 18/55 (32%), Positives = 26/55 (46%)
Frame = +1
Query: 41 LLVANSPSSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
LLV S +SDN +++ VS Q R + V L+ F + A +YL I
Sbjct: 163 LLVEFSKASDNTLSVSTVSDDQTLVQLGRPMLMAVSLSNSFFDDLPKAWFIYLFI 327
>AV417531
Length = 300
Score = 26.2 bits (56), Expect = 3.9
Identities = 12/47 (25%), Positives = 22/47 (46%), Gaps = 8/47 (17%)
Frame = +3
Query: 20 VLWTNEGSSTSQSDSNIFVESL--------LVANSPSSDNRVAIGHV 58
+LW NEG+ + +F+ ++ L N P + R +GH+
Sbjct: 87 ILWANEGNCDGEESVTLFLRAIGLREYSRYLCFNFPFTHERPLLGHL 227
>TC16417 homologue to UP|AAR83884 (AAR83884) Ly200 protein, complete
Length = 719
Score = 26.2 bits (56), Expect = 3.9
Identities = 20/69 (28%), Positives = 25/69 (35%), Gaps = 4/69 (5%)
Frame = -2
Query: 20 VLWTNEGSSTSQSDSNIFVESLLVA----NSPSSDNRVAIGHVSTSQLWTSCWRGVHPLV 75
V+ ST+ + N E LV N + AI H W CW HP V
Sbjct: 577 VIRAKSSRSTNHNPKNTNFEKDLVPCPYPN*EKQNTATAITHPLGCATWQVCW*QQHPTV 398
Query: 76 LLTTRLFSF 84
L + SF
Sbjct: 397 LHALQYPSF 371
>BP061637
Length = 492
Score = 26.2 bits (56), Expect = 3.9
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -2
Query: 63 LWTSCWRGVHPLVLLTTRLFSFVSLAML 90
++ SC VHPLVL T LF L+M+
Sbjct: 92 MYLSCSVCVHPLVLKTQSLFYIYCLSMV 9
>TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycoprotein
DZ-HRGP precursor, partial (8%)
Length = 584
Score = 25.8 bits (55), Expect = 5.1
Identities = 16/66 (24%), Positives = 30/66 (45%)
Frame = +3
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
WY +++ +S + VL T+ G++T+ + S + + +G +T
Sbjct: 183 WYGIFTISLIAIS---STPVLSTSNGATTTTTTSTSTTRTTTI-----------LGSTTT 320
Query: 61 SQLWTS 66
S LWTS
Sbjct: 321 SSLWTS 338
>TC12755 similar to UP|Q9C587 (Q9C587) Replication factor C large
subunit-like protein, partial (3%)
Length = 477
Score = 25.4 bits (54), Expect = 6.6
Identities = 13/41 (31%), Positives = 24/41 (57%), Gaps = 1/41 (2%)
Frame = -3
Query: 55 IGHVST-SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLD 94
+GH T S L CWRG+ L+L + + +SL ++ +++
Sbjct: 463 LGHPPTRSTLQFLCWRGISYLLLTSHQDTVIISLILISFVN 341
>TC19626 similar to UP|Q7XEQ1 (Q7XEQ1) Contains similarity to chromaffin
granule ATPase II homolog, partial (15%)
Length = 576
Score = 25.0 bits (53), Expect = 8.6
Identities = 11/31 (35%), Positives = 15/31 (47%)
Frame = +3
Query: 95 IHEYDASIFYYYTEWTFTLVMIYFALGTTVS 125
I Y I+++Y TFTL +F T S
Sbjct: 63 ISVYAVVIYFFYKNLTFTLTQFWFTFQTGFS 155
>TC18409 weakly similar to UP|Q944A5 (Q944A5) AT3g01590/F4P13_13, partial
(51%)
Length = 609
Score = 25.0 bits (53), Expect = 8.6
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = -2
Query: 24 NEGSSTSQSDSNIFVESLLVANSPSSDNR 52
+ G + +D N+F+ + +N PSS NR
Sbjct: 431 DRGEGSLSNDHNLFLANPWCSNKPSSPNR 345
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,960,198
Number of Sequences: 28460
Number of extensions: 52317
Number of successful extensions: 450
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 447
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 449
length of query: 192
length of database: 4,897,600
effective HSP length: 85
effective length of query: 107
effective length of database: 2,478,500
effective search space: 265199500
effective search space used: 265199500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC136973.11


