

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.10 + phase: 0
(330 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
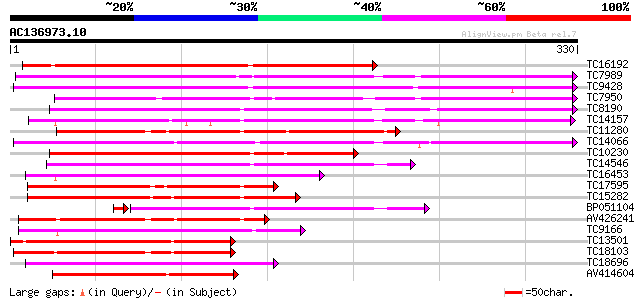
Score E
Sequences producing significant alignments: (bits) Value
TC16192 similar to UP|Q9SSZ9 (Q9SSZ9) Peroxidase 1, partial (49%) 238 1e-63
TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , par... 236 3e-63
TC9428 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), pa... 234 2e-62
TC7950 similar to UP|Q9XFI8 (Q9XFI8) Peroxidase (Fragment), part... 223 3e-59
TC8190 similar to UP|PER1_ARAHY (P22195) Cationic peroxidase 1 p... 219 4e-58
TC14157 similar to UP|O24080 (O24080) Peroxidase2 precursor , p... 217 2e-57
TC11280 similar to UP|Q9XFL5 (Q9XFL5) Peroxidase 4 (Fragment), p... 189 4e-49
TC14066 homologue to UP|O64970 (O64970) Cationic peroxidase 2 ,... 183 3e-47
TC10230 similar to UP|PE10_ARATH (Q9FX85) Peroxidase 10 precurso... 154 2e-38
TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxida... 151 2e-37
TC16453 similar to UP|Q40372 (Q40372) Peroxidase precursor, part... 140 2e-34
TC17595 similar to UP|Q9XFI7 (Q9XFI7) Peroxidase (Fragment), par... 137 2e-33
TC15282 similar to UP|Q9ZNZ6 (Q9ZNZ6) Peroxidase precursor (Fra... 133 4e-32
BP051104 129 6e-31
AV426241 126 4e-30
TC9166 similar to UP|Q9XFL4 (Q9XFL4) Peroxidase 3, partial (49%) 126 6e-30
TC13501 similar to UP|PE47_ARATH (Q9SZB9) Peroxidase 47 precurso... 125 7e-30
TC18103 similar to UP|PER2_ARAHY (P22196) Cationic peroxidase 2 ... 125 1e-29
TC18696 similar to UP|Q40372 (Q40372) Peroxidase precursor, part... 122 8e-29
AV414604 122 8e-29
>TC16192 similar to UP|Q9SSZ9 (Q9SSZ9) Peroxidase 1, partial (49%)
Length = 645
Score = 238 bits (607), Expect = 1e-63
Identities = 117/207 (56%), Positives = 151/207 (72%)
Frame = +2
Query: 8 ATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFH 67
A +V+VI ++ S L+ G+Y +C E IV+ V ++V+ NPGIAAGL+RMHFH
Sbjct: 32 AIIVLVIFLLNQNAYSE--LEVGYYSYSCGMAEFIVKDEVRRSVTKNPGIAAGLVRMHFH 205
Query: 68 DCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILA 127
DCF+RGCD SVLLDS P +E+D PAN PSLRGFEVI+ AKA++EA C VSCADI+A
Sbjct: 206 DCFIRGCDASVLLDSTPSNTAEKDSPANKPSLRGFEVIDSAKAKLEAVCKGVVSCADIIA 385
Query: 128 FAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVD 187
FAARDS G + Y VP+GRRDGR+S+ + +LPPPTF+ QL F +KGL+ +
Sbjct: 386 FAARDSVELAGG--VGYDVPAGRRDGRISLASDTRTDLPPPTFNVNQLTQLFAKKGLTQE 559
Query: 188 EMVTLSGAHSIGVSHCSSFSKRLYSFN 214
EMVTLSGAH+IG SHC++FS RLY+F+
Sbjct: 560 EMVTLSGAHTIGRSHCAAFSSRLYNFS 640
>TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , partial (85%)
Length = 1352
Score = 236 bits (603), Expect = 3e-63
Identities = 140/329 (42%), Positives = 193/329 (58%), Gaps = 2/329 (0%)
Frame = +3
Query: 4 ILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIR 63
I S + + +S + T L + FY ++C +E+I+R+ + K + AAGL+R
Sbjct: 69 ISSFLSFSHIRVSEAATAPIVKGLSWTFYDSSCPKLESIIRKELKKVFDKDIAQAAGLLR 248
Query: 64 MHFHDCFVRGCDGSVLLDSIPGIQSERDHPAN-NPSLRGFEVINEAKAQIEAACPKTVSC 122
+HFHDCFV+GCD SVLLD SE+D P N + F++I + ++QIE C + VSC
Sbjct: 249 LHFHDCFVQGCDASVLLDGSASGPSEKDAPPNLTLRAQAFKIIEDLRSQIEKKCGRVVSC 428
Query: 123 ADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVT-QNLPPPTFSAEQLIDNFDR 181
ADI A AARD A +SGG DY +P GRRDG VT NLP P + ++++
Sbjct: 429 ADITAIAARD-AVFLSGGP-DYELPLGRRDGLNFATRNVTLDNLPAPQSNTTTILNSLAT 602
Query: 182 KGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQ 241
K L ++V+LSG H+IG+SHCSSF+ RLY +DP MD F + LK CP +
Sbjct: 603 KNLDPTDVVSLSGGHTIGISHCSSFTDRLYP-----SKDPVMDQTFEKNLKLTCP---AS 758
Query: 242 SINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVK 301
+ + T VLD +PN DN YY L N +GL SDQ L T+ +V A + +++ K
Sbjct: 759 NTDNTTVLDLRSPNTFDNKYYVDLMNRQGLFFSDQDLYTDKRTKDIVTSFAVNQSLFFEK 938
Query: 302 FAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F AM+ MG L+VLTGS+GEIR CSV N
Sbjct: 939 FVVAMLKMGQLNVLTGSQGEIRANCSVRN 1025
>TC9428 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), partial
(97%)
Length = 1354
Score = 234 bits (596), Expect = 2e-62
Identities = 127/330 (38%), Positives = 188/330 (56%), Gaps = 2/330 (0%)
Frame = +2
Query: 3 SILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLI 62
S + +A + V++ +S L FYK TC +V +IVR V +P + LI
Sbjct: 44 SPIHLAVVFCVVIGALVPYSSDAQLDNSFYKDTCPNVHSIVREVVRNVSKSDPRMLGSLI 223
Query: 63 RMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSC 122
R+HFHDCFV+GCD S+LL+ I SE+ NN S+RG +V+N+ K +E ACP TVSC
Sbjct: 224 RLHFHDCFVQGCDASILLNDTSTIVSEQGAFPNNNSIRGLDVVNQIKTAVENACPNTVSC 403
Query: 123 ADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRK 182
ADILA AA S+ +G +D+ VP GRRD + QNLP P F+ QL +F +
Sbjct: 404 ADILALAAEISSVLANG--VDWKVPLGRRDSLTANRSLANQNLPGPNFNLSQLNASFAVQ 577
Query: 183 GLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQS 242
L++ ++V LSGAH+IG C F RLY+F+ T DP+++ + + L+S C P
Sbjct: 578 NLNITDLVALSGAHTIGRGQCQFFVNRLYNFSNTGNPDPTLNTTYLQTLRSIC--PHGGP 751
Query: 243 INPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKN--ARHAAIWNV 300
LD +TP+ LD+ Y+ L+ +GL SD L ++ + + N + + ++
Sbjct: 752 GTTLAHLDPTTPDTLDSQYFSNLQGGKGLFQSDPVLFSTSGAATVAIVNSFSSNPTLFFE 931
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F +M+ M + VLTGS+GEIR+ C+ VN
Sbjct: 932 NFKASMIKMSRIGVLTGSQGEIRKPCNFVN 1021
>TC7950 similar to UP|Q9XFI8 (Q9XFI8) Peroxidase (Fragment), partial (86%)
Length = 1286
Score = 223 bits (569), Expect = 3e-59
Identities = 130/305 (42%), Positives = 173/305 (56%), Gaps = 1/305 (0%)
Frame = +1
Query: 27 LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSIPGI 86
L + FY TC +E +VR + K + + G A GL+R+ FHDCFV+GCDGSVLLD PG
Sbjct: 106 LSFSFYSKTCPKLETVVRNHLKKVLKKDNGQAPGLLRIFFHDCFVQGCDGSVLLDGSPG- 282
Query: 87 QSERDHPAN-NPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRIDYS 145
ERD PAN + I + +A + C K VSCADI A+RD+ G DY+
Sbjct: 283 --ERDQPANIGIRPEALQTIEDIRALVHKQCGKIVSCADITILASRDAVFLTGGP--DYA 450
Query: 146 VPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSHCSS 205
VP GRRDG VS TQ LP P + + F + ++V LSGAH+ G +HC +
Sbjct: 451 VPLGRRDG-VSFSTVGTQKLPSPINNTTATLKAFADRNFDATDVVALSGAHTFGRAHCGT 627
Query: 206 FSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVLDGSTPNDLDNMYYKRL 265
F FN P DP+MD A+ L + CP +Q+ T LD TPN DN YY L
Sbjct: 628 F------FNRLSPLDPNMDKTLAKNLTATCP---AQNSTNTANLDIRTPNVFDNKYYLDL 780
Query: 266 KNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEGEIRER 325
N +G+ TSDQ LL+ T+ +V A + ++ KF A++ + LDVLTG++GEIR R
Sbjct: 781 MNRQGVFTSDQDLLSDKRTKGLVNAFAVNQTLFFEKFVDAVIKLSQLDVLTGNQGEIRGR 960
Query: 326 CSVVN 330
C+VVN
Sbjct: 961 CNVVN 975
>TC8190 similar to UP|PER1_ARAHY (P22195) Cationic peroxidase 1 precursor
(PNPC1) , partial (96%)
Length = 1202
Score = 219 bits (559), Expect = 4e-58
Identities = 124/308 (40%), Positives = 170/308 (54%), Gaps = 1/308 (0%)
Frame = +1
Query: 24 STSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSI 83
S L FY TC V A ++ VN AV+ + A L+R+HFHDCFV+GCD S+LLD
Sbjct: 133 SAQLSSTFYAKTCPLVLATIKTQVNLAVAKEARMGASLLRLHFHDCFVQGCDASILLDDT 312
Query: 84 PGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRID 143
E+ N S+RG++VI+ K+++E+ CP VSCADI+A AARDS V+ G
Sbjct: 313 SSFTGEKTAGPNANSVRGYDVIDTIKSKVESLCPGVVSCADIVAVAARDSV--VALGGFS 486
Query: 144 YSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSHC 203
++VP GRRD + LP P+ + + L F KG + EMV LSG+H+IG + C
Sbjct: 487 WAVPLGRRDSTTASLSSANSELPGPSSNLDGLNTAFSNKGFTTREMVALSGSHTIGQARC 666
Query: 204 SSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQS-INPTVVLDGSTPNDLDNMYY 262
F R+Y + +D FA+ L+ CP S ++P LD ++P DN YY
Sbjct: 667 LFFRTRIY-------YETHIDSTFAKNLQGNCPFNGGDSNLSP---LDTTSPTTFDNGYY 816
Query: 263 KRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEGEI 322
+ L++ +GL SDQ L N G T V + A + FA AMV MG+L LTGS G+I
Sbjct: 817 RNLQSQKGLFHSDQVLFNGGSTDSQVNSYVTNPASFKTDFANAMVKMGNLSPLTGSSGQI 996
Query: 323 RERCSVVN 330
R C N
Sbjct: 997 RTHCRKTN 1020
>TC14157 similar to UP|O24080 (O24080) Peroxidase2 precursor , partial
(96%)
Length = 1390
Score = 217 bits (553), Expect = 2e-57
Identities = 129/327 (39%), Positives = 188/327 (57%), Gaps = 9/327 (2%)
Frame = +1
Query: 12 IVILSVSTTLASST---SLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHD 68
+++LS+S TL T L Y +TC ++++IV+ V K +R+ FHD
Sbjct: 73 VLVLSLSLTLCFYTCFAQLSPNHYASTCPNLQSIVKGVVQKKFQQTFVTVPATLRLFFHD 252
Query: 69 CFVRGCDGSVLLDSIPGIQSERDHPANNPSLRG--FEVINEAKAQIEAA--CPKTVSCAD 124
CFV+GCD SV++ S ++E+DHP +N SL G F+ + +AKA ++A C VSCAD
Sbjct: 253 CFVQGCDASVMVASSGNNKAEKDHP-DNLSLAGDGFDTVIKAKAAVDAVPQCRNKVSCAD 429
Query: 125 ILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGL 184
ILA A RD V G Y+V GR DG VS +V LP P F+ QL F +GL
Sbjct: 430 ILALATRDVV--VLAGGPSYTVELGRFDGLVSRASDVNGRLPEPNFNLNQLNSLFASQGL 603
Query: 185 SVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSIN 244
+ +M+ LSGAH++G SHC+ FS R+YS P DP+++ N+A L+ CP +++N
Sbjct: 604 TQTDMIALSGAHTLGFSHCNRFSNRIYS----TPVDPTLNRNYATQLQQMCP----KNVN 759
Query: 245 PTVV--LDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKF 302
P + +D +TP DN+YYK L+ +GL TSDQ L ++ V A ++ +N F
Sbjct: 760 PQIAINMDPTTPRTFDNIYYKNLQQGKGLFTSDQILFTDQRSKATVNSFASNSNTFNANF 939
Query: 303 AKAMVHMGSLDVLTGSEGEIRERCSVV 329
A AM+ +G + V T G+IR CSV+
Sbjct: 940 AAAMIKLGRVGVKTARNGKIRTDCSVL 1020
>TC11280 similar to UP|Q9XFL5 (Q9XFL5) Peroxidase 4 (Fragment), partial
(68%)
Length = 614
Score = 189 bits (481), Expect = 4e-49
Identities = 107/200 (53%), Positives = 132/200 (65%)
Frame = +3
Query: 28 KYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSIPGIQ 87
+ GFY TC E+IVR V V+ +P +AAGL+RMHFHDCFV+GCD SVL I G
Sbjct: 27 RVGFYLGTCPRAESIVRSTVESHVNSDPTLAAGLLRMHFHDCFVQGCDASVL---IAGAG 197
Query: 88 SERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRIDYSVP 147
+ER N SLRGFEVI++AKA++EAACP VSCADILA AARDS V G + + VP
Sbjct: 198 TERT-AIPNLSLRGFEVIDDAKAKVEAACPGVVSCADILALAARDSV--VLSGGLSWQVP 368
Query: 148 SGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSHCSSFS 207
+GRRDGRVS +V NLP P S + F KGL+ ++VTL G H+IG + C FS
Sbjct: 369 TGRRDGRVSQASDV-NNLPAPFDSVDVQKQKFTAKGLNTQDLVTLVGGHTIGTTACQFFS 545
Query: 208 KRLYSFNLTFPQDPSMDPNF 227
RLY+F P DPS+D +F
Sbjct: 546 NRLYNFTSNGP-DPSIDASF 602
>TC14066 homologue to UP|O64970 (O64970) Cationic peroxidase 2 , partial
(92%)
Length = 1436
Score = 183 bits (465), Expect = 3e-47
Identities = 117/330 (35%), Positives = 169/330 (50%), Gaps = 2/330 (0%)
Frame = +2
Query: 3 SILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLI 62
S+LS++ L ++ S + T L FYK TC E I+ V + A +
Sbjct: 80 SLLSLSALSLLSPSSAQEDVQDTGLAMNFYKETCPQAEDIITEQVKLLYKRHKNTAFSWL 259
Query: 63 RMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSC 122
R FHDC V+ CD S+LLDS SE++ + LR F I K +E CP VSC
Sbjct: 260 RNIFHDCAVQSCDASLLLDSTRKTLSEKETDRSF-GLRNFRYIETIKEALERECPGVVSC 436
Query: 123 ADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRK 182
+DIL +ARD + G I + +GRRDGR S D V Q LP S ++D F
Sbjct: 437 SDILVLSARDGIAALGGPHIP--LRTGRRDGRRSRADVVEQFLPDHNESISAVLDKFGAM 610
Query: 183 GLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPP--PQS 240
G+ +V L GAHS+G +HC RLY DP+++P+ + KCP P
Sbjct: 611 GIDTPGVVALLGAHSVGRTHCVKLVHRLYP-----EVDPALNPDHIPHMLKKCPDAIPDP 775
Query: 241 QSINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNV 300
+++ V D TP LDN YY+ + +N+GLL D L N T+ V K A+ +
Sbjct: 776 KAVQ-YVRNDRGTPMILDNNYYRNILDNKGLLLVDHQLANDKRTKPYVKKMAKSQDYFFK 952
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
+F++A+ + + LTG++GEIR++C+V N
Sbjct: 953 EFSRAITLLSENNPLTGTKGEIRKQCNVAN 1042
>TC10230 similar to UP|PE10_ARATH (Q9FX85) Peroxidase 10 precursor (Atperox
P10) (ATP5a) , partial (42%)
Length = 753
Score = 154 bits (389), Expect = 2e-38
Identities = 85/180 (47%), Positives = 109/180 (60%)
Frame = +3
Query: 24 STSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSI 83
S+ L Y FY TTC ++ IVR + A+S + IAA L+R+HFHDCFV GCDGSVLLD
Sbjct: 216 SSQLYYNFYDTTCPNLTRIVRYNLLSAMSNDTRIAASLLRLHFHDCFVNGCDGSVLLDDT 395
Query: 84 PGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRID 143
+ E++ N S+RGFEVI+ K+ +E ACP TVSCADIL AAR++ G
Sbjct: 396 STQKGEKNALPNKNSIRGFEVIDTIKSALEKACPSTVSCADILTLAAREAVYLSRGP--F 569
Query: 144 YSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSHC 203
+SVP GRRDG + E NLP P E + F KGL ++ LSGAH+ G + C
Sbjct: 570 WSVPLGRRDGTTASESE-ANNLPSPFEPLENITAKFISKGLEKKDVAVLSGAHTFGFAQC 746
>TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (66%)
Length = 738
Score = 151 bits (381), Expect = 2e-37
Identities = 82/216 (37%), Positives = 117/216 (53%), Gaps = 1/216 (0%)
Frame = +1
Query: 22 ASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLD 81
+++ L FY +C ++ IVR A+ + + + A ++R+ FHDCFV GCDGS+LLD
Sbjct: 85 SANAQLTPTFYDRSCPKLQTIVRNAMVQTIKKEARMGASILRLFFHDCFVNGCDGSILLD 264
Query: 82 SI-PGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGG 140
I E++ N S RGFEVI+ K +EA+C TVSCADILA A RD + G
Sbjct: 265 DIGTTFVGEKNAAPNKNSARGFEVIDTIKTNVEASCNNTVSCADILALATRDGINLLGGP 444
Query: 141 RIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGV 200
+ VP GRRD R + + +P P+ LI F KGLS ++ LSG H+IG
Sbjct: 445 --TWQVPLGRRDARTASQSKANTEIPSPSSDLSTLISMFSAKGLSARDLTVLSGGHTIGQ 618
Query: 201 SHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCP 236
+ C F R+ + + ++D FA K+ CP
Sbjct: 619 AECQFFRSRVNN-------ETNIDAAFAASRKTNCP 705
>TC16453 similar to UP|Q40372 (Q40372) Peroxidase precursor, partial (49%)
Length = 581
Score = 140 bits (354), Expect = 2e-34
Identities = 77/177 (43%), Positives = 104/177 (58%), Gaps = 3/177 (1%)
Frame = +2
Query: 10 LVIVILSVSTTLASST--SLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFH 67
LV+V+++++T L ST L G+Y C I++ V KA++ I A L+R+HFH
Sbjct: 50 LVLVLVTLATFLIPSTLAGLTPGYYDKVCPQALPIIKSVVQKAINREQRIGASLLRLHFH 229
Query: 68 DCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPK-TVSCADIL 126
DCFV GCDGS+LLD E+ N S+RGFEV++E KA ++ AC + VSCADIL
Sbjct: 230 DCFVNGCDGSLLLDDTSSFVGEKTALPNLNSVRGFEVVDEIKAAVDRACKRPVVSCADIL 409
Query: 127 AFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKG 183
A AARDS + G + Y V GRRD R + NLPPP + QL+ F +G
Sbjct: 410 AIAARDSVAILGGKQYWYQVRLGRRDARSASQAAANSNLPPPFLNFTQLLPAFQAQG 580
>TC17595 similar to UP|Q9XFI7 (Q9XFI7) Peroxidase (Fragment), partial (42%)
Length = 657
Score = 137 bits (346), Expect = 2e-33
Identities = 75/146 (51%), Positives = 98/146 (66%)
Frame = +1
Query: 11 VIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCF 70
V+V+L ++S L+ GFY TC E+IV+ V AV+ +P +AA L+R+HFHDCF
Sbjct: 139 VLVLLFSFLIVSSDGQLQVGFYSNTCPHAESIVQAVVRGAVASDPNMAAVLLRLHFHDCF 318
Query: 71 VRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAA 130
V GCDGS+L+++ G QSE+ + +RGFEVI AKAQ EA+CP VSCADI+A AA
Sbjct: 319 VEGCDGSILIEN--GDQSEK-LAFGHQGVRGFEVIERAKAQSEASCPDVVSCADIVALAA 489
Query: 131 RDSARKVSGGRIDYSVPSGRRDGRVS 156
DS V G +Y VP+GRRDG VS
Sbjct: 490 TDSI--VMGNGPEYQVPTGRRDGSVS 561
>TC15282 similar to UP|Q9ZNZ6 (Q9ZNZ6) Peroxidase precursor (Fragment) ,
partial (47%)
Length = 604
Score = 133 bits (335), Expect = 4e-32
Identities = 72/159 (45%), Positives = 101/159 (63%)
Frame = +1
Query: 11 VIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCF 70
V+++ ++ ++ L+ GFY TC E I+ V++ + P +AA LIRM+FHDCF
Sbjct: 115 VLIVCLLALIASNHAQLEQGFYTKTCPKAEKIILDFVHEHIHNAPSLAAALIRMNFHDCF 294
Query: 71 VRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAA 130
VRGCD S+LL+S Q+ERD P N ++RGF+ I+ K+ +EA CP VSCADI+A AA
Sbjct: 295 VRGCDASILLNS-TSKQAERDAPP-NLTVRGFDFIDRIKSLVEAQCPGVVSCADIIALAA 468
Query: 131 RDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPT 169
RDS V+ G + VP+GRRDG +S E +P PT
Sbjct: 469 RDSI--VATGGPFWKVPTGRRDGVISNLVEARSQIPAPT 579
>BP051104
Length = 566
Score = 129 bits (323), Expect(2) = 6e-31
Identities = 70/174 (40%), Positives = 97/174 (55%)
Frame = +2
Query: 71 VRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAA 130
V GCD S+L D E+ ANN S RGF VI+ KA +E CP VSCAD+LA AA
Sbjct: 71 VNGCDASILXDDTSNFIGEQTAAANNRSARGFNVIDGIKANLEKQCPGVVSCADVLALAA 250
Query: 131 RDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMV 190
RDS ++ G + V GRRD + +P P S LI NF +GLSV ++V
Sbjct: 251 RDSVVQLGGP--SWEVGLGRRDSTTASRGTANNTIPGPFLSLSGLITNFANQGLSVTDLV 424
Query: 191 TLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSIN 244
LSGAH+IG++ C +F +Y+ D ++D ++A+ LK CP + ++N
Sbjct: 425 ALSGAHTIGLAQCKNFRAHIYN-------DSNIDASYAKFLKX*CPRSGNDNLN 565
Score = 21.6 bits (44), Expect(2) = 6e-31
Identities = 7/9 (77%), Positives = 7/9 (77%)
Frame = +3
Query: 61 LIRMHFHDC 69
L MHFHDC
Sbjct: 42 LTSMHFHDC 68
>AV426241
Length = 429
Score = 126 bits (317), Expect = 4e-30
Identities = 70/146 (47%), Positives = 95/146 (64%)
Frame = +3
Query: 6 SIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMH 65
S + +++L++ TS+ GFY +C S+E+IV+ V V + AAGL+R+H
Sbjct: 15 SFLLVFLIVLTLQAFAVHGTSV--GFYSKSCPSIESIVKSTVASHVKTDFEYAAGLLRLH 188
Query: 66 FHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADI 125
F DCFVRGCD S+L I G +E+ P N SL+G+EVI+EAKA++EA CP VSCADI
Sbjct: 189 FRDCFVRGCDASIL---IAGNGTEKQAPPNR-SLKGYEVIDEAKAKLEAQCPGVVSCADI 356
Query: 126 LAFAARDSARKVSGGRIDYSVPSGRR 151
LA AARDS V G + + VP+GRR
Sbjct: 357 LALAARDSV--VLSGGLSWQVPTGRR 428
>TC9166 similar to UP|Q9XFL4 (Q9XFL4) Peroxidase 3, partial (49%)
Length = 559
Score = 126 bits (316), Expect = 6e-30
Identities = 69/169 (40%), Positives = 97/169 (56%), Gaps = 2/169 (1%)
Frame = +3
Query: 6 SIATLVIVILSVSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIR 63
S A + ++ L + + SS S L FY C SV V+ V+ AV+ + L+R
Sbjct: 42 SAANIFVLSLFMLFLIGSSNSAQLSENFYVKKCPSVFNAVKSVVHSAVAKEARMGGSLLR 221
Query: 64 MHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCA 123
+ FHDCFV GCDGSVLLD + E+ P N+ SLRGF+VI+ K+++EA CP VSCA
Sbjct: 222 LFFHDCFVNGCDGSVLLDDTSSFKGEKTAPPNSNSLRGFDVIDAIKSKVEAVCPGVVSCA 401
Query: 124 DILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSA 172
D++A AARDS + G + V GRRD + + F+ + P FS+
Sbjct: 402 DVVAIAARDSVAILGGPY--WKVKLGRRDSKTASFNAANSGVIPSPFSS 542
>TC13501 similar to UP|PE47_ARATH (Q9SZB9) Peroxidase 47 precursor (Atperox
P47) (ATP32) , partial (33%)
Length = 519
Score = 125 bits (315), Expect = 7e-30
Identities = 62/131 (47%), Positives = 88/131 (66%)
Frame = +1
Query: 1 MHSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAG 60
M S+L++ L+I ++S + L +Y C E++V+ VN+A+ +P +AAG
Sbjct: 133 MASLLTVF-LLIEVISCGFGFGGNNGLNMNYYLMRCPFAESVVKNIVNRALQNDPTLAAG 309
Query: 61 LIRMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTV 120
LIRMHFHDCFV GCDGS+L+DS +E+D PA N SL+G+E+I+E K ++E CP V
Sbjct: 310 LIRMHFHDCFVEGCDGSILIDSTKDNTAEKDSPA-NLSLKGYEIIDEIKEELERQCPGVV 486
Query: 121 SCADILAFAAR 131
SCAD+LA AAR
Sbjct: 487 SCADVLAMAAR 519
>TC18103 similar to UP|PER2_ARAHY (P22196) Cationic peroxidase 2 precursor
(PNPC2) , partial (37%)
Length = 546
Score = 125 bits (314), Expect = 1e-29
Identities = 69/129 (53%), Positives = 88/129 (67%)
Frame = +1
Query: 3 SILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLI 62
S+ IA L++ + S+ T+ S + GFY+ TC E+IVR AV V + +AAGL+
Sbjct: 124 SLFRIAFLLLALASIVNTVHGQGS-RVGFYRRTCPRAESIVRSAVESHVKSDRTLAAGLL 300
Query: 63 RMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSC 122
RMHFHDCFV+GCD SVL I G +ER P N LRG+EVI++AKA++EAACP VSC
Sbjct: 301 RMHFHDCFVQGCDASVL---IAGAGTERTAPP-NLGLRGYEVIDDAKAKVEAACPGVVSC 468
Query: 123 ADILAFAAR 131
ADILA AAR
Sbjct: 469 ADILALAAR 495
>TC18696 similar to UP|Q40372 (Q40372) Peroxidase precursor, partial (47%)
Length = 500
Score = 122 bits (306), Expect = 8e-29
Identities = 61/148 (41%), Positives = 87/148 (58%), Gaps = 1/148 (0%)
Frame = +2
Query: 10 LVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDC 69
+V+V+++++ + S L FY C ++ V +A+ I A L+R+HFHDC
Sbjct: 38 IVLVMVTLTLVIPSKAQLSPSFYNKVCPQALPVINSVVRRAILRERRIGASLLRLHFHDC 217
Query: 70 FVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPK-TVSCADILAF 128
FV GCDGSVLLD E+ NN S++GF+V++E K ++ AC + VSCADILA
Sbjct: 218 FVNGCDGSVLLDDTRNFIGEKTAFPNNNSIKGFDVVDEIKKAVDKACKRPVVSCADILAI 397
Query: 129 AARDSARKVSGGRIDYSVPSGRRDGRVS 156
AARDS + G + Y V GRRD R +
Sbjct: 398 AARDSVAILGGPSLVYKVLLGRRDARTA 481
>AV414604
Length = 427
Score = 122 bits (306), Expect = 8e-29
Identities = 62/108 (57%), Positives = 77/108 (70%)
Frame = +2
Query: 26 SLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSIPG 85
SL+ FY+ +CS E IV+ + + VS P + A L+RMHFHDCFVRGCDGSVLL+S G
Sbjct: 83 SLRKQFYRKSCSQAEQIVKTTIQQHVSSRPELPAKLLRMHFHDCFVRGCDGSVLLNSTAG 262
Query: 86 IQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDS 133
+E+D N SL GF+VI+E K +EA CPK VSCADILA AARD+
Sbjct: 263 NTAEKD-AIPNLSLSGFDVIDEIKEALEAKCPKIVSCADILALAARDA 403
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.133 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,631,395
Number of Sequences: 28460
Number of extensions: 75328
Number of successful extensions: 517
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 444
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 444
length of query: 330
length of database: 4,897,600
effective HSP length: 91
effective length of query: 239
effective length of database: 2,307,740
effective search space: 551549860
effective search space used: 551549860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC136973.10


