

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
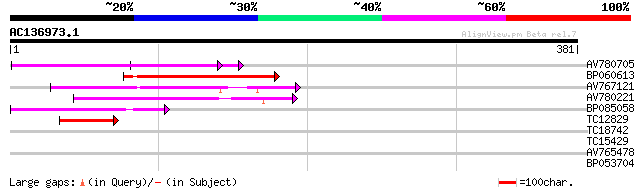
Query= AC136973.1 - phase: 0 /pseudo
(381 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV780705 105 1e-23
BP060613 91 2e-19
AV767121 80 7e-16
AV780221 50 5e-07
BP085058 46 9e-06
TC12829 44 4e-05
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 32 0.23
TC15429 similar to PIR|T00425|T00425 photolyase/blue-light recep... 30 0.87
AV765478 27 4.3
BP053704 26 9.6
>AV780705
Length = 524
Score = 105 bits (262), Expect = 1e-23
Identities = 50/143 (34%), Positives = 84/143 (57%), Gaps = 1/143 (0%)
Frame = -2
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLGLP++ GR+K + +I+++V +K+ W L++AG+EV+IK ++QAIPSY M+I
Sbjct: 484 YLGLPAIWGRNKSHSLVWIEEKVKEKLEGWKETLLNQAGKEVLIKAIIQAIPSYAMTIVH 305
Query: 62 LSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
T + + ++ FWW G K + K GG+GF+D N+A L +
Sbjct: 304 FPKTFCNSLYALVADFWWKSQGEGAVEYTGAVGTK*PLSKEKGGVGFRDLRTQNLAFLAR 125
Query: 121 QGWQLQTKPDSLVSKIFKARYYP 143
Q W++ T P++L ++ K+ Y+P
Sbjct: 124 QAWRVLTNPEALWVRVMKSLYFP 56
Score = 45.1 bits (105), Expect = 2e-05
Identities = 22/76 (28%), Positives = 35/76 (45%)
Frame = -1
Query: 82 GSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKARY 141
G +RG++W SW+K+++ GG G + + KP LV +
Sbjct: 242 GGRSRGIHWRSWDKMTLE*GKGGRGLQGLENSEPCLPS*ASLACANKPGGLVGTCHEISL 63
Query: 142 YPNSNFLDAKLGHNPS 157
+P +F+ A LG NPS
Sbjct: 62 FPYPSFMFAALGRNPS 15
>BP060613
Length = 378
Score = 91.3 bits (225), Expect = 2e-19
Identities = 40/105 (38%), Positives = 66/105 (62%)
Frame = +1
Query: 77 WWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKI 136
WW G+ RG++W W+ L+ K++GG+GFKDF N A+L KQ W++ PD+L +I
Sbjct: 46 WWSSKGT--RGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQI 219
Query: 137 FKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGAG 181
KA Y+P+ +FL S+VW S+L + ++ + +W++G+G
Sbjct: 220 LKALYFPHHDFLQTTKRTGASWVWSSLLHGRELLLKQGQWQLGSG 354
>AV767121
Length = 525
Score = 79.7 bits (195), Expect = 7e-16
Identities = 55/177 (31%), Positives = 85/177 (47%), Gaps = 9/177 (5%)
Frame = +2
Query: 28 ISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFRLSNTLLDE-IEMMNSFWWGHGGSSNR 86
+S W K +S GR +I+ VL A+P + +S F+L + + +M +F WG + N+
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 87 GLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKAR---YYP 143
+ + W + K GG+G KD FN A+LGK W+ T+PDSL ++ +A+ Y
Sbjct: 182 -IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQPDHYSC 358
Query: 144 NSNFLDAKLGHNPSFVWRSILS-----AKVVVRQGARWKIGAGFDIPIISEPWIGSG 195
S++ W ILS A G + +G G SE W+GSG
Sbjct: 359 GSSW------------WNDILSLCPEDADGWFSSGLKKLVGEGDQTKFWSEDWLGSG 493
>AV780221
Length = 461
Score = 50.4 bits (119), Expect = 5e-07
Identities = 35/157 (22%), Positives = 71/157 (44%), Gaps = 7/157 (4%)
Frame = +3
Query: 44 MIKYVLQAIPSYVMSIFRL-SNTLLDEIEMMNSFWWGHGGSSNRGLNWLSWEKLSVHKND 102
+I+ VL A+P + + F+L + + ++G + + W+ W + K++
Sbjct: 9 LIRSVLIALPLFHLPFFKLPTGGFYTNVNN**DPFFGREVEGGKKIAWVKWSTVCRPKDE 188
Query: 103 GGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRS 162
GG+G ++ FN A+LGK W++ + D L K+ +Y + + S W
Sbjct: 189 GGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKY-------KNAIPQSASSWWND 347
Query: 163 ILSAKVV------VRQGARWKIGAGFDIPIISEPWIG 193
+ S +++G +IG G ++ +E W+G
Sbjct: 348 LHSTCFEDGGGGWMQRGLCRRIGEGTEVKFWNENWLG 458
>BP085058
Length = 452
Score = 46.2 bits (108), Expect = 9e-06
Identities = 32/108 (29%), Positives = 52/108 (47%), Gaps = 1/108 (0%)
Frame = +2
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KY+ P G K F F DRV +++ + L+K R + K VL AI YVM +
Sbjct: 134 KYMSFPIHQGHV*KEDFVFTLDRVISRLAC*KAYLLNKPARLTLAKPVLSAISMYVMQLN 313
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGF 107
L ++ D+I ++ F W + G+ ++WE + K+ G+ F
Sbjct: 314 WLPQSICDQICTVVRHFVW----ENVNGIPLVAWETIQSRKSMVGLIF 445
>TC12829
Length = 448
Score = 43.9 bits (102), Expect = 4e-05
Identities = 21/40 (52%), Positives = 30/40 (74%)
Frame = +3
Query: 34 KCLSKAGREVMIKYVLQAIPSYVMSIFRLSNTLLDEIEMM 73
K L +AGREV+IK V +AIP+Y+MS F L +++ +IE M
Sbjct: 255 KFLYRAGREVLIKSVTKAIPTYIMSCFALPDSICSQIEGM 374
Score = 27.3 bits (59), Expect = 4.3
Identities = 16/64 (25%), Positives = 33/64 (51%)
Frame = +1
Query: 33 SKCLSKAGREVMIKYVLQAIPSYVMSIFRLSNTLLDEIEMMNSFWWGHGGSSNRGLNWLS 92
+ C+ + G+ + + + Q+ + +++ L +L + SF W G S R ++WL
Sbjct: 256 NSCIEREGKF*LSR*LKQSQLTSCLALLFLILFVLRLKAWLVSFIWS-GDVSRRSIHWLG 432
Query: 93 WEKL 96
W+KL
Sbjct: 433 WKKL 444
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 31.6 bits (70), Expect = 0.23
Identities = 15/47 (31%), Positives = 25/47 (52%)
Frame = -3
Query: 3 LGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVL 49
LG+P + + K + + DR K+ W+ + LS AGR +I +L
Sbjct: 154 LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLINSIL 14
>TC15429 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (17%)
Length = 717
Score = 29.6 bits (65), Expect = 0.87
Identities = 18/51 (35%), Positives = 22/51 (42%)
Frame = +3
Query: 79 GHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKP 129
G GGSSN G NWL +E L ++DF F QL+ P
Sbjct: 204 GGGGSSNTGTNWLMFELL----------WRDFFRFITKKYSSAKKQLEAAP 326
>AV765478
Length = 357
Score = 27.3 bits (59), Expect = 4.3
Identities = 13/25 (52%), Positives = 17/25 (68%), Gaps = 2/25 (8%)
Frame = +1
Query: 229 PSTFCSRNGPKHPE--YSSAPSGRE 251
PSTF R GP+H +++PSGRE
Sbjct: 274 PSTFALRAGPRHTAG*KTTSPSGRE 348
>BP053704
Length = 457
Score = 26.2 bits (56), Expect = 9.6
Identities = 15/43 (34%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Frame = -2
Query: 229 PSTFCSRNGPKHPEYSSAPSGREG*T---DLEGGEKWPLFCEK 268
P+T C KH E++ PS + *+ L E+WP C+K
Sbjct: 252 PTTLCLMRLCKHEEFNP-PSRKSE*SCNKTLRRKERWPYMCQK 127
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.339 0.148 0.518
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,881,722
Number of Sequences: 28460
Number of extensions: 118576
Number of successful extensions: 833
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 823
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 830
length of query: 381
length of database: 4,897,600
effective HSP length: 92
effective length of query: 289
effective length of database: 2,279,280
effective search space: 658711920
effective search space used: 658711920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC136973.1


