

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.18 - phase: 1 /pseudo
(305 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
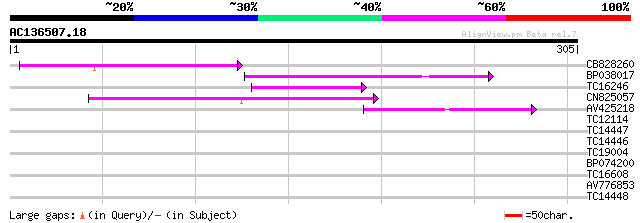
Score E
Sequences producing significant alignments: (bits) Value
CB828260 78 2e-15
BP038017 67 5e-12
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 44 3e-05
CN825057 44 4e-05
AV425218 40 5e-04
TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transp... 37 0.004
TC14447 homologue to UP|Q9M531 (Q9M531) Histone H2A, partial (86%) 28 1.9
TC14446 homologue to UP|H2A_LYCES (P25469) Histone H2A, partial ... 28 1.9
TC19004 26 7.3
BP074200 26 7.3
TC16608 weakly similar to UP|Q9FWS8 (Q9FWS8) F1B16.7 protein, pa... 26 9.6
AV776853 26 9.6
TC14448 homologue to UP|Q9M531 (Q9M531) Histone H2A, partial (95%) 26 9.6
>CB828260
Length = 557
Score = 78.2 bits (191), Expect = 2e-15
Identities = 39/122 (31%), Positives = 67/122 (53%), Gaps = 2/122 (1%)
Frame = +3
Query: 6 LVCERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQ--TKEWRLNILNGVHNHPMEP 63
L CE+ G ++ + GT ++K C F + G K ++WRL ++ G HNH
Sbjct: 192 LGCEKHGKYVPYRDPDLVEGTRTQKTECPFRLKGRPMKNGTDRDWRLKVMEGAHNHEPAR 371
Query: 64 ALEGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKA 123
+L GH GRL ++K+ V ++K+ +LPR +L+ LK P +T + Q+Y +++ K+
Sbjct: 372 SLLGHNFVGRLNSEEKEQVEKMSKSWVLPRKMLLTLKENNPLNLTTISQIYGACKRLRKS 551
Query: 124 NR 125
R
Sbjct: 552 LR 557
>BP038017
Length = 570
Score = 66.6 bits (161), Expect = 5e-12
Identities = 41/136 (30%), Positives = 65/136 (47%), Gaps = 2/136 (1%)
Frame = +2
Query: 127 DKKPLQYLISKLEEHKYTYYSRTQLES-NTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKT 185
D + L K++ +Y QL+ N + ++FWA S +N F ++ D+ Y+
Sbjct: 167 DAQNLLNYFKKMQGKNPGFYYAIQLDDENRMINVFWADARSRSAYNYFGDAVIFDTMYRP 346
Query: 186 NIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKV-IVTD 244
N Y++P GV G + E E +F W + R LS+ + P V I TD
Sbjct: 347 NQYQVPFAPFTGVNHHGQNVLFGCALLLDESESSFTW---LFRTWLSAMNDRPPVSITTD 517
Query: 245 RDMSLMKAVAHVFPES 260
+D ++ AVA VFPE+
Sbjct: 518 QDRAIQAAVAQVFPET 565
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 44.3 bits (103), Expect = 3e-05
Identities = 21/62 (33%), Positives = 31/62 (49%)
Frame = +2
Query: 131 LQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRM 190
L YL K + +Y T+ ++E++FW S +N F V+ DSTYK N Y
Sbjct: 164 LSYLEGKADNDPMFFYKFTKTGDESLENLFWCDGVSRMDYNVFGDVIAFDSTYKKNKYNK 343
Query: 191 PM 192
P+
Sbjct: 344 PL 349
>CN825057
Length = 721
Score = 43.5 bits (101), Expect = 4e-05
Identities = 40/163 (24%), Positives = 68/163 (41%), Gaps = 7/163 (4%)
Frame = +1
Query: 43 KQTKEWRLNILNGVHNHPMEPALEGHILAGR-LKEDDKKIVRDLTKNKMLPRNILIHLKN 101
K +W ++ HNH + PAL H R +K +K + L R + + +
Sbjct: 208 KPDGKWIIHEFIKEHNHELLPALAYHFRIHRNVKLAEKNNMDILHAVSERTRKMYVEMSR 387
Query: 102 QRPHCMTNVKQVYNERQQIWKA-----NRGDKKP-LQYLISKLEEHKYTYYSRTQLESNT 155
Q C+ V + Q K + GD + L+Y +E+ +YS E
Sbjct: 388 QSGGCLNIESLVGDLNDQFKKGQYLAMDEGDAQVMLEYFKHIQKENPNFFYSIDLNEEQR 567
Query: 156 IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGV 198
+ +IFW SI + +F V+ D++Y + ++P VGV
Sbjct: 568 LRNIFWIDAKSINDYLSFNDVVSFDTSYIKSNEKLPFAPFVGV 696
>AV425218
Length = 419
Score = 40.0 bits (92), Expect = 5e-04
Identities = 23/93 (24%), Positives = 44/93 (46%)
Frame = +2
Query: 191 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLM 250
P+ +VGV + G ++ E E+FVW+ + L +S P+ I+TD ++
Sbjct: 5 PLATLVGVNHHGQSVLFGCALLSSEDSESFVWLFQSLLHCMSGV--PPQGIITDHSEAMK 178
Query: 251 KAVAHVFPESYAMNCYFHVQANVKQRCVLDCKY 283
KA+ V P + C ++ + Q+ + +Y
Sbjct: 179 KAIETVLPSTRHRWCLSYIMKKLPQKLLGYAQY 277
>TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transposase-like
{Oryza sativa (japonica cultivar-group);}, partial (8%)
Length = 488
Score = 37.0 bits (84), Expect = 0.004
Identities = 22/95 (23%), Positives = 41/95 (43%)
Frame = +1
Query: 131 LQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRM 190
LQY KL E+ Y++ + + I ++FW + + F ++ +DSTY T+
Sbjct: 58 LQYFQRKLVENSPFYHAYQLDDEDQITNVFWVDARMLIDYGYFGDMVSLDSTYCTHSSNR 237
Query: 191 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLK 225
P+ G G + E E++ W+ +
Sbjct: 238 PLAVFSGFNHHRKAVIFGAALLYDETTESY*WLFE 342
>TC14447 homologue to UP|Q9M531 (Q9M531) Histone H2A, partial (86%)
Length = 755
Score = 28.1 bits (61), Expect = 1.9
Identities = 26/85 (30%), Positives = 40/85 (46%)
Frame = +2
Query: 17 PKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGRLKE 76
PKKKP + S K G F + G + K+ R G P+ A LA + E
Sbjct: 125 PKKKPV---SRSIKAGLQFPV-GRIGRYLKKGRYAQRVGT-GAPVYLAAVLEYLAAEVLE 289
Query: 77 DDKKIVRDLTKNKMLPRNILIHLKN 101
RD KN+++PR++L+ ++N
Sbjct: 290 LAGNAARDNKKNRIIPRHVLLAVRN 364
>TC14446 homologue to UP|H2A_LYCES (P25469) Histone H2A, partial (94%)
Length = 744
Score = 28.1 bits (61), Expect = 1.9
Identities = 26/85 (30%), Positives = 40/85 (46%)
Frame = +3
Query: 17 PKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGRLKE 76
PKKKP + S K G F + G + K+ R G P+ A LA + E
Sbjct: 153 PKKKPV---SRSVKAGLQFPV-GRIGRYLKKGRYAQRVGT-GAPVYLAAVLEYLAAEVLE 317
Query: 77 DDKKIVRDLTKNKMLPRNILIHLKN 101
RD KN+++PR++L+ ++N
Sbjct: 318 LAGNAARDNKKNRIIPRHVLLAVRN 392
>TC19004
Length = 445
Score = 26.2 bits (56), Expect = 7.3
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 2/45 (4%)
Frame = -3
Query: 47 EWRLNILNGVHNHPMEPALEGHIL--AGRLKEDDKKIVRDLTKNK 89
+W L+ HP P EG AGR + D KK+ + +NK
Sbjct: 350 KWSSVFLHYSTRHPTRPNREGLFW*RAGRFRSDPKKLDKPTGRNK 216
>BP074200
Length = 411
Score = 26.2 bits (56), Expect = 7.3
Identities = 9/30 (30%), Positives = 14/30 (46%)
Frame = +3
Query: 22 KHAGTGSRKCGCLFMISGYQSKQTKEWRLN 51
K + CGC+F++ Y S W L+
Sbjct: 144 KESAKNGSCCGCMFLVQQYSSFYIGNWELS 233
>TC16608 weakly similar to UP|Q9FWS8 (Q9FWS8) F1B16.7 protein, partial
(19%)
Length = 686
Score = 25.8 bits (55), Expect = 9.6
Identities = 10/22 (45%), Positives = 16/22 (72%), Gaps = 1/22 (4%)
Frame = -3
Query: 48 WRLNILNGVHNH-PMEPALEGH 68
W L+IL+ +H+H P++ L GH
Sbjct: 81 WSLSILHDMHDHSPVQQQLRGH 16
>AV776853
Length = 625
Score = 25.8 bits (55), Expect = 9.6
Identities = 22/92 (23%), Positives = 34/92 (36%), Gaps = 9/92 (9%)
Frame = +1
Query: 3 MFRLVCERSGSHIVPKKKPKHAGTGSRK--------CGCLFMISGYQSKQTKEWRLNILN 54
M + VC R G + KH RK CL + + + +W++
Sbjct: 352 MRQFVCNRQGL-----RSKKHYNRTDRKRDHKPVTHTNCLAKLRVHLDYKIGKWKVVSFE 516
Query: 55 GVHNHPMEPALEGHILAG-RLKEDDKKIVRDL 85
HNH + PA H + R+ D K D+
Sbjct: 517 ECHNHELTPARFVHFIPPYRVMNDADKAKGDM 612
>TC14448 homologue to UP|Q9M531 (Q9M531) Histone H2A, partial (95%)
Length = 715
Score = 25.8 bits (55), Expect = 9.6
Identities = 11/32 (34%), Positives = 20/32 (62%)
Frame = +3
Query: 70 LAGRLKEDDKKIVRDLTKNKMLPRNILIHLKN 101
LA + E RD KN+++PR++L+ ++N
Sbjct: 276 LAAEVLELAGNAARDNKKNRIIPRHVLLAVRN 371
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,662,776
Number of Sequences: 28460
Number of extensions: 76496
Number of successful extensions: 395
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 393
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 393
length of query: 305
length of database: 4,897,600
effective HSP length: 90
effective length of query: 215
effective length of database: 2,336,200
effective search space: 502283000
effective search space used: 502283000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC136507.18


