

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.1 - phase: 0
(236 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
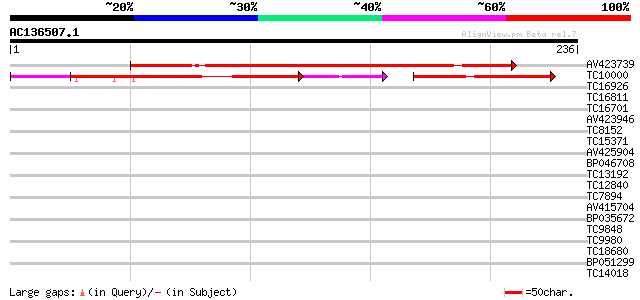
Score E
Sequences producing significant alignments: (bits) Value
AV423739 222 4e-59
TC10000 97 1e-33
TC16926 weakly similar to UP|Q9LHH2 (Q9LHH2) Ankyrin-like protei... 36 0.007
TC16811 35 0.011
TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome ... 34 0.025
AV423946 32 0.096
TC8152 31 0.21
TC15371 similar to GB|AAP37820.1|30725596|BT008461 At1g29790 {Ar... 30 0.37
AV425904 30 0.37
BP046708 30 0.48
TC13192 weakly similar to UP|Q8M0B6 (Q8M0B6) Orf102, partial (22%) 29 0.81
TC12840 similar to PIR|T02010|T02010 expansin homolog T15B16.16 ... 29 0.81
TC7894 weakly similar to UP|Q93YY0 (Q93YY0) 68 kDa protein HP68,... 28 1.1
AV415704 28 1.1
BP035672 28 1.4
TC9848 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lect... 28 1.8
TC9980 similar to UP|AAR23721 (AAR23721) At1g29470, partial (15%) 27 2.4
TC18680 27 2.4
BP051299 27 2.4
TC14018 similar to UP|Q43465 (Q43465) Protein kinase 2, partial ... 27 3.1
>AV423739
Length = 465
Score = 222 bits (566), Expect = 4e-59
Identities = 107/161 (66%), Positives = 130/161 (80%)
Frame = +1
Query: 51 FRKTLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMA 110
F + SRMSL+KK+L S+VEN ++E DS Q+LSVLDLP+L LECILE+LPPS+LCQMA
Sbjct: 1 FHCQIPSRMSLMKKSLSSRVENDDQE-DS--QDLSVLDLPDLALECILERLPPSALCQMA 171
Query: 111 GVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPRSLIKF 170
GVC SLRE CVSDHLWERHMKQKWG+VIG AYREWKWHVA+ +N+G L+HGK R L++F
Sbjct: 172 GVCRSLRESCVSDHLWERHMKQKWGKVIGPAAYREWKWHVASTRNVGGLKHGKQRGLMRF 351
Query: 171 MSLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLAL 211
+SL WPF WM++K D N +KQ S+ P DS M+WYLAL
Sbjct: 352 LSLRWPFPWMKLKGD---PNVRSKQGSFSPVDSVMSWYLAL 465
>TC10000
Length = 934
Score = 97.4 bits (241), Expect = 2e-21
Identities = 57/102 (55%), Positives = 68/102 (65%), Gaps = 5/102 (4%)
Frame = +1
Query: 26 SWTNEVRLISLLFHKDF-CLKSFCKL----FRKTLVSRMSLVKKTLVSKVENVEEEVDSL 80
SWT+ +RLISLLF+K+ C+KSF F + ++MSL KKTL SK ENVE+ + L
Sbjct: 148 SWTSGMRLISLLFYKELLCVKSFRNCLWLSFLYQIPTKMSLAKKTLTSKFENVEDTREML 327
Query: 81 QQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVS 122
VLECILE+LP SSLCQMA VCHSLRERCVS
Sbjct: 328 ------------VLECILERLPQSSLCQMASVCHSLRERCVS 417
Score = 95.1 bits (235), Expect(2) = 1e-33
Identities = 68/161 (42%), Positives = 84/161 (51%), Gaps = 4/161 (2%)
Frame = +3
Query: 1 MLLYFITCVSFFLFLKPLLLLKPLPS----WTNEVRLISLLFHKDFCLKSFCKLFRKTLV 56
MLLYFITCV FFLF KPLL LKPLPS W +EV L +L C ++
Sbjct: 69 MLLYFITCVPFFLFSKPLLFLKPLPSIMD*W-DEVDLSFVLQRAFVC----------EIL 215
Query: 57 SRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSL 116
+ L +L + +NV + DS + V + PS G C
Sbjct: 216 *ELPLA*FSLSNPYKNVSCKEDSYFKV*KC*GHSRDVSVGMHT*EAPSIFTLSNGQCLPF 395
Query: 117 RERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIG 157
E + ++LWERH KQKWGRV G VAY EWKWHVA+K+NIG
Sbjct: 396 LEGEMCEYLWERHTKQKWGRV-GLVAYMEWKWHVASKRNIG 515
Score = 63.9 bits (154), Expect(2) = 1e-33
Identities = 28/59 (47%), Positives = 41/59 (69%)
Frame = +2
Query: 169 KFMSLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNREV 227
+ + L+ PFSWMRMKV +YS+ K SY+PPDSFMTWY+AL+ G+F + ++ +
Sbjct: 518 RLVPLH*PFSWMRMKVSTSYSS---KHRSYVPPDSFMTWYVALDKGSFWYLPYCFSESI 685
>TC16926 weakly similar to UP|Q9LHH2 (Q9LHH2) Ankyrin-like protein, partial
(8%)
Length = 801
Score = 35.8 bits (81), Expect = 0.007
Identities = 23/77 (29%), Positives = 33/77 (41%), Gaps = 1/77 (1%)
Frame = +2
Query: 60 SLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRER 119
SL +K++ N+ + S +S D PE V CIL L PS + A
Sbjct: 89 SL*QKSITEMATNLNDPNSSSSSVVSFSDFPEDVQLCILSFLGPSEIATFACTSKRFLSL 268
Query: 120 CVSD-HLWERHMKQKWG 135
C +D LW ++WG
Sbjct: 269 CTNDAKLWFTLCHRRWG 319
>TC16811
Length = 509
Score = 35.0 bits (79), Expect = 0.011
Identities = 22/78 (28%), Positives = 35/78 (44%)
Frame = +2
Query: 79 SLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
SL + D+PE + +L L P +C++A V + +D LWE + + ++
Sbjct: 203 SLPSKTGLDDVPENCISSLLMSLDPQEICKLARVNKAFHRASSADFLWESKLPPGYKFLV 382
Query: 139 GSVAYREWKWHVATKKNI 156
V E K TKK I
Sbjct: 383 NKVLGEE-KLGSMTKKEI 433
>TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome 5, BAC
clone:F24B18, partial (24%)
Length = 691
Score = 33.9 bits (76), Expect = 0.025
Identities = 27/74 (36%), Positives = 36/74 (48%), Gaps = 6/74 (8%)
Frame = +3
Query: 3 LYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKL------FRKTLV 56
L+ T F+L L LL L + ++ L+ LFH F +K CKL FR L
Sbjct: 399 LHVRTLERFWLILSGLLCL--ISFALSDSTLVGALFHLSFPMKILCKLHAS*FSFRFLLD 572
Query: 57 SRMSLVKKTLVSKV 70
RM L++K LV V
Sbjct: 573 RRM*LIRKGLVEMV 614
>AV423946
Length = 426
Score = 32.0 bits (71), Expect = 0.096
Identities = 15/58 (25%), Positives = 27/58 (45%)
Frame = +1
Query: 70 VENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
VE +E +S + D+PE + + L P +C++A V + +D +WE
Sbjct: 184 VEKNDEGSNSTSPRSGLGDIPESCISSLFMNLDPPDICKLARVNRAFHRASSADFVWE 357
>TC8152
Length = 715
Score = 30.8 bits (68), Expect = 0.21
Identities = 13/54 (24%), Positives = 26/54 (48%)
Frame = +1
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSV 141
+LPE + IL P +CQ+A + + R +D +WE + + ++ +
Sbjct: 196 ELPESCVALILGYADPPQICQLATLNRAFRAASSADFVWESKLPANYRAIVRKI 357
>TC15371 similar to GB|AAP37820.1|30725596|BT008461 At1g29790 {Arabidopsis
thaliana;}, partial (13%)
Length = 735
Score = 30.0 bits (66), Expect = 0.37
Identities = 11/43 (25%), Positives = 25/43 (57%), Gaps = 2/43 (4%)
Frame = +2
Query: 123 DHLWER--HMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGK 163
DH + + +++ + +IG + Y++ KW K + G ++HG+
Sbjct: 56 DHFFSKGVDLEKVYAPLIGKLGYKKVKWATGNKTDAGGVKHGE 184
>AV425904
Length = 429
Score = 30.0 bits (66), Expect = 0.37
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +2
Query: 198 YLPPDSFMTWYLALETGNFSFPAQVY 223
+LP +S + Y +L+ G F FPAQ Y
Sbjct: 2 FLPNNSMLALYFSLQGGKFWFPAQFY 79
>BP046708
Length = 498
Score = 29.6 bits (65), Expect = 0.48
Identities = 12/47 (25%), Positives = 25/47 (52%), Gaps = 7/47 (14%)
Frame = -2
Query: 124 HLWERH-------MKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGK 163
+LW H +++ + +IG + Y++ KW K + G ++HG+
Sbjct: 344 YLWLDHFFSXGVDLEKVYAPLIGKLGYKKVKWATGNKTDAGGVKHGE 204
>TC13192 weakly similar to UP|Q8M0B6 (Q8M0B6) Orf102, partial (22%)
Length = 550
Score = 28.9 bits (63), Expect = 0.81
Identities = 17/62 (27%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Frame = -2
Query: 155 NIGSLRHGKPRSLIKFMSLYWPFSWMRMK-VDAAYSNSANKQNSYLPPDSFMTWYLALET 213
N+G + HG+ + F L+W + W + + +NS+NK + Y T Y+ + T
Sbjct: 186 NLGLIGHGRTVPVFSFFLLFW-WPWSSISFCSSGLANSSNKGSEYKEES---TKYIXITT 19
Query: 214 GN 215
N
Sbjct: 18 RN 13
>TC12840 similar to PIR|T02010|T02010 expansin homolog T15B16.16 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(60%)
Length = 648
Score = 28.9 bits (63), Expect = 0.81
Identities = 11/43 (25%), Positives = 22/43 (50%), Gaps = 1/43 (2%)
Frame = +2
Query: 107 CQMAGV-CHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKW 148
CQ+ G C +L ++ W + +W R+ + AY++ +W
Sbjct: 122 CQVQGWNCSNLSQKSTVQEKWRHQIYHQWERLF*ASAYKQCRW 250
>TC7894 weakly similar to UP|Q93YY0 (Q93YY0) 68 kDa protein HP68, partial
(9%)
Length = 1599
Score = 28.5 bits (62), Expect = 1.1
Identities = 13/28 (46%), Positives = 15/28 (53%)
Frame = +3
Query: 11 FFLFLKPLLLLKPLPSWTNEVRLISLLF 38
FF+ KPLL L PLP+W L F
Sbjct: 861 FFIQRKPLLWLSPLPTWHFSFMTTELFF 944
>AV415704
Length = 355
Score = 28.5 bits (62), Expect = 1.1
Identities = 14/46 (30%), Positives = 21/46 (45%)
Frame = +1
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKW 134
LP+ V I L LC + R+ C SD +WE ++ +W
Sbjct: 10 LPQDVTFKITSLLQVRDLCALGCCSRFCRDLCFSDCIWESLVRTRW 147
>BP035672
Length = 485
Score = 28.1 bits (61), Expect = 1.4
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = -3
Query: 11 FFLFLKPLLLLKPLPSW 27
FF+ KPLL L PLP+W
Sbjct: 468 FFIQRKPLLWLSPLPTW 418
>TC9848 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (19%)
Length = 394
Score = 27.7 bits (60), Expect = 1.8
Identities = 14/50 (28%), Positives = 22/50 (44%)
Frame = +1
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
LPE + IL P C+ + + +LR SD LW + + +I
Sbjct: 40 LPEECVSEILSHTSPPDACRFSMLSSTLRSAANSDMLWRSFLPSDYSDII 189
>TC9980 similar to UP|AAR23721 (AAR23721) At1g29470, partial (15%)
Length = 709
Score = 27.3 bits (59), Expect = 2.4
Identities = 12/35 (34%), Positives = 20/35 (56%)
Frame = -2
Query: 4 YFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLF 38
+++TC+ LFLKP + K +E R+I +F
Sbjct: 636 FYLTCIPNILFLKPSCIYKRKVK*VSEKRIIHNIF 532
>TC18680
Length = 492
Score = 27.3 bits (59), Expect = 2.4
Identities = 11/47 (23%), Positives = 22/47 (46%)
Frame = +1
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKW 134
D+PE + + L P +C +A + + R +D +WE + +
Sbjct: 178 DIPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWETKLPSNY 318
>BP051299
Length = 539
Score = 27.3 bits (59), Expect = 2.4
Identities = 14/41 (34%), Positives = 19/41 (46%)
Frame = +2
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERH 129
LP+ + IL +LPP +L + VC S R S H
Sbjct: 5 LPQEIWAEILHRLPPKTLVKCTSVCKSWRSLITSTSFISLH 127
>TC14018 similar to UP|Q43465 (Q43465) Protein kinase 2, partial (27%)
Length = 559
Score = 26.9 bits (58), Expect = 3.1
Identities = 16/39 (41%), Positives = 20/39 (51%)
Frame = -3
Query: 17 PLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTL 55
P LL P + L+ L FH+ FC SF FR+TL
Sbjct: 272 PCLLNNPHDFFNTLA*LVLLCFHETFCF-SFNDFFRQTL 159
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,495,592
Number of Sequences: 28460
Number of extensions: 91535
Number of successful extensions: 766
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 757
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 760
length of query: 236
length of database: 4,897,600
effective HSP length: 87
effective length of query: 149
effective length of database: 2,421,580
effective search space: 360815420
effective search space used: 360815420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC136507.1


