

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
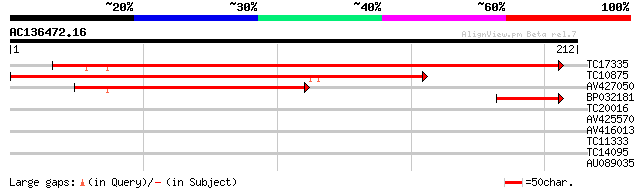
Query= AC136472.16 - phase: 0
(212 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17335 similar to UP|Q9M2C0 (Q9M2C0) Nodulin-like protein, part... 290 1e-79
TC10875 similar to UP|NO21_SOYBN (P16313) Nodulin 21 (N-21), par... 158 8e-40
AV427050 132 4e-32
BP032181 42 1e-04
TC20016 similar to UP|Q9MAW5 (Q9MAW5) Ribosomal protein L29, com... 31 0.18
AV425570 27 3.5
AV416013 26 4.5
TC11333 similar to GB|AAO42855.1|28416649|BT004609 At4g10170 {Ar... 26 4.5
TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein... 25 7.7
AU089035 25 7.7
>TC17335 similar to UP|Q9M2C0 (Q9M2C0) Nodulin-like protein, partial (85%)
Length = 911
Score = 290 bits (742), Expect = 1e-79
Identities = 155/200 (77%), Positives = 170/200 (84%), Gaps = 9/200 (4%)
Frame = +3
Query: 17 SKVYNMDVEKK---GEEENDEK------DYTQRAQWLRAAVLGANDGLLSTASLMMGVGA 67
SKV+N+DVE + EE+ + DYTQRAQWLRAAVLGANDGLLSTASLMMGVGA
Sbjct: 69 SKVFNIDVEGQLAHAEEDGGDALSHSAHDYTQRAQWLRAAVLGANDGLLSTASLMMGVGA 248
Query: 68 VTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDKLPNPYYA 127
V KDV+TMILTGIAGLVAGACSMAIGEFVSVYSQYDIE AQMKR+GN KDKLPNPYYA
Sbjct: 249 VKKDVRTMILTGIAGLVAGACSMAIGEFVSVYSQYDIELAQMKREGNTQNKDKLPNPYYA 428
Query: 128 AFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSL 187
A ASA+AFAVGA VPLL AAF+ +YKVR+GVVV V+LAL FG LSA LGK+PLVKSSL
Sbjct: 429 ALASALAFAVGAGVPLLAAAFITEYKVRIGVVVAAVTLALVVFGGLSAFLGKSPLVKSSL 608
Query: 188 RVLIGGWLAMSLTFGLTKLV 207
RVL GGWLAM++TFGLTKLV
Sbjct: 609 RVLFGGWLAMAVTFGLTKLV 668
>TC10875 similar to UP|NO21_SOYBN (P16313) Nodulin 21 (N-21), partial (58%)
Length = 581
Score = 158 bits (399), Expect = 8e-40
Identities = 86/168 (51%), Positives = 109/168 (64%), Gaps = 12/168 (7%)
Frame = +1
Query: 1 MAEIQHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTAS 60
MA +H T+ + + + + + + DY QRAQWLRAAVLGANDGL+S AS
Sbjct: 73 MAAGAPNHVGTIPNPANGTTDQNKQTQVTDVPRVVDYWQRAQWLRAAVLGANDGLVSVAS 252
Query: 61 LMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKR------QGN 114
LMMGVGAV KD K M+L G AGLVAGAC MAIGEFVSV++QY++E QMKR QG
Sbjct: 253 LMMGVGAVNKDAKAMLLAGFAGLVAGACGMAIGEFVSVFTQYEVEVGQMKRDMIKSEQGE 432
Query: 115 ------ISQKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRL 156
+ ++ +PNP AA ASA +F++G VPLL +FV YK+RL
Sbjct: 433 RDLEMAMEKRKSIPNPMQAALASAFSFSIGGLVPLLSGSFVSVYKIRL 576
>AV427050
Length = 420
Score = 132 bits (332), Expect = 4e-32
Identities = 67/90 (74%), Positives = 77/90 (85%), Gaps = 2/90 (2%)
Frame = +1
Query: 25 EKKGEEENDEK--DYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAG 82
E +G EE++ DY+QRAQWLRAAVLGANDGL+S ASLMMGVGAV +DV M+L G AG
Sbjct: 112 ESEGVEESNNNYIDYSQRAQWLRAAVLGANDGLVSVASLMMGVGAVKEDVSAMLLAGFAG 291
Query: 83 LVAGACSMAIGEFVSVYSQYDIEFAQMKRQ 112
LVAGACSMAIGEFVSVY+QYDIE AQ+KR+
Sbjct: 292 LVAGACSMAIGEFVSVYTQYDIEIAQIKRE 381
>BP032181
Length = 460
Score = 41.6 bits (96), Expect = 1e-04
Identities = 17/25 (68%), Positives = 23/25 (92%)
Frame = -2
Query: 183 VKSSLRVLIGGWLAMSLTFGLTKLV 207
++S +RVL GGWLAM++TFGLTKL+
Sbjct: 456 LRSCVRVLXGGWLAMAITFGLTKLI 382
>TC20016 similar to UP|Q9MAW5 (Q9MAW5) Ribosomal protein L29, complete
Length = 430
Score = 30.8 bits (68), Expect = 0.18
Identities = 23/54 (42%), Positives = 29/54 (53%), Gaps = 4/54 (7%)
Frame = -1
Query: 150 KDYKVRLGVVVGVVSLALF---GFGLLSAVLGKAPLVKSSLRV-LIGGWLAMSL 199
K K+R+ + V S LF GF L VL + PLV LRV +GG +AM L
Sbjct: 268 KKSKLRILIAPSVTST*LFFSDGFSTLLVVLPRVPLVPQKLRVHPLGGSVAMPL 107
>AV425570
Length = 420
Score = 26.6 bits (57), Expect = 3.5
Identities = 25/90 (27%), Positives = 40/90 (43%), Gaps = 8/90 (8%)
Frame = +1
Query: 86 GACSMAIGEFVSVYSQYDIEFAQM--------KRQGNISQKDKLPNPYYAAFASAIAFAV 137
G CS+ I F++ + + AQ+ KR+G +K L YA+ AI A+
Sbjct: 49 GLCSLGIVPFINAQIVFQL-LAQVYPKLQDLQKREGEAGRKKILQYTRYASVGFAIVQAI 225
Query: 138 GAFVPLLGAAFVKDYKVRLGVVVGVVSLAL 167
G + L +V D+ V+ V+ L L
Sbjct: 226 GQVLFL--RPYVNDFSTE-WVISSVILLTL 306
>AV416013
Length = 309
Score = 26.2 bits (56), Expect = 4.5
Identities = 16/61 (26%), Positives = 29/61 (47%), Gaps = 1/61 (1%)
Frame = -1
Query: 39 QRAQWLRAAVLGA-NDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVS 97
QR + +R A+ A GLL + +G+G DV+ + + V C + E++S
Sbjct: 270 QRVRVVRTALRAALGAGLLCSVQPRLGIGHAH*DVRVALRLTLGVEVGNLCKELVAEYLS 91
Query: 98 V 98
+
Sbjct: 90 L 88
>TC11333 similar to GB|AAO42855.1|28416649|BT004609 At4g10170 {Arabidopsis
thaliana;}, partial (14%)
Length = 487
Score = 26.2 bits (56), Expect = 4.5
Identities = 13/29 (44%), Positives = 18/29 (61%), Gaps = 6/29 (20%)
Frame = -1
Query: 37 YTQRAQWLRAAVLGAN------DGLLSTA 59
YTQ+ QWL VLGA +GL+S++
Sbjct: 121 YTQQHQWLEQCVLGATTSDECFEGLISSS 35
>TC14095 homologue to UP|RS2_ARATH (P49688) 40S ribosomal protein S2,
partial (88%)
Length = 1134
Score = 25.4 bits (54), Expect = 7.7
Identities = 16/56 (28%), Positives = 26/56 (45%)
Frame = +1
Query: 32 NDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGA 87
N + T + +WL D +++A + T+ V+T + AGLVAGA
Sbjct: 22 NSRRSLTTKPKWLNVVAEIVVDSAVASAVV------ATEAVETEAVADAAGLVAGA 171
>AU089035
Length = 460
Score = 25.4 bits (54), Expect = 7.7
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +1
Query: 181 PLVKSSLRVLIGGWLAMSLTFGLTKL 206
P VK L+ WL MSL+F LT+L
Sbjct: 325 PRVKWXXSNLLXCWLPMSLSFELTEL 402
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,888,547
Number of Sequences: 28460
Number of extensions: 28273
Number of successful extensions: 152
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 150
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 151
length of query: 212
length of database: 4,897,600
effective HSP length: 86
effective length of query: 126
effective length of database: 2,450,040
effective search space: 308705040
effective search space used: 308705040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC136472.16


