

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.12 - phase: 0 /pseudo
(346 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
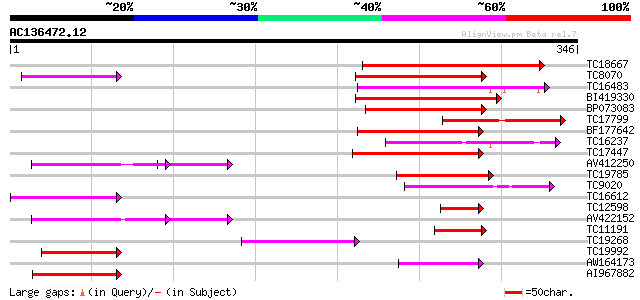
Score E
Sequences producing significant alignments: (bits) Value
TC18667 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein,... 108 2e-24
TC8070 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (73%) 86 1e-17
TC16483 homologue to UP|O24091 (O24091) MtN21 protein, partial (... 85 2e-17
BI419330 84 3e-17
BP073083 84 5e-17
TC17799 weakly similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (8%) 78 2e-15
BF177642 65 2e-11
TC16237 weakly similar to UP|Q84TV1 (Q84TV1) Nodulin-like protei... 64 3e-11
TC17447 weakly similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like ... 64 5e-11
AV412250 61 8e-11
TC19785 weakly similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (... 62 2e-10
TC9020 weakly similar to PIR|H84792|H84792 nodulin-like protein ... 59 2e-09
TC16612 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, part... 52 1e-07
TC12598 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, part... 49 1e-06
AV422152 45 2e-06
TC11191 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (9%) 48 3e-06
TC19268 similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein, partia... 47 5e-06
TC19992 similar to UP|Q8LE45 (Q8LE45) Nodulin-like protein, part... 45 2e-05
AW164173 44 3e-05
AI967882 44 5e-05
>TC18667 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein, partial
(27%)
Length = 911
Score = 108 bits (269), Expect = 2e-24
Identities = 50/112 (44%), Positives = 78/112 (69%), Gaps = 1/112 (0%)
Frame = +3
Query: 216 YAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLI 275
Y G+V SG + +++WC+ ++GPL+AS F PL+L++VA+AG L EKL+LGSIIG++LI
Sbjct: 570 YTGVVASGVVWVLMAWCLRLKGPLYASAFNPLLLVIVAIAGSLFLEEKLYLGSIIGSILI 749
Query: 276 VCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIV-VRSIEKDKSNLNNNN 326
V GLY V+WGK KE+K + + E +P+EIV + I+ +++ N+
Sbjct: 750 VLGLYIVLWGKGKELKSHMEQKHKNDSVEVEPLEIVTTKQIDGKNVDIDGND 905
Score = 33.5 bits (75), Expect = 0.054
Identities = 25/87 (28%), Positives = 43/87 (48%)
Frame = +2
Query: 70 QEEEGKDDLDNTVSVISMRFIWGIISSKLLLTSPDFNFSNICIRNGQPYTSCNLYHGCFL 129
+EE+ KD ++ +S IS+ IWG + + + S D +I + C++
Sbjct: 50 KEEQAKDHNESALSSISLWIIWGNHTREFVCGSSDLGRCDI--------S*CHV------ 187
Query: 130 RYGKIEFENKSREGKDIRYTNRNWWGN 156
GKIE NK+ EG+ + +RN W +
Sbjct: 188 --GKIEH*NKNWEGQGGWHFDRN*WSH 262
>TC8070 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (73%)
Length = 1758
Score = 85.5 bits (210), Expect = 1e-17
Identities = 38/80 (47%), Positives = 54/80 (67%)
Frame = +1
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG+V SG AV WC+ GP+F +V+ P+ L+VA+ L E+ +LG IIG
Sbjct: 880 FTILYAGVVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASLALGEEFYLGGIIG 1059
Query: 272 AVLIVCGLYAVVWGKSKEMK 291
A+LIV GLY V+WGK+ E K
Sbjct: 1060AILIVVGLYLVLWGKNAERK 1119
Score = 39.7 bits (91), Expect = 8e-04
Identities = 20/61 (32%), Positives = 35/61 (56%)
Frame = +3
Query: 8 CNVVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILE 67
C+V + L+ M+ +Q +AG +V+ + A+N G+S + YR I A + P A+ LE
Sbjct: 99 CSVPERLQLHGAMLALQFGYAGFHVVSRAALNMGISKLVFPVYRNIIALLLLLPFAYFLE 278
Query: 68 R 68
+
Sbjct: 279 K 281
>TC16483 homologue to UP|O24091 (O24091) MtN21 protein, partial (24%)
Length = 739
Score = 84.7 bits (208), Expect = 2e-17
Identities = 50/129 (38%), Positives = 78/129 (59%), Gaps = 12/129 (9%)
Frame = +1
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ AYAGIV S V + +GP+FA+ F+PLM+++VA+ G IL E++ LG IIG+
Sbjct: 31 AAAYAGIVTSSISYYVQGLVIRKKGPVFATAFSPLMMIIVAIMGSFILAEQIFLGGIIGS 210
Query: 273 VLIVCGLYAVVWGKSKEMKK----KNQLVPSK----NPHEFDPVEIVVRSIEKDKSN--- 321
+LIV GLY+V+WGK KE + ++ +P K N ++ PV+ + ++ KSN
Sbjct: 211 ILIVIGLYSVLWGKHKEQVETKVGEDIALPVKSGQFNGNQGTPVDSTDQCCDEAKSNEKV 390
Query: 322 -LNNNNQMV 329
N N+ +V
Sbjct: 391 EANQNSSIV 417
>BI419330
Length = 470
Score = 84.3 bits (207), Expect = 3e-17
Identities = 39/89 (43%), Positives = 56/89 (62%)
Frame = +1
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
++ AY+GIV S V + +GP+F + F PL +++V C +L+EKL+LGS+IG
Sbjct: 190 FACAYSGIVTSAVQFYVQGTVIKTKGPVFVTAFNPLRMIIVTALACILLSEKLYLGSLIG 369
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQLVPSK 300
V++V GLY VVWGK+KE K P K
Sbjct: 370 GVVVVMGLYLVVWGKAKEQKNLIPPCPEK 456
>BP073083
Length = 352
Score = 83.6 bits (205), Expect = 5e-17
Identities = 39/74 (52%), Positives = 54/74 (72%)
Frame = -2
Query: 218 GIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVC 277
G V SG + +++WC+ ++GPL AS F PL+L++VAL G KL+LGSIIG++LIV
Sbjct: 348 GGVASGVVWVLMAWCLRLKGPL*ASAFHPLLLVLVALPGSLFFEGKLYLGSIIGSILIVL 169
Query: 278 GLYAVVWGKSKEMK 291
GLY V+WG KE+K
Sbjct: 168 GLYLVLWG*GKELK 127
>TC17799 weakly similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (8%)
Length = 497
Score = 78.2 bits (191), Expect = 2e-15
Identities = 40/75 (53%), Positives = 56/75 (74%)
Frame = +1
Query: 265 HLGSIIGAVLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNN 324
++GSIIGAVLIVCGLYAV+W KSKEMK LV S +E D VEIV+RS ++++ +N
Sbjct: 19 YIGSIIGAVLIVCGLYAVIWVKSKEMKMNTVLVSS---NESDRVEIVLRSTGSEENSNSN 189
Query: 325 NNQMVKDNEDSQTDE 339
Q+++D+EDS ++
Sbjct: 190 GIQVIRDDEDSSAED 234
>BF177642
Length = 482
Score = 65.1 bits (157), Expect = 2e-11
Identities = 27/77 (35%), Positives = 48/77 (62%)
Frame = +1
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+V Y+ I V V +W +GP++ ++F+PL +++ + G L + L+LGS+IGA
Sbjct: 136 AVCYSAIFVVTMRSVVYAWAFRKKGPIYVAMFSPLGMVIAVVLGVLFLGDILYLGSLIGA 315
Query: 273 VLIVCGLYAVVWGKSKE 289
V+I G Y V+W +++E
Sbjct: 316 VIIAIGFYGVIWAQAQE 366
>TC16237 weakly similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein, partial
(11%)
Length = 695
Score = 64.3 bits (155), Expect = 3e-11
Identities = 41/111 (36%), Positives = 66/111 (58%), Gaps = 4/111 (3%)
Frame = +3
Query: 230 SWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGLYAVVWGKSKE 289
+W + ++GPL+AS F PL L++VA+AG L+EKL+LGS+IGA+LIV L A+ +
Sbjct: 78 AWVLRLKGPLYASSFNPLFLVIVAIAGSLFLDEKLYLGSVIGALLIV--LXAIXLSCGVK 251
Query: 290 MKK----KNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQ 336
+K +Q + + + + +EIV S KS ++++ V D D Q
Sbjct: 252 VKS*RAMLSQSIKMTHLQKLNXLEIVTTSPIHGKS---DDDKSVDDGNDMQ 395
>TC17447 weakly similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like protein,
partial (24%)
Length = 590
Score = 63.5 bits (153), Expect = 5e-11
Identities = 30/80 (37%), Positives = 49/80 (60%)
Frame = +3
Query: 210 AYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSI 269
A S+ +GI S +AV + +H++G ++ ++F PL + + + G L + LHLGS+
Sbjct: 63 ALASILCSGIFGSILNLAVHTHMLHLKGAVYVAMFKPLSIAIAVVLGVMFLGDTLHLGSV 242
Query: 270 IGAVLIVCGLYAVVWGKSKE 289
IGA +I G Y V+WGK+ E
Sbjct: 243 IGATVISIGFYTVMWGKATE 302
>AV412250
Length = 403
Score = 61.2 bits (147), Expect(2) = 8e-11
Identities = 34/86 (39%), Positives = 52/86 (59%), Gaps = 1/86 (1%)
Frame = +3
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEE 73
+K LLM +VQ AFAG N+LYK+ + DGM + I+ YR I + AF +AFI ER
Sbjct: 9 MKAFLLMFVVQAAFAGNNILYKIVIFDGMPVSIITVYRQIISAAFTLSLAFIFER----- 173
Query: 74 GKDDLDN-TVSVISMRFIWGIISSKL 98
+++ N T ++S+ F+ G++ L
Sbjct: 174--NNMPNLTFKLLSISFLNGLLGGSL 245
Score = 21.6 bits (44), Expect(2) = 8e-11
Identities = 12/46 (26%), Positives = 19/46 (41%)
Frame = +2
Query: 91 WGIISSKLLLTSPDFNFSNICIRNGQPYTSCNLYHGCFLRYGKIEF 136
WG K + N S++ + QPY+ G +R+G F
Sbjct: 233 WGKSVPKFIF*IIKLNISHLRHGSVQPYSGHYFCFGHLIRFGNSRF 370
>TC19785 weakly similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (16%)
Length = 514
Score = 61.6 bits (148), Expect = 2e-10
Identities = 28/59 (47%), Positives = 38/59 (63%)
Frame = +2
Query: 237 GPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGLYAVVWGKSKEMKKKNQ 295
GP+ AS++ PL L+VAL + E LG IIGA LI+ GLY VVWG+++E K +
Sbjct: 5 GPVLASIYLPLQTLLVALMSSIVFGEDFFLGGIIGAFLIMTGLYLVVWGRTQETKSAQE 181
>TC9020 weakly similar to PIR|H84792|H84792 nodulin-like protein [imported]
- Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(16%)
Length = 759
Score = 58.5 bits (140), Expect = 2e-09
Identities = 33/91 (36%), Positives = 51/91 (55%)
Frame = +3
Query: 242 SVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGLYAVVWGKSKEMKKKNQLVPSKN 301
+ F PL +++VA+ G L E+++LG +IGA++I+ GLY VVWGK+K+ P N
Sbjct: 3 TAFNPLCMVIVAILGSFCLAEQMYLGRVIGAIVILLGLYLVVWGKNKDYDSPPS--PIIN 176
Query: 302 PHEFDPVEIVVRSIEKDKSNLNNNNQMVKDN 332
H P + S KDK +N+ + N
Sbjct: 177 EHVL-PDKQTTESNAKDKEKPSNHELITISN 266
>TC16612 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (39%)
Length = 589
Score = 52.0 bits (123), Expect = 1e-07
Identities = 26/68 (38%), Positives = 38/68 (55%)
Frame = +3
Query: 1 MMMKNNLCNVVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIA 60
M +N V Q KP L MV +Q +AG+ ++ ++ GMS ++ YR I A F+A
Sbjct: 117 MTSENGFTKVFQKFKPYLAMVSLQFGYAGMYIITMVSFKHGMSHWVLSVYRHIIAFCFMA 296
Query: 61 PIAFILER 68
P A +LER
Sbjct: 297 PFALVLER 320
>TC12598 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (8%)
Length = 349
Score = 48.9 bits (115), Expect = 1e-06
Identities = 19/26 (73%), Positives = 25/26 (96%)
Frame = +2
Query: 264 LHLGSIIGAVLIVCGLYAVVWGKSKE 289
+HLGSIIGA+++VCGLY VVWGKS++
Sbjct: 2 IHLGSIIGAIILVCGLYPVVWGKSQD 79
>AV422152
Length = 465
Score = 45.4 bits (106), Expect(2) = 2e-06
Identities = 26/85 (30%), Positives = 47/85 (54%)
Frame = +2
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEE 73
+KP L +V +Q ++G+ V+ ++ GMS I+ YR + A IAP AF+LER +
Sbjct: 104 VKPYLAIVSLQFGYSGMYVITMVSFKHGMSHWILSVYRHVVALIIIAPFAFVLER--KIR 277
Query: 74 GKDDLDNTVSVISMRFIWGIISSKL 98
K L + ++++ F+ ++ L
Sbjct: 278 PKMTLPIFLRIVALGFLEPVLDQNL 352
Score = 22.3 bits (46), Expect(2) = 2e-06
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +1
Query: 96 SKLLLTSPDFNFSNICIRNGQPYTSCNLYHGCFLRYGKIEF 136
S+L+ + + +NICIR+ +S Y+G + G EF
Sbjct: 343 SELVQHGYEDDLNNICIRHC*CPSSHYFYNGTHFQVGDSEF 465
>TC11191 similar to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (9%)
Length = 544
Score = 47.8 bits (112), Expect = 3e-06
Identities = 22/32 (68%), Positives = 26/32 (80%)
Frame = +2
Query: 260 LNEKLHLGSIIGAVLIVCGLYAVVWGKSKEMK 291
L E+ +LG IIGAVLIV GLY V+WGKS+E K
Sbjct: 2 LGEEFYLGGIIGAVLIVVGLYLVLWGKSEEKK 97
>TC19268 similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein, partial (8%)
Length = 505
Score = 47.0 bits (110), Expect = 5e-06
Identities = 29/72 (40%), Positives = 37/72 (51%)
Frame = +1
Query: 142 EGKDIRYTNRNWWGNGIDIR*RC*N*DGIIPFKSFTSSKWCCNTHYLHC*YHFGFSMCLG 201
EG D Y N NWWGNG D *R N D ++P + F K +T + * G S+
Sbjct: 1 EG*DSGYNNWNWWGNGSDFG*RQGNQDRVLPREPFAPPKRYTSTRRWYR*EPLGCSLQPS 180
Query: 202 **HLLCTLAYYS 213
HLL +A+YS
Sbjct: 181 QWHLLLIVAHYS 216
>TC19992 similar to UP|Q8LE45 (Q8LE45) Nodulin-like protein, partial (46%)
Length = 615
Score = 44.7 bits (104), Expect = 2e-05
Identities = 21/49 (42%), Positives = 33/49 (66%)
Frame = +2
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
+V +Q +AG++VL K A+N+GMS ++V YR A +AP A IL++
Sbjct: 59 VVFIQSGYAGMDVLCKAALNNGMSNYVLVVYRHAVAFVVMAPFALILDK 205
>AW164173
Length = 377
Score = 44.3 bits (103), Expect = 3e-05
Identities = 18/52 (34%), Positives = 29/52 (55%)
Frame = +3
Query: 238 PLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGLYAVVWGKSKE 289
P F +++ L + LH G+++GAV++ G YAV+WGK+KE
Sbjct: 114 PCVYITFQASTIVIATSMSVIFLGDALHFGTVVGAVILSIGFYAVIWGKAKE 269
Score = 32.3 bits (72), Expect = 0.12
Identities = 12/35 (34%), Positives = 22/35 (62%)
Frame = +1
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPL 247
S+ Y+G +G V +W + ++GP++ S+F PL
Sbjct: 40 SILYSGFFSTGLSCLVHTWGLRLKGPVYISLFKPL 144
>AI967882
Length = 381
Score = 43.5 bits (101), Expect = 5e-05
Identities = 21/54 (38%), Positives = 33/54 (60%)
Frame = +1
Query: 15 KPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
K L + MV + + G +V+ K+A+NDG++ + YR + A +APIAF ER
Sbjct: 94 KSHLGLAMVHLLYGGYHVITKVALNDGVNQLVFCFYRDLLAFCILAPIAFFKER 255
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.145 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,265,650
Number of Sequences: 28460
Number of extensions: 92926
Number of successful extensions: 902
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 896
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 901
length of query: 346
length of database: 4,897,600
effective HSP length: 91
effective length of query: 255
effective length of database: 2,307,740
effective search space: 588473700
effective search space used: 588473700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC136472.12


