

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.2 - phase: 0 /pseudo
(355 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
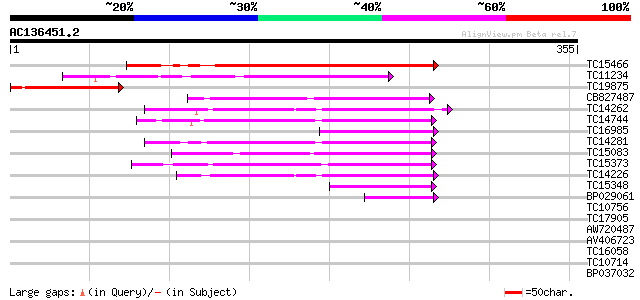
TC15466 homologue to UP|Q9LKH7 (Q9LKH7) Cytochrome P450, partial... 141 1e-34
TC11234 weakly similar to UP|Q9FQY4 (Q9FQY4) Ent-kaurene oxidase... 86 9e-18
TC19875 similar to UP|C90C_ARATH (Q9M066) Cytochrome P450 90C1 ... 82 1e-16
CB827487 63 9e-11
TC14262 UP|Q9MBE5 (Q9MBE5) Cytochrome P450, complete 60 7e-10
TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment)... 57 4e-09
TC16985 similar to UP|Q9LVY7 (Q9LVY7) Cytochrome P450-like, part... 55 1e-08
TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygen... 54 4e-08
TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, p... 51 3e-07
TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 hom... 50 8e-07
TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete 42 1e-04
TC15348 weakly similar to GB|AAA32913.1|166949|AVOCYP cytochrome... 41 3e-04
BP029061 40 8e-04
TC10756 weakly similar to UP|Q9SML3 (Q9SML3) Cytochrome P450 mon... 39 0.002
TC17905 weakly similar to UP|Q9STH8 (Q9STH8) Flavonoid 3', 5'-hy... 37 0.005
AW720487 35 0.025
AV406723 35 0.025
TC16058 weakly similar to UP|Q9LU04 (Q9LU04) Flavonoid 3',5'-hyd... 35 0.025
TC10714 similar to UP|Q9LSF8 (Q9LSF8) Cytochrome p450, partial (... 34 0.043
BP037032 33 0.095
>TC15466 homologue to UP|Q9LKH7 (Q9LKH7) Cytochrome P450, partial (57%)
Length = 1014
Score = 141 bits (356), Expect = 1e-34
Identities = 76/195 (38%), Positives = 122/195 (61%)
Frame = +3
Query: 74 KSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSN 133
K++KA+ ++ + + IV +R K E ++K ND++ ALL +S +
Sbjct: 72 KAIKARTKVAEALTLIVRQRRK-------EGEKK----NDMLGALL---------ASGDH 191
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
E I ++ ++ G ET T MTLA+KFLT+ PLAL++L EE+ +++ +K+
Sbjct: 192 FSDEQIVDFMLALLVAGYETTSTIMTLAVKFLTENPLALAQLKEEHDQIRAKKSYPEAPL 371
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
WTDY S+ FTQ V++ETLR+ANI+ GI+R+A+ D+ IKGY IPK W V AS +VH++
Sbjct: 372 EWTDYKSMAFTQCVVNETLRVANIIGGIFRRAMTDINIKGYTIPKGWKVFASFRAVHLNP 551
Query: 254 KNYENPYKFDPWRWE 268
++++ F+PWRW+
Sbjct: 552 DHFKDARTFNPWRWQ 596
>TC11234 weakly similar to UP|Q9FQY4 (Q9FQY4) Ent-kaurene oxidase, partial
(16%)
Length = 595
Score = 85.9 bits (211), Expect = 9e-18
Identities = 62/211 (29%), Positives = 104/211 (48%), Gaps = 4/211 (1%)
Frame = +2
Query: 34 IKIPKGNSGWPLLGETLDF----IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRI 89
I I G+S ++ E +D + +G S P+ + +K+LKA++++ K++ +
Sbjct: 2 IHILMGSSIHHMIIENMDTSFAELTNGILSAPIN---APGFVFHKALKARKKLAKILQSV 172
Query: 90 VEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIP 149
V+ER +L N E +KV +D LL K E + ++ +I+Q +
Sbjct: 173 VDER-RLRSKNGQEGKDKV-----FLDNLLEAKDENGRKRDDEYIVDVLIAQ-----LFA 319
Query: 150 GEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVIS 209
G ET TA+ + +LT +P L K +E E+ K + + + + + VI
Sbjct: 320 GHETSATALMWTILYLTQHPHILEKAKKEQEEIMKARVSSQGRLNLQEIKQMVYLSQVID 499
Query: 210 ETLRMANIVNGIWRKAVKDVEIKGYLIPKDW 240
ETLR ANIV +R+A+ DV I GY+IP W
Sbjct: 500 ETLRCANIVFTTFREAISDVNINGYVIPNGW 592
>TC19875 similar to UP|C90C_ARATH (Q9M066) Cytochrome P450 90C1
(ROTUNDIFOLIA3) , partial (16%)
Length = 623
Score = 82.4 bits (202), Expect = 1e-16
Identities = 37/71 (52%), Positives = 54/71 (75%)
Frame = +2
Query: 1 MEWFIGVYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSC 60
M W +G++ S++ +L+ WF+ +K+ +++ K+PKGN GWPLLGETLDFIA GY+S
Sbjct: 275 MAWIMGLW-SLMGILLCWWFLGVKKDQSRVKKNGKVPKGNMGWPLLGETLDFIACGYTSN 451
Query: 61 PVTFMEKRKSI 71
PV+FM KRKS+
Sbjct: 452 PVSFMTKRKSL 484
>CB827487
Length = 569
Score = 62.8 bits (151), Expect = 9e-11
Identities = 45/155 (29%), Positives = 78/155 (50%)
Frame = +3
Query: 112 NDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLA 171
+D+VD LL+ + +Q S S +L E I +++ +I +T A + L P A
Sbjct: 126 DDIVDTLLQLR---NQGSLSIDLTDEHIKAFMMDLLIGSTDTSVAASVWLMTGLMKNPTA 296
Query: 172 LSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEI 231
+ K+ EE L K D D L + + VI ETLR + I R+ +K++ +
Sbjct: 297 MKKVQEEIRNLCVNK----DFIDEVDIQKLEYLKAVIKETLRFYPLAPLIPRETIKNIIV 464
Query: 232 KGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWR 266
GY IP V ++ ++H D + +++P++F+P R
Sbjct: 465 DGYEIPAKTIVYVNVWAIHRDPEAWKDPHEFNPDR 569
>TC14262 UP|Q9MBE5 (Q9MBE5) Cytochrome P450, complete
Length = 1960
Score = 59.7 bits (143), Expect = 7e-10
Identities = 50/195 (25%), Positives = 92/195 (46%), Gaps = 2/195 (1%)
Frame = +1
Query: 85 MVGRIVEERMKLIMDNNAEDDEKVGVVNDVV--DALLRDKGELSQSSSSSNLMVEMISQN 142
++ +++++R ++I + E VV D LL + + + E I
Sbjct: 805 VIEKVIKKRQEIIKRRKERNGELEEGEQSVVFLDTLLEYAAD---ENMEIKITKEQIKGL 975
Query: 143 IIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLP 202
+++F G ++ A AL L + P L K EE ++ K D +D +LP
Sbjct: 976 VVDFFSAGTDSTAVATDWALAELINNPRVLKKAREE-VDSVVGKDRLVDE---SDIQNLP 1143
Query: 203 FTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKF 262
+ + ++ ET RM + + RK V++ E+ GY+IP+ V+ ++ +V D K ++ P +F
Sbjct: 1144YIRAIVKETFRMHPPLPVVKRKCVQECELNGYVIPEGALVLFNVWAVQRDPKYWKTPLEF 1323
Query: 263 DPWRWEIIYLHEAII 277
P R +L EA I
Sbjct: 1324RPER----FLEEADI 1356
>TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment) , complete
Length = 1753
Score = 57.4 bits (137), Expect = 4e-09
Identities = 50/192 (26%), Positives = 88/192 (45%), Gaps = 4/192 (2%)
Frame = +1
Query: 80 ERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN----DVVDALLRDKGELSQSSSSSNLM 135
E + + + RI+E +I D+ A K G V D++D LL K E S + +L
Sbjct: 733 EYLHQKMDRILET---IIDDHKANSRTKEGQVEGGEEDLIDVLL--KYENSSTDQDFHLT 897
Query: 136 VEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTW 195
+ I + + I G ET T + + + P+ L K +E E+ +++ +
Sbjct: 898 IRNIKAILFDIFIAGSETSATTINWTMAEMMKDPILLKKAQDEVREIFQRRGKVDE---- 1065
Query: 196 TDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKN 255
T L + + I+E LR+ ++R+ + EI GY IP V+ + ++ DSK
Sbjct: 1066TCIYELKYLKAFINEVLRLHPPGPLVFRECRQACEINGYHIPAKSTVLVNTFAIGTDSKY 1245
Query: 256 YENPYKFDPWRW 267
+ P +F P R+
Sbjct: 1246WAEPERFCPERF 1281
>TC16985 similar to UP|Q9LVY7 (Q9LVY7) Cytochrome P450-like, partial (25%)
Length = 942
Score = 55.5 bits (132), Expect = 1e-08
Identities = 26/74 (35%), Positives = 40/74 (53%)
Frame = +1
Query: 195 WTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSK 254
W D + ++ NV E +R+A + G +R+A+ D G+ IPK W + S S H +
Sbjct: 4 WDDVNQMRYSWNVACEVMRVAPPLQGGFREALHDFMFNGFSIPKGWKLYWSANSTHKSPE 183
Query: 255 NYENPYKFDPWRWE 268
+ P KFDP R+E
Sbjct: 184 YFPEPEKFDPTRFE 225
>TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygenaseCYP93D1,
partial (97%)
Length = 1768
Score = 53.9 bits (128), Expect = 4e-08
Identities = 43/183 (23%), Positives = 83/183 (44%)
Frame = +3
Query: 85 MVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNII 144
M+ ++++E + A+ D K D+ D LL + + + L + +
Sbjct: 798 MMEKVLKEHEEARATEGADSDRK----KDLFDILLN---LIEADGADNKLTRDSAKAFAL 956
Query: 145 EFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFT 204
+ + G + + +L L P L K EE + ++ + +D +LP+
Sbjct: 957 DMFLAGTNGPASVLEWSLAELIRNPHVLKKAREEIDTVVGKERLVKE----SDIPNLPYL 1124
Query: 205 QNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDP 264
Q V+ ETLRM R+A++ ++ GY IP + + S ++ DS+ +ENP+ +DP
Sbjct: 1125QAVVKETLRMHPPTPIFAREAMRSCQVDGYDIPANSKIFISAYAIGRDSQYWENPHVYDP 1304
Query: 265 WRW 267
R+
Sbjct: 1305ERF 1313
>TC15083 similar to UP|GAR1_YEAST (P28007) snoRNP protein GAR1, partial (7%)
Length = 1367
Score = 50.8 bits (120), Expect = 3e-07
Identities = 45/167 (26%), Positives = 77/167 (45%), Gaps = 1/167 (0%)
Frame = +1
Query: 102 AEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLA 161
A++ + V V VD LL + + + + MV + S E + G +T TA+
Sbjct: 814 AKEGKPVNDVVSYVDTLLELELPEEKRKLNESEMVTLCS----EILNAGTDTTSTALEWV 981
Query: 162 LKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGI 221
L L YP +++E ++ K + + D + LP+ + VI E LR + +
Sbjct: 982 LANLDKYPRVQKNIVDEISDVMKGRED--KEVKEEDLVKLPYLKAVILEGLRRHPPGHFV 1155
Query: 222 WRKAV-KDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
AV +D + GYL+PKD V + + D + +E P +F P R+
Sbjct: 1156LPHAVTEDTVLDGYLVPKDGTVNFMVAEMGWDPQVWEEPMEFKPERF 1296
>TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 homolog,
partial (61%)
Length = 1059
Score = 49.7 bits (117), Expect = 8e-07
Identities = 42/192 (21%), Positives = 83/192 (42%), Gaps = 1/192 (0%)
Frame = +1
Query: 77 KAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMV 136
K R ++ +++ ER+K+ E + K D + LL K E S + L +
Sbjct: 151 KVVPRFDRIFEKMIGERVKM------ESEGKRSESKDFLQFLLNLKEE---GDSKTPLSI 303
Query: 137 EMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWT 196
+ +++ + G +T + A+ + P + ++ EE + + + +
Sbjct: 304 THVKALLMDMLAGGSDTSSNTVEFAMAEMMQKPEVMKRVQEELEGVVGRDNMVEESHIH- 480
Query: 197 DYMSLPFTQNVISETLRMANIVNGIWRKAVKDV-EIKGYLIPKDWGVMASLTSVHMDSKN 255
LP+ V+ ETLR+ ++ + + GY IPK V ++ ++H D
Sbjct: 481 ---KLPYLLAVMKETLRLHPVLPLLVPHCPSETTSAGGYTIPKGSRVFVNVWAIHRDPSI 651
Query: 256 YENPYKFDPWRW 267
+ENP +FDP R+
Sbjct: 652 WENPLEFDPTRF 687
>TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete
Length = 1878
Score = 42.4 bits (98), Expect = 1e-04
Identities = 38/165 (23%), Positives = 73/165 (44%), Gaps = 1/165 (0%)
Frame = +3
Query: 105 DEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKF 164
D+K N ++D LL Q S ++I + ++ G ++ + ++
Sbjct: 831 DKKQRTANTMIDHLLT-----LQESQPEYYTDQIIKGLALAMLLAGTDSSAVTLEWSMCN 995
Query: 165 LTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIW-R 223
+ +YP L K+ E ++ + D +D L + +NVI+ETLR+ +
Sbjct: 996 VLNYPEVLEKIKAE-LDTHVGQDRLVDE---SDIPKLTYLKNVINETLRLYTPAPLLLPH 1163
Query: 224 KAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
A D I GY +P+D V+ + ++H D + + F P R++
Sbjct: 1164SASDDCTIGGYKVPRDTIVLINAWALHRDPQLWTEATTFKPERFD 1298
>TC15348 weakly similar to GB|AAA32913.1|166949|AVOCYP cytochrome
P-450LXXIA1 (cyp71A1) {Persea americana;} , partial
(23%)
Length = 614
Score = 40.8 bits (94), Expect = 3e-04
Identities = 20/67 (29%), Positives = 39/67 (57%)
Frame = +2
Query: 201 LPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPY 260
L + + +I ETLR I R+ +K + + GY IP V ++ ++H D + +++P+
Sbjct: 11 LEYLKAIIKETLRFYPPAPLIPRETIKSIIVDGYEIPAKTIVYVNVWAIHRDPEAWKDPH 190
Query: 261 KFDPWRW 267
+F+P R+
Sbjct: 191 EFNPDRF 211
>BP029061
Length = 442
Score = 39.7 bits (91), Expect = 8e-04
Identities = 20/46 (43%), Positives = 25/46 (53%)
Frame = -3
Query: 223 RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
R+A+KD Y IPK W + S S H D + NP FDP R+E
Sbjct: 440 REALKDFTYADYDIPKGWKLHWSTGSSHKDPTLFSNPETFDPSRFE 303
>TC10756 weakly similar to UP|Q9SML3 (Q9SML3) Cytochrome P450 monooxygenase
(Fragment), partial (37%)
Length = 887
Score = 38.5 bits (88), Expect = 0.002
Identities = 34/113 (30%), Positives = 56/113 (49%), Gaps = 8/113 (7%)
Frame = +3
Query: 201 LPFTQNVISETLRMANIVNGIW-RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENP 259
LP+ Q ++ E R+ V + R+A D+++ GY IPK V+ ++ + D ++NP
Sbjct: 48 LPYLQAIVKEIFRLHPAVPLLLPRQAEVDLDMHGYTIPKGSQVLVNVWANGRDPNLWDNP 227
Query: 260 YKFDPWRW---EIIYLHEAIIYTMFYA----CRKLELCLVTIALPRLGVDIDC 305
F P R+ EI + + T F A C L L + + L LG+ I+C
Sbjct: 228 NLFSPERFLGSEIDFKGRSFELTPFGAGRRICPGLPLAIRLLFL-MLGLLINC 383
>TC17905 weakly similar to UP|Q9STH8 (Q9STH8) Flavonoid 3', 5'-hydroxylase
like protein (Flavonoid 3, 5-hydroxylase like protein),
partial (30%)
Length = 597
Score = 37.0 bits (84), Expect = 0.005
Identities = 22/68 (32%), Positives = 34/68 (49%), Gaps = 1/68 (1%)
Frame = +2
Query: 201 LPFTQNVISETLRMANIVNGIWRKAVKDV-EIKGYLIPKDWGVMASLTSVHMDSKNYENP 259
LP+ V+ ETLR+ V + + GY IPK V ++ ++H D +E P
Sbjct: 8 LPYLLAVMKETLRLHPTVPLLVPHCPSEATSTGGYTIPKGSRVFVNVWAIHRDPSIWEKP 187
Query: 260 YKFDPWRW 267
+FDP R+
Sbjct: 188 LEFDPERF 211
>AW720487
Length = 575
Score = 34.7 bits (78), Expect = 0.025
Identities = 18/61 (29%), Positives = 37/61 (60%)
Frame = +1
Query: 1 MEWFIGVYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSC 60
+E + +V I + L SL++ K ++ N +P G G+P++GE+L+F+++G++
Sbjct: 79 VEVMVHAFVVISLSLFSLFY--KHRSPFTAPN---LPPGRMGFPVIGESLEFLSTGWNGH 243
Query: 61 P 61
P
Sbjct: 244 P 246
>AV406723
Length = 430
Score = 34.7 bits (78), Expect = 0.025
Identities = 19/59 (32%), Positives = 36/59 (60%)
Frame = +1
Query: 10 SIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
SI + L SL++ K ++ N +P G G+P++GE+L+F+++G++ P F+ R
Sbjct: 106 SISLSLFSLFY--KHRSPFTAPN---LPPGRMGFPVIGESLEFLSTGWNGHPEKFIFDR 267
>TC16058 weakly similar to UP|Q9LU04 (Q9LU04) Flavonoid
3',5'-hydroxylase-like, cytochrome P450, partial (26%)
Length = 553
Score = 34.7 bits (78), Expect = 0.025
Identities = 14/37 (37%), Positives = 22/37 (58%)
Frame = +1
Query: 231 IKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
+ GY IPK V ++ ++ D N+E P +FDP R+
Sbjct: 46 VGGYTIPKGSQVFVNVWAIQRDPSNWEKPLEFDPTRF 156
>TC10714 similar to UP|Q9LSF8 (Q9LSF8) Cytochrome p450, partial (16%)
Length = 701
Score = 33.9 bits (76), Expect = 0.043
Identities = 17/56 (30%), Positives = 30/56 (53%)
Frame = +2
Query: 223 RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEIIYLHEAIIY 278
R+A D + GY +PK ++ +L + D + + NP +F P R+ + HE I +
Sbjct: 26 REATDDCHVAGYQVPKGTRLLINLWKLQRDPQIWSNPDEFQPERF--LNAHEDISF 187
>BP037032
Length = 577
Score = 32.7 bits (73), Expect = 0.095
Identities = 20/74 (27%), Positives = 35/74 (47%), Gaps = 1/74 (1%)
Frame = -2
Query: 197 DYMSLPFTQNVISETLRMANIVNGIW-RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKN 255
D +P+ ++ ET R + + A ++ E+ GY +PKD V + D
Sbjct: 459 DVEKMPYLSAIVKETFRRHPPSHFVLSHAATEETELGGYTVPKDASVEFYTAWLSEDPDM 280
Query: 256 YENPYKFDPWRWEI 269
+E+P +F P R+ I
Sbjct: 279 WEDPDEFRPERFLI 238
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,034,313
Number of Sequences: 28460
Number of extensions: 77917
Number of successful extensions: 505
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 501
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 503
length of query: 355
length of database: 4,897,600
effective HSP length: 91
effective length of query: 264
effective length of database: 2,307,740
effective search space: 609243360
effective search space used: 609243360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC136451.2


