

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.10 - phase: 0 /pseudo
(820 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
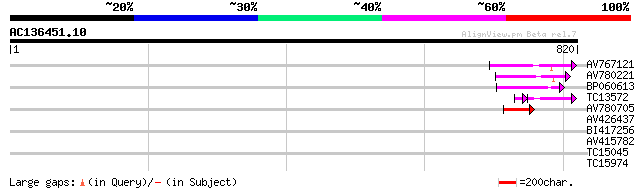
AV767121 97 1e-20
AV780221 95 4e-20
BP060613 51 8e-07
TC13572 45 1e-06
AV780705 45 5e-05
AV426437 32 0.41
BI417256 30 1.5
AV415782 29 3.4
TC15045 similar to AAR25638 (AAR25638) At1g55300, partial (66%) 29 3.4
TC15974 similar to UP|Q94KL4 (Q94KL4) BEL1-like homeodomain 1, p... 27 10.0
>AV767121
Length = 525
Score = 97.1 bits (240), Expect = 1e-20
Identities = 50/129 (38%), Positives = 70/129 (53%), Gaps = 3/129 (2%)
Frame = +2
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F GG+++ KI V W VC KE+GGLG++ + FN ALLGK W L E +SLW RV
Sbjct: 149 FLWGGSENENKIA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRV 328
Query: 754 LVARYGEEGGRLLDGGRTASAWWRAIATL---HSEAWFGTNVKRSVGNGENIYFWSDVWV 810
+ A+ D S+WW I +L ++ WF + +K+ VG G+ FWS+ W+
Sbjct: 329 IEAQ--------PDHYSCGSSWWNDILSLCPEDADGWFSSGLKKLVGEGDQTKFWSEDWL 484
Query: 811 GGVSFKERF 819
G RF
Sbjct: 485 GSGLLSARF 511
>AV780221
Length = 461
Score = 95.1 bits (235), Expect = 4e-20
Identities = 44/113 (38%), Positives = 65/113 (56%), Gaps = 4/113 (3%)
Frame = +3
Query: 703 KKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEG 762
KKI WV W+ VC K+ GGLGV+ + FN ALLGKW W +L E++ LWY+VL+ +Y
Sbjct: 138 KKIAWVKWSTVCRPKDEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKYKNA- 314
Query: 763 GRLLDGGRTASAWWRAIATLHSE----AWFGTNVKRSVGNGENIYFWSDVWVG 811
++AS+WW + + E W + R +G G + FW++ W+G
Sbjct: 315 -----IPQSASSWWNDLHSTCFEDGGGGWMQRGLCRRIGEGTEVKFWNENWLG 458
>BP060613
Length = 378
Score = 50.8 bits (120), Expect = 8e-07
Identities = 29/99 (29%), Positives = 48/99 (48%), Gaps = 1/99 (1%)
Frame = +1
Query: 705 ITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGR 764
+ W W+ + K+ GGLG + N ALL K +W +L ++LW ++L A Y
Sbjct: 73 VHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQILKALYFPHHDF 252
Query: 765 LLDGGRTASAW-WRAIATLHSEAWFGTNVKRSVGNGENI 802
L RT ++W W ++ LH + +G+GE +
Sbjct: 253 LQTTKRTGASWVWSSL--LHGRELLLKQGQWQLGSGETV 363
>TC13572
Length = 534
Score = 45.1 bits (105), Expect(2) = 1e-06
Identities = 24/73 (32%), Positives = 37/73 (49%), Gaps = 3/73 (4%)
Frame = -3
Query: 750 WYRVLVARYGEEGGRLLDGGRT-ASAWWRAIATLHS--EAWFGTNVKRSVGNGENIYFWS 806
W RV++A+YG DGG T S+WWR + T WF V + + +G + FW
Sbjct: 478 WCRVVLAKYG-------DGGCTNISSWWRDVQTAGGGENGWFEEGVHKVLRSGNHTRFWL 320
Query: 807 DVWVGGVSFKERF 819
+ W G +E++
Sbjct: 319 ENWTGVGILREKY 281
Score = 24.6 bits (52), Expect(2) = 1e-06
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = -1
Query: 731 NSALLGKWSWMVLVEKESL 749
N ALLGKW W + SL
Sbjct: 534 NKALLGKWLWRLKTVPNSL 478
>AV780705
Length = 524
Score = 45.1 bits (105), Expect = 5e-05
Identities = 21/44 (47%), Positives = 28/44 (62%)
Frame = -2
Query: 715 LSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARY 758
LSKE GG+G R +R N A L + +W VL E+LW RV+ + Y
Sbjct: 193 LSKEKGGVGFRDLRTQNLAFLARQAWRVLTNPEALWVRVMKSLY 62
>AV426437
Length = 303
Score = 32.0 bits (71), Expect = 0.41
Identities = 10/27 (37%), Positives = 19/27 (70%)
Frame = +1
Query: 699 NDDHKKITWVDWNYVCLSKEVGGLGVR 725
+++ +KI W++W +C K + GLGV+
Sbjct: 211 DENQRKIAWINWETICKPKWLRGLGVK 291
>BI417256
Length = 449
Score = 30.0 bits (66), Expect = 1.5
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = +2
Query: 473 MDKRMYRHDNSIGFGQWVPNRRVPFEEGFEARRPA 507
MD+ Y+ ++ FG W N +PF + FE+ R A
Sbjct: 74 MDEYYYKRNHVPAFGSWDWNDNLPFTQCFESARQA 178
>AV415782
Length = 170
Score = 28.9 bits (63), Expect = 3.4
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = -3
Query: 767 DGGRTASAWWRAIATLHSEAWFG 789
DGGR A +WWRA+A + W G
Sbjct: 105 DGGRVA*SWWRAVA---EDTW*G 46
>TC15045 similar to AAR25638 (AAR25638) At1g55300, partial (66%)
Length = 1050
Score = 28.9 bits (63), Expect = 3.4
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = -2
Query: 767 DGGRTASAWWRAIATLHSEAWFG 789
DGGR A +WWRA+A + W G
Sbjct: 98 DGGRVA*SWWRAVA---EDTW*G 39
>TC15974 similar to UP|Q94KL4 (Q94KL4) BEL1-like homeodomain 1, partial (5%)
Length = 742
Score = 27.3 bits (59), Expect = 10.0
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +2
Query: 745 EKESLWYRVLVARYGEEGGRLLDGGRTASAWW 776
E ESLW R+ + G EGG ++G + +W
Sbjct: 620 ELESLWLRLYIIFVGSEGGNGIEGLMS*ILFW 715
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.361 0.164 0.647
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,527,977
Number of Sequences: 28460
Number of extensions: 276120
Number of successful extensions: 2654
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1948
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 659
Number of HSP's gapped (non-prelim): 2064
length of query: 820
length of database: 4,897,600
effective HSP length: 98
effective length of query: 722
effective length of database: 2,108,520
effective search space: 1522351440
effective search space used: 1522351440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC136451.10


