

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
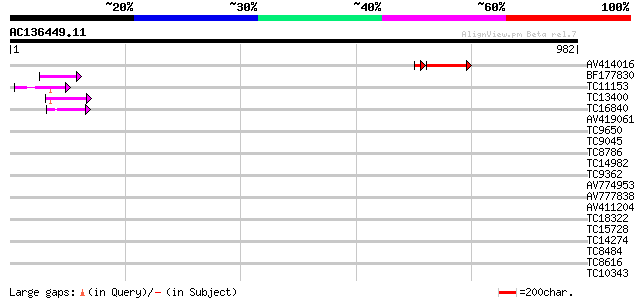
Query= AC136449.11 - phase: 0 /pseudo
(982 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV414016 150 3e-41
BF177830 62 4e-10
TC11153 similar to UP|Q9FJJ6 (Q9FJJ6) Genomic DNA, chromosome 5,... 60 2e-09
TC13400 similar to UP|BAD05829 (BAD05829) Zinc finger (C3HC4-typ... 52 5e-07
TC16840 homologue to UP|Q9SHZ4 (Q9SHZ4) At2g22120 protein, parti... 42 5e-04
AV419061 39 0.003
TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogen... 34 0.13
TC9045 similar to UP|O81373 (O81373) Cotton fiber expressed prot... 34 0.13
TC8786 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, pa... 33 0.17
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 33 0.17
TC9362 similar to PIR|F86340|F86340 protein F2D10.34 [imported] ... 33 0.17
AV774953 33 0.22
AV777838 33 0.22
AV411204 33 0.29
TC18322 similar to GB|AAP13390.1|30023714|BT006282 At5g44170 {Ar... 33 0.29
TC15728 similar to AAS09986 (AAS09986) MYB transcription factor,... 33 0.29
TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.3... 33 0.29
TC8484 similar to UP|Q9XGP1 (Q9XGP1) ESTs AU070372(S13446), part... 33 0.29
TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like pro... 32 0.37
TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (... 32 0.37
>AV414016
Length = 417
Score = 150 bits (379), Expect(2) = 3e-41
Identities = 73/79 (92%), Positives = 77/79 (97%)
Frame = +2
Query: 722 YVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 781
YVIWTAVAG+RYS++QIRKRR SVLL Q+WKWCSIVVKSSALLSIWIFVIPVLIGLLFEL
Sbjct: 179 YVIWTAVAGLRYSLDQIRKRRISVLLGQVWKWCSIVVKSSALLSIWIFVIPVLIGLLFEL 358
Query: 782 LVIVPMRVPVDESPVFLLY 800
LVIVPMRVPVDESPVFLLY
Sbjct: 359 LVIVPMRVPVDESPVFLLY 415
Score = 36.2 bits (82), Expect(2) = 3e-41
Identities = 18/20 (90%), Positives = 18/20 (90%)
Frame = +3
Query: 701 FALFFCWSLLG*LSLSSTLP 720
FALFF W LLG*LSLSSTLP
Sbjct: 9 FALFFYW*LLG*LSLSSTLP 68
>BF177830
Length = 476
Score = 62.0 bits (149), Expect = 4e-10
Identities = 25/73 (34%), Positives = 41/73 (55%)
Frame = +2
Query: 52 APPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQ 111
AP + D++ +EE VCRIC + D N + C+C G + VH++CL++W + ++
Sbjct: 182 APVESAADEEIPEEEAVCRICFDVCDERNTFKMECSCKGDLTLVHEECLIKWFSLKGNKK 361
Query: 112 CEVCKHPFSFSPV 124
C+VC PV
Sbjct: 362 CDVCGQEVQNLPV 400
>TC11153 similar to UP|Q9FJJ6 (Q9FJJ6) Genomic DNA, chromosome 5, TAC
clone:K19B1 (AT5g62460/K19B1_7), partial (17%)
Length = 451
Score = 60.1 bits (144), Expect = 2e-09
Identities = 37/103 (35%), Positives = 50/103 (47%), Gaps = 5/103 (4%)
Frame = +1
Query: 8 PPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAK-YDDDDEDEE 66
P +D A PS SPS ++ A A + TAPP+ + +D DEDE
Sbjct: 142 PVPVDPVAQPAQLPSEPSPSLDAA-----------AAAAGPSGTAPPADQDVEDADEDEP 288
Query: 67 DV----CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLN 105
+ CRIC+ N L PCACSGS+K+ H+ C+ W N
Sbjct: 289 LIQMAECRICQEEDQVSN-LETPCACSGSLKYAHRKCVQHWCN 414
>TC13400 similar to UP|BAD05829 (BAD05829) Zinc finger (C3HC4-type RING
finger) protein-like, partial (25%)
Length = 553
Score = 52.0 bits (123), Expect = 5e-07
Identities = 32/92 (34%), Positives = 45/92 (48%), Gaps = 12/92 (13%)
Frame = +2
Query: 63 EDEEDV-------CRIC-RNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNA---RQ 111
+DEEDV CRIC + D ++ L PC C G+ +FVH+ CL W +
Sbjct: 248 KDEEDVEAGSLPCCRICLESDSDPEDELISPCMCKGTQQFVHRSCLDHWRSVKEGFAFSH 427
Query: 112 CEVCKHPFSFS-PVYAENAPARLPFQEFVVGM 142
C CK F Y +N+ ++ F+ FV GM
Sbjct: 428 CTTCKAQFHLRVESYEDNSWRKIKFRLFVEGM 523
>TC16840 homologue to UP|Q9SHZ4 (Q9SHZ4) At2g22120 protein, partial (39%)
Length = 576
Score = 42.0 bits (97), Expect = 5e-04
Identities = 27/79 (34%), Positives = 34/79 (42%), Gaps = 4/79 (5%)
Frame = +3
Query: 65 EEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNA---RQCEVCKHPFSF 121
E+ CRIC D PC C G+ K+VH+DCL W C CK P+
Sbjct: 258 EQIQCRICLETDGRD--FIAPCKCKGTSKYVHRDCLDHWRAIKEGFAFAHCTTCKAPYHL 431
Query: 122 S-PVYAENAPARLPFQEFV 139
V A+ L F+ FV
Sbjct: 432 RVHVAADRKWRTLKFRFFV 488
>AV419061
Length = 407
Score = 39.3 bits (90), Expect = 0.003
Identities = 27/83 (32%), Positives = 36/83 (42%), Gaps = 8/83 (9%)
Frame = +2
Query: 65 EEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNA---RQCEVCKHPFSF 121
++ CRIC + G D L PC C G+ K+VH+ CL W + C C+ F
Sbjct: 86 DQPQCRICLDIGGED--LIAPCHCKGTQKYVHRSCLDNWRSTKEGFAFSHCTECRAVF-- 253
Query: 122 SPVYAENAP-----ARLPFQEFV 139
+ N P RL FQ V
Sbjct: 254 --ILRANVPPDRWWLRLKFQFLV 316
>TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogenase-like
protein , partial (47%)
Length = 1041
Score = 33.9 bits (76), Expect = 0.13
Identities = 21/58 (36%), Positives = 29/58 (49%), Gaps = 8/58 (13%)
Frame = +2
Query: 4 ANEPPPSLDGTPIAATTPSSSSPSSS--------SSSPRGSKGKEIDAEAVATASTAP 53
++ PPPS + AT P+ SS SS SSSPR S + ++ + STAP
Sbjct: 98 SHSPPPSPSPSAATATVPAPSSSSSPPPSTPNPPSSSPRSSAKRVSNSSRTSPTSTAP 271
>TC9045 similar to UP|O81373 (O81373) Cotton fiber expressed protein 1,
partial (17%)
Length = 1201
Score = 33.9 bits (76), Expect = 0.13
Identities = 22/54 (40%), Positives = 31/54 (56%), Gaps = 6/54 (11%)
Frame = -1
Query: 7 PPPSLDGTPIAATTPSSSSPSSSS--SSPRG----SKGKEIDAEAVATASTAPP 54
PPPS +P T+P+SSSP S+ ++PR S I + A +TAS +PP
Sbjct: 610 PPPS---SPCRRTSPTSSSPDRSNPPAAPRSAAAESPSSPISSSASSTASLSPP 458
>TC8786 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(33%)
Length = 594
Score = 33.5 bits (75), Expect = 0.17
Identities = 19/55 (34%), Positives = 28/55 (50%)
Frame = +1
Query: 4 ANEPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKY 58
AN PPP + A T PSS+ P++ S+P S I A +++ PP A +
Sbjct: 112 ANPPPPP---STAARTPPSSAPPATPKSTPPTSSPPAIHVSLSAKSASKPPLASH 267
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 33.5 bits (75), Expect = 0.17
Identities = 20/47 (42%), Positives = 23/47 (48%)
Frame = +3
Query: 9 PSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPS 55
PS AA+ PS+S SSSSSS S +TA T PPS
Sbjct: 159 PSTAAPAAAASAPSASG*SSSSSSSSSSSAVLASPSTSSTAPTTPPS 299
>TC9362 similar to PIR|F86340|F86340 protein F2D10.34 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(58%)
Length = 591
Score = 33.5 bits (75), Expect = 0.17
Identities = 20/48 (41%), Positives = 24/48 (49%)
Frame = +2
Query: 7 PPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPP 54
PPP L P + T SSSSP SS+ S S A V+ A +PP
Sbjct: 62 PPPHLIPPPPSTPTSSSSSPLSSALSSASSAWSPSPAAPVSGAYASPP 205
>AV774953
Length = 579
Score = 33.1 bits (74), Expect = 0.22
Identities = 23/77 (29%), Positives = 35/77 (44%)
Frame = +1
Query: 4 ANEPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDE 63
A PP + P + SSS S ++ S +E AE+ A +TA P+ D DD+
Sbjct: 271 AKSTPPPAETDPPKKRSRKSSSGSEKDNTETESASEEGKAESAAEENTADPN---DADDK 441
Query: 64 DEEDVCRICRNPGDADN 80
E D ++ + DN
Sbjct: 442 PETDAEETKQSNVEGDN 492
>AV777838
Length = 565
Score = 33.1 bits (74), Expect = 0.22
Identities = 26/60 (43%), Positives = 28/60 (46%), Gaps = 7/60 (11%)
Frame = +1
Query: 4 ANEPPPSLDGTPIAA-----TTP--SSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSA 56
A PPP +AA T P SSSS SSSSSS S A T TAPPS+
Sbjct: 145 AKPPPPRASSRELAAGHTTITGPPSSSSSSSSSSSSSSSSSSSMRATAAGHTTITAPPSS 324
>AV411204
Length = 427
Score = 32.7 bits (73), Expect = 0.29
Identities = 26/94 (27%), Positives = 43/94 (45%), Gaps = 3/94 (3%)
Frame = +2
Query: 27 SSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRN---PGDADNPLR 83
+ + SS RG+ AV+ + P + ++ E VC IC++ PG N L
Sbjct: 113 AENDSSRRGAP-----PAAVSFVNNLPRIVIGKEHEKHGELVCAICKDVLAPGTEVNQL- 274
Query: 84 YPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKH 117
PC+ H C+L WL+ N+ C +C++
Sbjct: 275 -PCS-----HLYHPSCILPWLSARNS--CPLCRY 352
>TC18322 similar to GB|AAP13390.1|30023714|BT006282 At5g44170 {Arabidopsis
thaliana;}, partial (41%)
Length = 427
Score = 32.7 bits (73), Expect = 0.29
Identities = 19/46 (41%), Positives = 25/46 (54%), Gaps = 1/46 (2%)
Frame = +1
Query: 14 TPIAATTPSSSSPSSSSSSPR-GSKGKEIDAEAVATASTAPPSAKY 58
TP + PS +PSSSS+SP G K T+ST+P SA +
Sbjct: 94 TPCTSEPPSGHAPSSSSNSPNAGPKPHPKPPTLTPTSSTSPASAPW 231
Score = 28.1 bits (61), Expect = 7.1
Identities = 15/51 (29%), Positives = 30/51 (58%)
Frame = +1
Query: 8 PPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKY 58
PP+L TP ++T+P+S+ SS++++ + A ++ T+PPS +
Sbjct: 181 PPTL--TPTSSTSPASAPWSSAAAAASPAWASTSSASLT*SSPTSPPSCPH 327
>TC15728 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(34%)
Length = 1207
Score = 32.7 bits (73), Expect = 0.29
Identities = 15/31 (48%), Positives = 21/31 (67%)
Frame = +1
Query: 1 MEIANEPPPSLDGTPIAATTPSSSSPSSSSS 31
+++ PPPS D +P ++ S SSPSSSSS
Sbjct: 958 LKLQPSPPPSKDHSPATSSHSSPSSPSSSSS 1050
>TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.35
{Arabidopsis thaliana;} , partial (16%)
Length = 1208
Score = 32.7 bits (73), Expect = 0.29
Identities = 20/48 (41%), Positives = 26/48 (53%)
Frame = +3
Query: 9 PSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSA 56
PS TP P+ SSP S+SSSP S I + + S+APP+A
Sbjct: 276 PSRSSTPNPPIPPNPSSPPSASSSPT-STTSMIPPASANSPSSAPPAA 416
>TC8484 similar to UP|Q9XGP1 (Q9XGP1) ESTs AU070372(S13446), partial (76%)
Length = 1121
Score = 32.7 bits (73), Expect = 0.29
Identities = 23/66 (34%), Positives = 33/66 (49%), Gaps = 14/66 (21%)
Frame = +1
Query: 4 ANEPPPSLDGTPIA----------ATTPSSSSPSSSSSSPRGSKGKEIDAEA----VATA 49
++ PPPS +P A +TTP+ S PS S +SPRG + + A +A
Sbjct: 235 SHPPPPSPPPSPSATNSLNPPYPTSTTPARSKPSPSLNSPRGRRQSSSPSPARSPRLARR 414
Query: 50 STAPPS 55
ST+P S
Sbjct: 415 STSPGS 432
>TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like protein III,
complete
Length = 2302
Score = 32.3 bits (72), Expect = 0.37
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = +1
Query: 882 YPLVVNSAVYRFAWLGCLSFSFVCFCAKR 910
+ ++ N++ F CLSFSFVCFC +
Sbjct: 2155 FSILSNTSFVAFVAAACLSFSFVCFCCMK 2241
>TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (Fibrillarin
2) (AtFib2), partial (85%)
Length = 1116
Score = 32.3 bits (72), Expect = 0.37
Identities = 19/56 (33%), Positives = 31/56 (54%)
Frame = -1
Query: 1 MEIANEPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSA 56
+ + N PSL + A T SS++P+SSSSS S K + + +++S PS+
Sbjct: 258 VRLNNNSAPSLHSSSSATTASSSATPASSSSSTATSCLKRGTSTSSSSSSITSPSS 91
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.141 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,102,970
Number of Sequences: 28460
Number of extensions: 300005
Number of successful extensions: 4722
Number of sequences better than 10.0: 166
Number of HSP's better than 10.0 without gapping: 3439
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4218
length of query: 982
length of database: 4,897,600
effective HSP length: 99
effective length of query: 883
effective length of database: 2,080,060
effective search space: 1836692980
effective search space used: 1836692980
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC136449.11


