

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136286.2 - phase: 0
(251 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
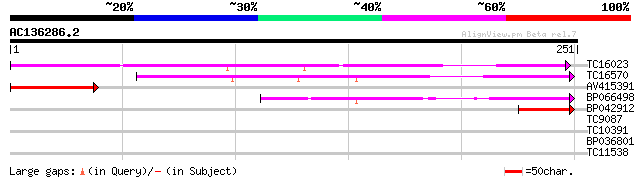
TC16023 174 1e-44
TC16570 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (18%) 144 2e-35
AV415391 79 6e-16
BP066498 76 6e-15
BP042912 43 5e-05
TC9087 homologue to UP|AAR83868 (AAR83868) 60S ribosomal protein... 30 0.52
TC10391 26 5.7
BP036801 26 5.7
TC11538 25 9.7
>TC16023
Length = 894
Score = 174 bits (442), Expect = 1e-44
Identities = 103/253 (40%), Positives = 135/253 (52%), Gaps = 5/253 (1%)
Frame = +2
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP PR++ DML+LP GN++I+NGA +G AG++NA N L P LY P +F +
Sbjct: 41 MEYMPEPRLLHDMLILPNGNILIINGAKHGCAGYDNARNASLEPYLYSP-KKRLGKRFRV 217
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDF-QAKYPTELSLDAYYPDYLRPELDT 119
L RMYHSSA LLPDGR+LV G NPH Y F YPTEL L A+ P Y+ +
Sbjct: 218 LKSTKIARMYHSSATLLPDGRVLVAGGNPHGRYSFHNVAYPTELRLQAFVPHYMHRRYHS 397
Query: 120 LRPVIVAVEV----VNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRL 175
RP + V+ + Y + F V FL+ + N + S+ AP FTTHS AMNQR+
Sbjct: 398 WRPSNLTVKSGGGGERGVIGYGAEFRVRFLV-GTRPSNEVAFSVYAPPFTTHSVAMNQRM 574
Query: 176 LFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLF 235
L L ++ D A + PPS VAP GYY+L
Sbjct: 575 LKLRCRSMVRSSGGWVD-----------------------AVLEAPPSPRVAPSGYYLLT 685
Query: 236 VIHVGIPSVATWV 248
V++ GIPSV+ W+
Sbjct: 686 VVNGGIPSVSQWI 724
>TC16570 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (18%)
Length = 794
Score = 144 bits (362), Expect = 2e-35
Identities = 80/197 (40%), Positives = 112/197 (56%), Gaps = 3/197 (1%)
Frame = +3
Query: 57 KFELLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQA-KYPTELSLDAYYPDYLRP 115
+FEL P TP MYHS+AVLL DGR+LV GSNPH Y+F + +PTELSL+A+ P YL P
Sbjct: 33 QFELQNPTKTPWMYHSTAVLLRDGRVLVAGSNPHEYYNFSSVLFPTELSLEAFSPPYLEP 212
Query: 116 ELDTLRPVIVA-VEVVNSTLSYESLFSVSFLLREVKDV-NRIRVSMVAPSFTTHSFAMNQ 173
+ +RP IV+ + L Y + F + + + + ++M+AP F THSF+MNQ
Sbjct: 213 QYTNVRPRIVSPTPQAQTKLKYGQGLRIRFQVTALLAAPDSVSMTMLAPPFNTHSFSMNQ 392
Query: 174 RLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYM 233
RLL LE L++V S+++ V P S +APPG+Y+
Sbjct: 393 RLLVLEPNGLKDV-----------------------GESIHEVEVTAPGSATLAPPGFYL 503
Query: 234 LFVIHVGIPSVATWVHV 250
+FV+H GIPS WV +
Sbjct: 504 VFVVHQGIPSEGIWVQI 554
>AV415391
Length = 138
Score = 79.3 bits (194), Expect = 6e-16
Identities = 35/39 (89%), Positives = 38/39 (96%)
Frame = +2
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAAN 39
ME MP+PRVMPDM+LLPTGNV+ILNGAANGTAGWENAAN
Sbjct: 20 METMPIPRVMPDMILLPTGNVMILNGAANGTAGWENAAN 136
>BP066498
Length = 512
Score = 75.9 bits (185), Expect = 6e-15
Identities = 49/141 (34%), Positives = 71/141 (49%), Gaps = 2/141 (1%)
Frame = -1
Query: 112 YLRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDV--NRIRVSMVAPSFTTHSF 169
YL + D P+I E L Y F F +++ ++ N I+V+M P FTTH F
Sbjct: 512 YLDVDFDIYXPMINEDESTKE-LRYGQFFETVFAVQDGAELTMNDIKVTMYFPPFTTHGF 336
Query: 170 AMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPP 229
+M QRLL L++ L +V+S PG +Y+ + PP+ +APP
Sbjct: 335 SMGQRLLVLKLDKL--MVDS----------------PG-----LYRVRIAAPPNSVIAPP 225
Query: 230 GYYMLFVIHVGIPSVATWVHV 250
GYY+LFV H G+P WVH+
Sbjct: 224 GYYLLFVNHRGVPGKGMWVHI 162
>BP042912
Length = 223
Score = 43.1 bits (100), Expect = 5e-05
Identities = 15/25 (60%), Positives = 20/25 (80%)
Frame = -3
Query: 226 VAPPGYYMLFVIHVGIPSVATWVHV 250
+AP GYY+LFV+H G+PS +WV V
Sbjct: 209 IAPAGYYLLFVVHAGVPSTGSWVQV 135
>TC9087 homologue to UP|AAR83868 (AAR83868) 60S ribosomal protein L12,
complete
Length = 787
Score = 29.6 bits (65), Expect = 0.52
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = -3
Query: 14 LLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNP 54
LLL ++ +N AANG AG EN N + ++P L +P
Sbjct: 521 LLLQILRILAVNSAANGDAGPENLLNGAGEILRHRPRLHHP 399
>TC10391
Length = 565
Score = 26.2 bits (56), Expect = 5.7
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = -1
Query: 113 LRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIR 156
LRP + RP+ V+ + TLSY+ +S + + E+ RIR
Sbjct: 298 LRPRVMVTRPLEVSDMSITQTLSYQ--WSGNLCINEMNTATRIR 173
>BP036801
Length = 540
Score = 26.2 bits (56), Expect = 5.7
Identities = 14/28 (50%), Positives = 19/28 (67%)
Frame = +1
Query: 59 ELLAPASTPRMYHSSAVLLPDGRILVGG 86
EL ++PR Y S+ +L PDGRI+V G
Sbjct: 457 ELPQNLTSPRWYASNQIL-PDGRIIVVG 537
>TC11538
Length = 542
Score = 25.4 bits (54), Expect = 9.7
Identities = 13/29 (44%), Positives = 15/29 (50%)
Frame = -3
Query: 68 RMYHSSAVLLPDGRILVGGSNPHRLYDFQ 96
R Y LLP +L G SNPH Y+ Q
Sbjct: 420 RSYWKHRWLLPHNILLPGISNPHIQYNLQ 334
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,024,333
Number of Sequences: 28460
Number of extensions: 52365
Number of successful extensions: 255
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 245
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 246
length of query: 251
length of database: 4,897,600
effective HSP length: 88
effective length of query: 163
effective length of database: 2,393,120
effective search space: 390078560
effective search space used: 390078560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC136286.2


