

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136142.6 + phase: 0
(147 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
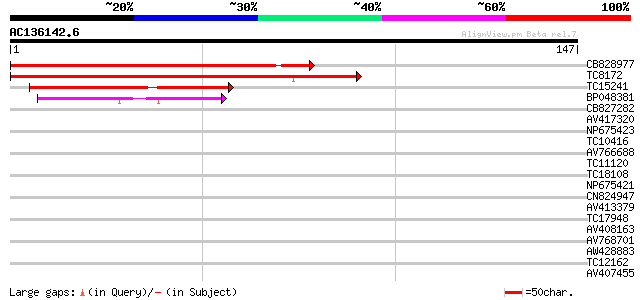
Score E
Sequences producing significant alignments: (bits) Value
CB828977 97 1e-21
TC8172 similar to AAS10008 (AAS10008) MYB transcription factor, ... 96 3e-21
TC15241 weakly similar to UP|AAR85480 (AAR85480) ANTHER INDEHISC... 49 5e-07
BP048381 38 6e-04
CB827282 28 0.001
AV417320 36 0.002
NP675423 transcription factor MYB101 [Lotus japonicus] 35 0.005
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 34 0.012
AV766688 34 0.012
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 33 0.015
TC18108 similar to UP|Q08707 (Q08707) Box-P binding factor 1 (BP... 33 0.020
NP675421 transcription factor MYB103 [Lotus japonicus] 32 0.044
CN824947 32 0.058
AV413379 31 0.099
TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose... 30 0.17
AV408163 30 0.22
AV768701 30 0.22
AW428883 28 0.49
TC12162 similar to AAS09999 (AAS09999) MYB transcription factor,... 28 0.49
AV407455 28 0.64
>CB828977
Length = 528
Score = 97.1 bits (240), Expect = 1e-21
Identities = 49/79 (62%), Positives = 57/79 (72%)
Frame = +1
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MG KQKWTAEEE AL GV K+GAGKW+ IL DPEF+ L +RSN+DLKDKWRN+NV
Sbjct: 52 MGNQKQKWTAEEEDALHRGVQKYGAGKWKNILKDPEFAPSLTSRSNIDLKDKWRNLNVGT 231
Query: 61 IWGSRQKAKLALKNSPPAP 79
GS K++ LK PAP
Sbjct: 232 GQGSNVKSR-TLKPKLPAP 285
>TC8172 similar to AAS10008 (AAS10008) MYB transcription factor, partial
(30%)
Length = 1353
Score = 95.9 bits (237), Expect = 3e-21
Identities = 50/92 (54%), Positives = 61/92 (65%), Gaps = 1/92 (1%)
Frame = +1
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MG KQKWTAEEE AL GV K+GAGKW+ IL DPEF+ L +RSN+DLKDKWRN+NV
Sbjct: 100 MGNQKQKWTAEEEDALHRGVQKYGAGKWKNILKDPEFAPSLTSRSNIDLKDKWRNLNVVT 279
Query: 61 IWGSRQKAKLAL-KNSPPAPKTDNNQLALGKV 91
GS K++ + K + P+ N Q A V
Sbjct: 280 GQGSNIKSRTSKPKLTAPSTPVPNQQNATPSV 375
>TC15241 weakly similar to UP|AAR85480 (AAR85480) ANTHER INDEHISCENCE1,
partial (13%)
Length = 655
Score = 48.5 bits (114), Expect = 5e-07
Identities = 23/53 (43%), Positives = 34/53 (63%)
Frame = +2
Query: 6 QKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINV 58
++W+ E+ L AGV K G G W IL + + + R++VDLKDKWRN+N+
Sbjct: 200 KRWSQLEKETLLAGVNKFGEGNWTFILSTHK--DVFKGRTSVDLKDKWRNMNL 352
>BP048381
Length = 513
Score = 38.1 bits (87), Expect = 6e-04
Identities = 24/55 (43%), Positives = 31/55 (55%), Gaps = 6/55 (10%)
Frame = -1
Query: 8 WTAEEEAALKAGVVKHGAGK----WRTILMDPEF--SSILRTRSNVDLKDKWRNI 56
WT +EE ALK GV+K WR IL EF + +TR+ DLKDKW+ +
Sbjct: 480 WTDDEEKALKEGVLKFSLENQNIPWRKIL---EFGCGTFDKTRAPSDLKDKWKKM 325
>CB827282
Length = 533
Score = 28.5 bits (62), Expect(2) = 0.001
Identities = 15/44 (34%), Positives = 23/44 (52%)
Frame = +2
Query: 6 QKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDL 49
++W+ E+ L AGV K G G W IL + + R++V L
Sbjct: 62 KRWSQLEKETLLAGVNKFGEGNWTFIL--STHKDVFKGRTSVSL 187
Score = 27.7 bits (60), Expect(2) = 0.001
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = +3
Query: 47 VDLKDKWRNINV 58
VDLKDKWRN+N+
Sbjct: 270 VDLKDKWRNMNL 305
>AV417320
Length = 369
Score = 36.2 bits (82), Expect = 0.002
Identities = 19/55 (34%), Positives = 30/55 (54%)
Frame = +3
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G K WT+EE+ L + + KHG G WR + P+ + +LR + L +W N
Sbjct: 105 IGLKKGPWTSEEDQILISYIQKHGHGNWRAL---PKHAGLLRCGKSCRL--RWIN 254
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 35.0 bits (79), Expect = 0.005
Identities = 19/55 (34%), Positives = 28/55 (50%)
Frame = +1
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G K WT EE+A L + KHG G WR + P+ + + R + L +W N
Sbjct: 28 IGLKKGPWTPEEDAILVDYIQKHGHGSWRAL---PKLAGLNRCGKSCRL--RWTN 177
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 33.9 bits (76), Expect = 0.012
Identities = 19/54 (35%), Positives = 28/54 (51%)
Frame = +2
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WTAEE+ L A V ++G WR + P+F+ + R + L +W N
Sbjct: 122 GMRKGTWTAEEDKKLIAYVTRYGCWNWRQL---PKFAGLARCGKSCRL--RWLN 268
>AV766688
Length = 608
Score = 33.9 bits (76), Expect = 0.012
Identities = 19/54 (35%), Positives = 28/54 (51%)
Frame = +2
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WTAEE+ L A V ++G WR + P+F+ + R + L +W N
Sbjct: 107 GLRKGTWTAEEDRKLIAYVTRYGCWNWRQL---PKFAGLARCGKSCRL--RWLN 253
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 33.5 bits (75), Expect = 0.015
Identities = 18/54 (33%), Positives = 26/54 (47%)
Frame = +1
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WT EE+ L + KHG G WR + P+ + + R + L +W N
Sbjct: 97 GLKKGPWTPEEDQKLVEHIQKHGHGSWRAL---PKLAGLNRCGKSCRL--RWTN 243
>TC18108 similar to UP|Q08707 (Q08707) Box-P binding factor 1 (BPF-1),
partial (15%)
Length = 700
Score = 33.1 bits (74), Expect = 0.020
Identities = 20/56 (35%), Positives = 31/56 (54%)
Frame = +1
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
++ +T E AL V + G G+WR + + ++ RT VDLKDKW+ + TA
Sbjct: 40 RRPFTVYEVEALVHAVEELGTGRWRDVKLRSFENADPRTY--VDLKDKWKTLVHTA 201
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 32.0 bits (71), Expect = 0.044
Identities = 17/54 (31%), Positives = 26/54 (47%)
Frame = +1
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K W+ EE+ L + KHG G WR + P+ + + R + L +W N
Sbjct: 31 GMKKGPWSQEEDKVLVDYIQKHGHGSWRAL---PQLAGLNRCGKSCRL--RWTN 177
>CN824947
Length = 738
Score = 31.6 bits (70), Expect = 0.058
Identities = 17/55 (30%), Positives = 29/55 (51%)
Frame = +1
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G + +WTAEE+ L + G G WR++ P+ + +LR + L +W N
Sbjct: 178 VGLKRGRWTAEEDELLTKYIQASGEGSWRSL---PKNAGLLRCGKSCRL--RWIN 327
>AV413379
Length = 416
Score = 30.8 bits (68), Expect = 0.099
Identities = 18/51 (35%), Positives = 27/51 (52%)
Frame = +3
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
K WT EE+ L A + HG G WR++ P+ + +LR + L +W N
Sbjct: 141 KGAWTKEEDDRLIAYIRAHGEGCWRSL---PKSAGLLRCGKSCRL--RWIN 278
>TC17948 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (50%)
Length = 487
Score = 30.0 bits (66), Expect = 0.17
Identities = 17/51 (33%), Positives = 24/51 (46%)
Frame = +2
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
K W+AEE+ L V +HGA W I S ++ RS + +W N
Sbjct: 170 KGPWSAEEDRILTRLVERHGARNWSLI------SRYIKGRSGKSCRLRWCN 304
>AV408163
Length = 425
Score = 29.6 bits (65), Expect = 0.22
Identities = 17/51 (33%), Positives = 27/51 (52%)
Frame = +2
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
K WT EE+ L + + HG G WR++ P+ + +LR + L +W N
Sbjct: 116 KGAWTKEEDDRLISYIRAHGEGCWRSL---PKAAGLLRCGKSCRL--RWIN 253
>AV768701
Length = 292
Score = 29.6 bits (65), Expect = 0.22
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = -3
Query: 32 LMDPEFSSILRTRSNVDLKDKWRNI 56
L++ S + R+N DLKDKWRN+
Sbjct: 284 LIENFHKSTFKDRTNKDLKDKWRNL 210
>AW428883
Length = 430
Score = 28.5 bits (62), Expect = 0.49
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +1
Query: 8 WTAEEEAALKAGVVKHGAGKWRTI 31
WT EE G+ K+G G WR+I
Sbjct: 139 WTEEEHRLFLLGLDKYGKGDWRSI 210
>TC12162 similar to AAS09999 (AAS09999) MYB transcription factor, partial
(56%)
Length = 751
Score = 28.5 bits (62), Expect = 0.49
Identities = 11/24 (45%), Positives = 13/24 (53%)
Frame = +1
Query: 8 WTAEEEAALKAGVVKHGAGKWRTI 31
WT EE G+ K+G G WR I
Sbjct: 580 WTEEEHRLFLLGLKKYGKGDWRNI 651
>AV407455
Length = 399
Score = 28.1 bits (61), Expect = 0.64
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = -1
Query: 52 KWRNINVTAIWGSRQKAKLALK 73
+WR++ V IWGS ++ + +LK
Sbjct: 141 RWRSVGVDGIWGSGERTEWSLK 76
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,562,245
Number of Sequences: 28460
Number of extensions: 30771
Number of successful extensions: 146
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 144
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 144
length of query: 147
length of database: 4,897,600
effective HSP length: 82
effective length of query: 65
effective length of database: 2,563,880
effective search space: 166652200
effective search space used: 166652200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 51 (24.3 bits)
Medicago: description of AC136142.6


