

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.5 - phase: 0
(736 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
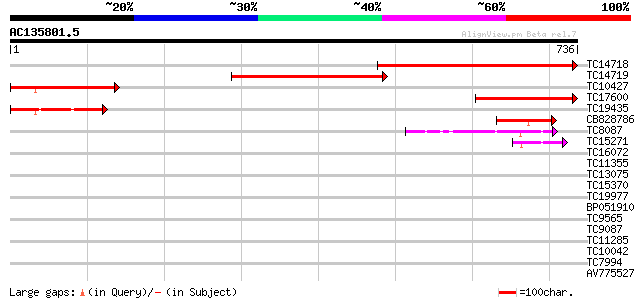
Sequences producing significant alignments: (bits) Value
TC14718 similar to UP|NCPR_CATRO (Q05001) NADPH--cytochrome P450... 499 e-142
TC14719 similar to UP|Q9AU07 (Q9AU07) NADPH-cytochrome P450 oxyd... 363 e-101
TC10427 similar to UP|Q9AU06 (Q9AU06) NADPH-cytochrome P450 oxyd... 218 3e-57
TC17600 similar to PIR|A47298|A47298 NADPH-ferrihemoprotein redu... 208 2e-54
TC19435 similar to UP|Q39036 (Q39036) NADPH-ferrihemoprotein red... 170 7e-43
CB828786 144 7e-35
TC8087 homologue to UP|FENR_PEA (P10933) Ferredoxin--NADP reduct... 60 1e-09
TC15271 similar to PIR|T06773|T06773 ferredoxin-NADP reductase ... 44 1e-04
TC16072 32 0.049
TC11355 33 0.16
TC13075 similar to UP|O23442 (O23442) Dynein light chain like pr... 30 1.1
TC15370 weakly similar to UP|Q8S8R0 (Q8S8R0) Predicted protein, ... 30 1.4
TC19977 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (20%) 29 2.4
BP051910 29 2.4
TC9565 similar to UP|Q9MAQ2 (Q9MAQ2) CDS, partial (19%) 29 3.1
TC9087 homologue to UP|AAR83868 (AAR83868) 60S ribosomal protein... 29 3.1
TC11285 28 4.0
TC10042 28 5.3
TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomeras... 28 5.3
AV775527 28 6.9
>TC14718 similar to UP|NCPR_CATRO (Q05001) NADPH--cytochrome P450 reductase
(CPR) (P450R) , partial (36%)
Length = 1163
Score = 499 bits (1284), Expect = e-142
Identities = 239/259 (92%), Positives = 253/259 (97%)
Frame = +1
Query: 478 WVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVH 537
WVIASQRSLLEVMAEF SAKPPIGVFFA++APRLQPR+YSISSSPR+APSRIHVTCALV+
Sbjct: 1 WVIASQRSLLEVMAEFPSAKPPIGVFFAAIAPRLQPRFYSISSSPRMAPSRIHVTCALVN 180
Query: 538 DKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGL 597
DKMPTGRIH+GVCSTWMKNS PLEKSQDCSWAPIFVRQSNF+LPADNKVPIIMIGPGTGL
Sbjct: 181 DKMPTGRIHRGVCSTWMKNSVPLEKSQDCSWAPIFVRQSNFKLPADNKVPIIMIGPGTGL 360
Query: 598 APFRGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSELIVAFSR 657
APFRGFLQERLALKE+GAELGPSVLFFGCRNRQ+DYIYEDELNHFV+ GALSELIVAFSR
Sbjct: 361 APFRGFLQERLALKEDGAELGPSVLFFGCRNRQMDYIYEDELNHFVNSGALSELIVAFSR 540
Query: 658 EGPTKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSK 717
EGPTKEYVQHKM+EKASDIWNMISQGAYIYVCGDAKGMA+DVHRTLHTILQEQGSLD+SK
Sbjct: 541 EGPTKEYVQHKMMEKASDIWNMISQGAYIYVCGDAKGMARDVHRTLHTILQEQGSLDSSK 720
Query: 718 TESMVKNLQMTGRYLRDVW 736
E MVKNLQ+ GRYLRDVW
Sbjct: 721 AEGMVKNLQLNGRYLRDVW 777
>TC14719 similar to UP|Q9AU07 (Q9AU07) NADPH-cytochrome P450 oxydoreductase
isoform 2, partial (28%)
Length = 633
Score = 363 bits (933), Expect = e-101
Identities = 177/202 (87%), Positives = 189/202 (92%)
Frame = +3
Query: 289 EDDATVATPYTASVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELH 348
+DD TV+TPYTA+VLEYRVVI D LDA+VDEKK N NGHA+VDA HPVR+NVAVRKELH
Sbjct: 27 DDDTTVSTPYTAAVLEYRVVIHDPLDASVDEKKWHNVNGHAIVDAQHPVRSNVAVRKELH 206
Query: 349 TPASDRSCTHLEFDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDD 408
TP SDRSCTHLEFDISGTGV YETGDHVGVYCENLS+TVEEA R+LGLSPDTYFSVHTDD
Sbjct: 207 TPVSDRSCTHLEFDISGTGVAYETGDHVGVYCENLSETVEEAVRLLGLSPDTYFSVHTDD 386
Query: 409 EDGKPLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLAS 468
EDGKPL GSSLPP FPPCTLRTA+A+YADVLSSPKKS LLALAAHAS+PSEADRLRHLAS
Sbjct: 387 EDGKPLSGSSLPPTFPPCTLRTAIARYADVLSSPKKSGLLALAAHASNPSEADRLRHLAS 566
Query: 469 PAGKDEYAEWVIASQRSLLEVM 490
PAGKDEY+EWVIASQRSLLEVM
Sbjct: 567 PAGKDEYSEWVIASQRSLLEVM 632
>TC10427 similar to UP|Q9AU06 (Q9AU06) NADPH-cytochrome P450 oxydoreductase
isoform 3, partial (17%)
Length = 565
Score = 218 bits (555), Expect = 3e-57
Identities = 112/146 (76%), Positives = 125/146 (84%), Gaps = 4/146 (2%)
Frame = +3
Query: 1 MQDSSSMKFSPLDLMSALIKGKIDPSNGTVPA---SLILENREFVMILTTSIAVLIGCVV 57
M++SSSMK SPLDLMSA+IKG +DPSN + + S+ LENREFVM+LTTSIAVLIGCVV
Sbjct: 126 MEESSSMKISPLDLMSAMIKGTLDPSNVSSTSGAGSVFLENREFVMVLTTSIAVLIGCVV 305
Query: 58 VLIWRRSNSQKPKPIEVPKRVIEKLP-ELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEA 116
V IWRRS K K IE PKRV+EKL E E+DDGT+KVT+FFGTQTGTAEGFAKAIAEEA
Sbjct: 306 VFIWRRSTGNKAKSIEPPKRVVEKLSDEAEVDDGTRKVTIFFGTQTGTAEGFAKAIAEEA 485
Query: 117 KARYEKAKFKVVDMDDYAADDDEYEE 142
K RYEKAKFK+VDMDDYA DDDEYEE
Sbjct: 486 KVRYEKAKFKIVDMDDYAQDDDEYEE 563
>TC17600 similar to PIR|A47298|A47298 NADPH-ferrihemoprotein reductase -
mung bean {Vigna radiata;} , partial (19%)
Length = 717
Score = 208 bits (530), Expect = 2e-54
Identities = 94/132 (71%), Positives = 120/132 (90%)
Frame = +3
Query: 605 QERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSELIVAFSREGPTKEY 664
QER ALKE+G ELGP++LFFGCRNR++D+IYEDELN+F+ G+LSELIVAFSREG KEY
Sbjct: 3 QERFALKEDGIELGPAILFFGCRNRRMDFIYEDELNNFLEQGSLSELIVAFSREGSEKEY 182
Query: 665 VQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVKN 724
VQHKM++KA+ +W++ISQGAY+YVCGDAKGMA+DVH TLHTI+Q+Q ++++SK E++VK
Sbjct: 183 VQHKMMDKAAYLWSLISQGAYLYVCGDAKGMARDVHHTLHTIVQQQENVESSKAEAIVKK 362
Query: 725 LQMTGRYLRDVW 736
LQ+ GRYLRDVW
Sbjct: 363 LQLDGRYLRDVW 398
>TC19435 similar to UP|Q39036 (Q39036) NADPH-ferrihemoprotein reductase ,
partial (12%)
Length = 459
Score = 170 bits (431), Expect = 7e-43
Identities = 91/130 (70%), Positives = 105/130 (80%), Gaps = 3/130 (2%)
Frame = +2
Query: 1 MQDSSSMKFSPLDLMSALIKGKIDPSNGTVPA---SLILENREFVMILTTSIAVLIGCVV 57
M+ SSSMK SPLD+MSA+IKG +DPSN + + S+ILEN EFVM+LTTSIAVLIGCVV
Sbjct: 74 MEGSSSMKISPLDMMSAMIKGTLDPSNVSSTSGAGSIILEN-EFVMVLTTSIAVLIGCVV 250
Query: 58 VLIWRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAK 117
VLIWRRS K IE PKRV++ E E+DDGTKKVT+FFGTQTG AEGFAKA+ EAK
Sbjct: 251 VLIWRRSTGNNAKSIEPPKRVVD---EAEVDDGTKKVTLFFGTQTGNAEGFAKALVAEAK 421
Query: 118 ARYEKAKFKV 127
RYEKAKFK+
Sbjct: 422 VRYEKAKFKI 451
>CB828786
Length = 456
Score = 144 bits (362), Expect = 7e-35
Identities = 75/107 (70%), Positives = 78/107 (72%), Gaps = 28/107 (26%)
Frame = +1
Query: 632 DYIYEDELNHFVHGGALSELIVAFSREGPTKEYVQHKMIEK------------------- 672
DYIYEDELNHFV+ GALSELIVAFSREGPTKEYVQHKM+EK
Sbjct: 4 DYIYEDELNHFVNSGALSELIVAFSREGPTKEYVQHKMMEKVPILSQDAFNYQFTIYSFS 183
Query: 673 ---------ASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQ 710
ASDIWNMISQGAYIYVCGDAKGMA+DVHRTLHTILQEQ
Sbjct: 184 Y*TLYLCGQASDIWNMISQGAYIYVCGDAKGMARDVHRTLHTILQEQ 324
>TC8087 homologue to UP|FENR_PEA (P10933) Ferredoxin--NADP reductase, leaf
isozyme, chloroplast precursor (FNR) , partial (94%)
Length = 1449
Score = 60.1 bits (144), Expect = 1e-09
Identities = 58/209 (27%), Positives = 95/209 (44%), Gaps = 11/209 (5%)
Frame = +1
Query: 514 RYYSISSSPR--VAPSRIHVTCA--LVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWA 569
R YSI+SS S+ C LV+ +G I +GVCS ++ + P S+
Sbjct: 559 RLYSIASSALGDFGDSKTVSLCVKRLVYTN-DSGEIVKGVCSNFLCDLKP--GSEVTITG 729
Query: 570 PIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAEL-GPSVLFFGCRN 628
P+ +P D +IM+ GTG+APFR FL + K + + G + LF G
Sbjct: 730 PV---GKEMLMPKDPNATVIMLATGTGIAPFRSFLWKMFFEKHDDYKFNGLAWLFLGVPT 900
Query: 629 RQVDYIYEDELNHFVHGGALS-ELIVAFSREGPT----KEYVQHKMIEKASDIWNMISQ- 682
+Y++E + L A SRE K Y+Q +M + A ++W ++ +
Sbjct: 901 SS-SLLYKEEFEKMKEKSPENFRLDFAVSREQTNDKGEKMYIQTRMAQYAEELWQLLKKD 1077
Query: 683 GAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
++Y+CG KGM K + + ++ G
Sbjct: 1078NTFVYMCG-LKGMEKGIDDIMVSLAARDG 1161
>TC15271 similar to PIR|T06773|T06773 ferredoxin-NADP reductase - garden
pea (fragment) {Pisum sativum;} ,
partial (27%)
Length = 729
Score = 43.5 bits (101), Expect = 1e-04
Identities = 24/76 (31%), Positives = 40/76 (52%), Gaps = 4/76 (5%)
Frame = +2
Query: 653 VAFSREGPTKE----YVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQ 708
+A SRE + YVQ K+ E + +I+ ++ GA+IY CG +GM + TL +
Sbjct: 62 IALSREQKNQNGGQMYVQDKIEEYSDEIFKLLDNGAHIYFCG-LRGMMPGIQETLKRVAD 238
Query: 709 EQGSLDNSKTESMVKN 724
++G + K + KN
Sbjct: 239 QRGENWDEKLSQLKKN 286
>TC16072
Length = 743
Score = 32.0 bits (71), Expect(2) = 0.049
Identities = 14/27 (51%), Positives = 17/27 (62%)
Frame = +2
Query: 179 EEDSFKNLQYGVFGLGNRQYEHFNKVC 205
E F+NL LGNRQY++FN VC
Sbjct: 335 ERRRFRNLHMLFLVLGNRQYDNFNMVC 415
Score = 21.6 bits (44), Expect(2) = 0.049
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = +3
Query: 231 SDIYHSTQVAKIVDDKLLE 249
S + Q+A +VD+KLLE
Sbjct: 492 SSLCELIQMANVVDEKLLE 548
>TC11355
Length = 679
Score = 33.1 bits (74), Expect = 0.16
Identities = 22/111 (19%), Positives = 48/111 (42%)
Frame = -2
Query: 65 NSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARYEKAK 124
+S P+PI +P +++ KK+T++ QTG A A + KAR + ++
Sbjct: 402 HSDLPEPILLP*KILGVAFSRINSSNLKKITLYPNMQTGITLTLASGFANDHKARKKSSR 223
Query: 125 FKVVDMDDYAADDDEYEEKLKKETMALFFLATYGDGEPTDNAARFYKWFEE 175
+ + + + + + + +K F +P N + +Y+ F +
Sbjct: 222 IEQIKLLNV*IETNFASQ*IKPCLSCHFLNVFQRSMDPASNDSAYYQCFSQ 70
>TC13075 similar to UP|O23442 (O23442) Dynein light chain like protein,
partial (86%)
Length = 514
Score = 30.4 bits (67), Expect = 1.1
Identities = 26/79 (32%), Positives = 35/79 (43%), Gaps = 7/79 (8%)
Frame = +2
Query: 407 DDEDGKPLGGSSLPPPFPPCTLRTA------LAKYADVLSSPKKSAL-LALAAHASDPSE 459
+ ED KP+ S PPP PP T A + K AD+L +K A+ +A+AA E
Sbjct: 107 NSEDRKPMLSSLSPPP-PPQTPSAAPAPKKVIIKSADMLPDMQKEAVDIAVAAFERLNVE 283
Query: 460 ADRLRHLASPAGKDEYAEW 478
D H+ K W
Sbjct: 284 KDVAEHIKKEFDKRHGPTW 340
>TC15370 weakly similar to UP|Q8S8R0 (Q8S8R0) Predicted protein, partial
(38%)
Length = 714
Score = 30.0 bits (66), Expect = 1.4
Identities = 25/85 (29%), Positives = 35/85 (40%), Gaps = 12/85 (14%)
Frame = -1
Query: 494 SSAKPPIGVFFASVAPRLQPRYYSI----SSSPRVAPSRIHVTCALVHDKMPTGRIHQGV 549
SS P I V ++ +L +I S+SP + R+H VH T R+H G+
Sbjct: 483 SSPTPLINVVACIISLQLDMNQETILTTTSTSPSLPKQRLHSMECAVHPLQNTTRVHMGM 304
Query: 550 CSTWMKNSAPL--------EKSQDC 566
S + L EKSQ C
Sbjct: 303 TSQGKDKATTLKQRHV*SIEKSQIC 229
>TC19977 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (20%)
Length = 452
Score = 29.3 bits (64), Expect = 2.4
Identities = 15/37 (40%), Positives = 19/37 (50%)
Frame = -1
Query: 321 KQLNGNGHAVVDAHHPVRANVAVRKELHTPASDRSCT 357
K L G+GH +VD H N V++EL RS T
Sbjct: 308 KDLGGHGHDIVDEVHYYSRNSKVKQELELNPCRRSIT 198
>BP051910
Length = 429
Score = 29.3 bits (64), Expect = 2.4
Identities = 11/53 (20%), Positives = 24/53 (44%)
Frame = -3
Query: 684 AYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVKNLQMTGRYLRDVW 736
A +Y+ G + M DV I+ ++ + ++ L+ +G+Y + W
Sbjct: 427 AAVYIAGSSTQMPADVTSAFEEIISKENEVSKEDAVRWIRALEKSGKYHVEAW 269
>TC9565 similar to UP|Q9MAQ2 (Q9MAQ2) CDS, partial (19%)
Length = 872
Score = 28.9 bits (63), Expect = 3.1
Identities = 17/55 (30%), Positives = 23/55 (40%)
Frame = +3
Query: 394 LGLSPDTYFSVHTDDEDGKPLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALL 448
LGLSP ++ S GG PP PP T + + SS ++ LL
Sbjct: 303 LGLSPSSFSSPSPSTNATTGGGGGGRAPPLPPPTSSSGYLSISSAASSLERYCLL 467
>TC9087 homologue to UP|AAR83868 (AAR83868) 60S ribosomal protein L12,
complete
Length = 787
Score = 28.9 bits (63), Expect = 3.1
Identities = 28/92 (30%), Positives = 42/92 (45%)
Frame = +1
Query: 438 VLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVIASQRSLLEVMAEFSSAK 497
+L +P SA++ +S P + ASPA + A V + +RS+ V + S K
Sbjct: 19 LLQNPNLSAIVPQCRRSSIPLRSSTFTS-ASPAERS--APPVPSLRRSVPSVSPQRRSEK 189
Query: 498 PPIGVFFASVAPRLQPRYYSISSSPRVAPSRI 529
+PR QPR +S SP +P RI
Sbjct: 190 ---------TSPRRQPRTGRVSVSPSSSPCRI 258
>TC11285
Length = 456
Score = 28.5 bits (62), Expect = 4.0
Identities = 17/40 (42%), Positives = 19/40 (47%)
Frame = +2
Query: 504 FASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKMPTG 543
FA+ AP L P SP V+P RIH L K P G
Sbjct: 158 FATAAPSLPP-------SPSVSPQRIHRVLILQVRKFPNG 256
>TC10042
Length = 575
Score = 28.1 bits (61), Expect = 5.3
Identities = 16/64 (25%), Positives = 30/64 (46%), Gaps = 4/64 (6%)
Frame = +3
Query: 32 ASLILENREFVMILTTSIAVLIGCVVVLIWRRSNSQKPKPIEVPKRVIEKL----PELEI 87
+S L++ ++ + +T + + I C LI + IE+P R+I + EL I
Sbjct: 162 SSFFLDDSQYFLSITLTSCIKISCSTFLITSSATKASTSSIELPGRIITGIC**EIELSI 341
Query: 88 DDGT 91
+ T
Sbjct: 342 SEST 353
>TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomerase
precursor , partial (81%)
Length = 1355
Score = 28.1 bits (61), Expect = 5.3
Identities = 36/116 (31%), Positives = 48/116 (41%), Gaps = 3/116 (2%)
Frame = +2
Query: 413 PLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALA--AHASDPSEADRLRHLASPA 470
PL S+P P P T R + SSP S+ A A +S PS A SP
Sbjct: 365 PLPNPSVPSPSPKTTSRDSPPTRPSTTSSPAWSSASAPAPPPPSSSPSSASFSPPANSPT 544
Query: 471 GKDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISS-SPRVA 525
A +++RS + A S P AS++P P ++SS S RVA
Sbjct: 545 SS---ASPPPSARRSRRDPSA---SRSPSSTTIPASISPSTAPTRSTLSSTSSRVA 694
>AV775527
Length = 466
Score = 27.7 bits (60), Expect = 6.9
Identities = 18/69 (26%), Positives = 28/69 (40%)
Frame = +1
Query: 397 SPDTYFSVHTDDEDGKPLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASD 456
SP + SV + P SLPPP PP + T + + + S + A H
Sbjct: 79 SPSSSSSVPSSSSSNLP----SLPPPPPPSPVSTPTSTSSPMPSPSPVPTCASRATHLPA 246
Query: 457 PSEADRLRH 465
P+ + + H
Sbjct: 247 PASSSAVSH 273
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,794,049
Number of Sequences: 28460
Number of extensions: 163677
Number of successful extensions: 914
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 899
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 909
length of query: 736
length of database: 4,897,600
effective HSP length: 97
effective length of query: 639
effective length of database: 2,136,980
effective search space: 1365530220
effective search space used: 1365530220
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135801.5


