

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
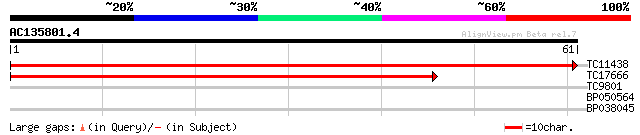
Query= AC135801.4 + phase: 0
(61 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11438 homologue to UP|RUXF_ARATH (Q9SUM2) Probable small nucle... 120 5e-29
TC17666 similar to GB|AAP37664.1|30725284|BT008305 At2g43810 {Ar... 50 1e-07
TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like... 30 0.12
BP050564 26 1.8
BP038045 25 3.0
>TC11438 homologue to UP|RUXF_ARATH (Q9SUM2) Probable small nuclear
ribonucleoprotein F (snRNP-F) (Sm protein F) (Sm-F)
(SmF), partial (97%)
Length = 585
Score = 120 bits (301), Expect = 5e-29
Identities = 57/61 (93%), Positives = 59/61 (96%)
Frame = +2
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDAAD 60
MEYKGYLVSVDSYMNLQLANTEEYIEG FTGNLGEILIRCNNVLY+RGVPEDEEIEDA +
Sbjct: 194 MEYKGYLVSVDSYMNLQLANTEEYIEGQFTGNLGEILIRCNNVLYLRGVPEDEEIEDAPE 373
Query: 61 D 61
D
Sbjct: 374 D 376
>TC17666 similar to GB|AAP37664.1|30725284|BT008305 At2g43810 {Arabidopsis
thaliana;}, partial (95%)
Length = 770
Score = 49.7 bits (117), Expect = 1e-07
Identities = 20/46 (43%), Positives = 29/46 (62%)
Frame = +3
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
++Y+G L +D YMN+ + TEEY+ G G+ IR NNVLY+
Sbjct: 234 VDYRGILACLDGYMNIAMEQTEEYVNGQLKNKYGDAFIRGNNVLYI 371
>TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like, complete
Length = 981
Score = 29.6 bits (65), Expect = 0.12
Identities = 23/63 (36%), Positives = 35/63 (55%), Gaps = 4/63 (6%)
Frame = +2
Query: 3 YKGYLVSVDSYMNLQLANTEEYIEGNFTGN----LGEILIRCNNVLYMRGVPEDEEIEDA 58
Y G+LV+ D++MN+ L E I N G+ + E IR N + Y+R VP DE I+
Sbjct: 170 YNGHLVNCDTWMNIHL---REVICTNKDGDRFWRMPECYIRGNTIKYLR-VP-DEVIDKV 334
Query: 59 ADD 61
++
Sbjct: 335 QEE 343
>BP050564
Length = 490
Score = 25.8 bits (55), Expect = 1.8
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +2
Query: 2 EYKGYLVSVDSYMNLQLANT 21
E+KGY + DS+MN +AN+
Sbjct: 419 EFKGYKLGGDSWMNTGIANS 478
>BP038045
Length = 556
Score = 25.0 bits (53), Expect = 3.0
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = +1
Query: 4 KGYLVSVDSYMNLQLANT 21
+G L SVD Y+N++L NT
Sbjct: 106 RGTLHSVDQYLNIKLENT 159
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.137 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 739,044
Number of Sequences: 28460
Number of extensions: 5742
Number of successful extensions: 29
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 29
length of query: 61
length of database: 4,897,600
effective HSP length: 37
effective length of query: 24
effective length of database: 3,844,580
effective search space: 92269920
effective search space used: 92269920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 49 (23.5 bits)
Medicago: description of AC135801.4


