

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135799.3 + phase: 0
(242 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
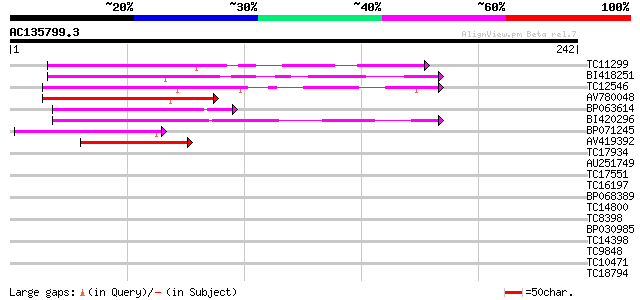
Score E
Sequences producing significant alignments: (bits) Value
TC11299 87 3e-18
BI418251 80 2e-16
TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat ... 78 2e-15
AV780048 69 7e-13
BP063614 54 2e-08
BI420296 53 5e-08
BP071245 49 6e-07
AV419392 47 4e-06
TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase ... 36 0.005
AU251749 35 0.009
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 30 0.49
TC16197 similar to PIR|H96696|H96696 protein F1N21.16 [imported]... 28 1.1
BP068389 27 2.4
TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromo... 27 2.4
TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific... 27 2.4
BP030985 27 3.2
TC14398 similar to UP|Q96477 (Q96477) LRR protein, partial (86%) 27 3.2
TC9848 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lect... 27 3.2
TC10471 weakly similar to UP|Q84QE1 (Q84QE1) Avr9/Cf-9 rapidly e... 27 3.2
TC18794 27 4.1
>TC11299
Length = 498
Score = 86.7 bits (213), Expect = 3e-18
Identities = 56/164 (34%), Positives = 90/164 (54%), Gaps = 1/164 (0%)
Frame = +1
Query: 17 VADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSK 76
+ D+IS LPD+++ +ILS + T+ AVAT++LSKRW +LW S+ +DF+++++ + +
Sbjct: 79 MVDRISALPDEIICHILSFLPTENAVATAVLSKRWTHLWRSVSALDFSSVRMYEPDDHRF 258
Query: 77 FI-DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVN 135
F D VYSVL+ + A I +F L++ D+ + N+ KWVN
Sbjct: 259 FFSDIVYSVLLFRNAATP----IKKFSLEL-----------DDDVAVANLDIANIHKWVN 393
Query: 136 LVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRT 179
V Q R+G HL + D LP+LPI+I +C+T
Sbjct: 394 FVTQ---------CRVGGIEHLRLHFMDFIQLPQLPISILSCKT 498
>BI418251
Length = 546
Score = 80.5 bits (197), Expect = 2e-16
Identities = 58/172 (33%), Positives = 91/172 (52%), Gaps = 3/172 (1%)
Frame = +1
Query: 17 VADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN---IKIDSIES 73
+AD+IS+LPD+VL ILS + T++AVATS+L KRW++LWLS+P +DF++ +K ++
Sbjct: 31 MADRISKLPDEVLCQILSFLPTEDAVATSVLCKRWSSLWLSVPTLDFDDYRYLKGKKLKL 210
Query: 74 NSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKW 133
S FI+ VY++++S I F L V + P+ +V W
Sbjct: 211 QSSFINFVYAIILSRAL----HQPIKNFTLSV--------ISEECPY-------PDVKVW 333
Query: 134 VNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
+N +QR+++ L + L L LP +I +C TLV L L
Sbjct: 334 LNAAMQRQVENLDICLSLS----------------TLPCSILSCTTLVVLKL 441
>TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat shock
transcription factor HSF30, partial (9%)
Length = 755
Score = 77.8 bits (190), Expect = 2e-15
Identities = 61/185 (32%), Positives = 91/185 (48%), Gaps = 14/185 (7%)
Frame = +2
Query: 15 LAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDS---- 70
+ + D IS LPD +L +ILS +++KEAVATS+LS+RW LW S+P + F + +
Sbjct: 137 MKMEDGISTLPDALLCHILSFLTSKEAVATSVLSRRWIPLWRSVPTLHFKDANYHTDIGH 316
Query: 71 ------IESNSKFIDSVYSVLVSPDTAIGGSHF---INRFCLDVQFRNGNPLYGYDNPHL 121
+ S+ + SVY+V++S D + F +N C P Y
Sbjct: 317 ADHDIVKDVRSRHVQSVYAVILSRDFQLPIKKFYLRLNDVC--------QPFY------- 451
Query: 122 LYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPI-TIFTCRTL 180
NV WVN VVQR+L++L ++L Y P+ + +IF+CRTL
Sbjct: 452 ----DPANVSVWVNAVVQRQLEHLDISL-----------PYPMLSTPRANLSSIFSCRTL 586
Query: 181 VSLDL 185
V L L
Sbjct: 587 VVLKL 601
>AV780048
Length = 501
Score = 68.9 bits (167), Expect = 7e-13
Identities = 34/76 (44%), Positives = 53/76 (69%), Gaps = 1/76 (1%)
Frame = +2
Query: 15 LAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIK-IDSIES 73
+ + D+ S LPD +L +ILSL++TKEAV TSILSKRW LW S+P +DF++ E
Sbjct: 239 MKMEDRFSSLPDPILCHILSLLTTKEAVTTSILSKRWIPLWRSVPTLDFDDSNWYGGDEV 418
Query: 74 NSKFIDSVYSVLVSPD 89
++ +++VY V+++ D
Sbjct: 419 YARLVEAVYKVILARD 466
>BP063614
Length = 508
Score = 53.9 bits (128), Expect = 2e-08
Identities = 31/79 (39%), Positives = 46/79 (57%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D++S LPD +L +ILS V K AV T ILS RW +LW LP++ + + +S +KF+
Sbjct: 167 DRLSDLPDCILLHILSFVKAKAAVQTCILSTRWKDLWKRLPSLILLSSNFWTYKSFTKFV 346
Query: 79 DSVYSVLVSPDTAIGGSHF 97
+ + L TA+ G F
Sbjct: 347 SRLLT-LRDGSTALHGLDF 400
>BI420296
Length = 592
Score = 52.8 bits (125), Expect = 5e-08
Identities = 45/167 (26%), Positives = 70/167 (40%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D++S LPD VL ++L +STK+AV T +LS RW +LW + + N+ + S+F+
Sbjct: 194 DRLSDLPDLVLLHVLKFMSTKQAVQTCVLSTRWKDLWKGVTTLALNSSDFATAPRFSEFL 373
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
V S + ++ + C++ + N Y V+ V
Sbjct: 374 SCVLSHR-NDSVSLHNLDLRRKGCVEPELLNRVMSYA------------------VSHDV 496
Query: 139 QRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
Q L L+LG+ LH IF+CRTL L L
Sbjct: 497 QSLTIEFNLYLKLGFKLH---------------PCIFSCRTLTYLKL 592
>BP071245
Length = 552
Score = 49.3 bits (116), Expect = 6e-07
Identities = 28/70 (40%), Positives = 39/70 (55%), Gaps = 5/70 (7%)
Frame = +1
Query: 3 RQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNI- 61
++ R S + D++S LPD VL +ILS V+ K AV T LS RW +LW LP++
Sbjct: 334 KRGRESESESENEENKDRLSDLPDCVLLHILSFVNAKYAVQTCTLSTRWKDLWKRLPSLI 513
Query: 62 ----DFNNIK 67
DF+ K
Sbjct: 514 LHSSDFSTFK 543
>AV419392
Length = 318
Score = 46.6 bits (109), Expect = 4e-06
Identities = 20/48 (41%), Positives = 32/48 (66%)
Frame = +3
Query: 31 YILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
+ILS V TK+A+ S+LSKRW +LW +LP ++F++ S + F+
Sbjct: 3 HILSYVETKDAMQCSVLSKRWRSLWRALPVLNFDSASFTHYSSFANFV 146
>TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 564
Score = 36.2 bits (82), Expect = 0.005
Identities = 16/35 (45%), Positives = 23/35 (65%)
Frame = -1
Query: 21 ISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLW 55
I++LPD + ILS + EA TSILS++W +W
Sbjct: 564 INQLPDGIPGAILSKLPINEAAKTSILSRKWRYMW 460
>AU251749
Length = 362
Score = 35.4 bits (80), Expect = 0.009
Identities = 20/44 (45%), Positives = 30/44 (67%)
Frame = +2
Query: 2 QRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATS 45
+R+K RR + + + D IS LPD +L++ILS + TK+ VATS
Sbjct: 92 ERRKCRR*FTLSRMV--DWISALPDTILNFILSFLPTKQVVATS 217
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 29.6 bits (65), Expect = 0.49
Identities = 15/47 (31%), Positives = 29/47 (60%)
Frame = +2
Query: 21 ISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIK 67
IS LPDD++ L+ V + S++ +RW++L L + DF++++
Sbjct: 128 ISSLPDDIILDFLTRVPPSSLPSLSLVCRRWSHL---LHSPDFSDLR 259
>TC16197 similar to PIR|H96696|H96696 protein F1N21.16 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 582
Score = 28.5 bits (62), Expect = 1.1
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 5/58 (8%)
Frame = +3
Query: 5 KRRRRSSKASLAVADKISRLPDDVLHYILSLVST-----KEAVATSILSKRWNNLWLS 57
++R++ S A+ + D LPDD++ ILS +S+ + + I KR N+L L+
Sbjct: 114 RKRQKKSPAATSGDDHFESLPDDLVISILSKLSSSSTSPSDFASVLITCKRLNSLGLN 287
>BP068389
Length = 557
Score = 27.3 bits (59), Expect = 2.4
Identities = 17/47 (36%), Positives = 24/47 (50%)
Frame = -3
Query: 134 VNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTL 180
+ L+ R LK L Y + L+ D YD PK+P+ +CRTL
Sbjct: 351 MRLL*NRSLKNL*KTQTRTYGITLECD-YDTMVFPKIPLPGDSCRTL 214
>TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromosome 5, TAC
clone:K14B20, partial (7%)
Length = 782
Score = 27.3 bits (59), Expect = 2.4
Identities = 25/97 (25%), Positives = 41/97 (41%), Gaps = 19/97 (19%)
Frame = +3
Query: 24 LPDDVLHYILSLVSTKEAVATSILSKRWNNL-------------WLSLPNIDFNN----I 66
LPD+++ ILS + K + ++SK W + LS N DF + I
Sbjct: 48 LPDELIIEILSWLPVKSLLQFRVVSKTWKSFISDPQFVKLHLLHRLSFRNADFEHTSLLI 227
Query: 67 KIDSIESNSKFIDS--VYSVLVSPDTAIGGSHFINRF 101
K + + +I S V S+L SP + I+ +
Sbjct: 228 KCHTDDFGRPYISSRTVSSLLESPSAIVASRSCISGY 338
>TC8398 weakly similar to UP|Q84KK1 (Q84KK1) S haplotype-specific F-box
protein a, partial (9%)
Length = 1602
Score = 27.3 bits (59), Expect = 2.4
Identities = 22/78 (28%), Positives = 38/78 (48%)
Frame = +3
Query: 24 LPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFIDSVYS 83
L D+++ ILSL+ K V + K W L +S P +++ S ++ + F DS
Sbjct: 78 LIDELIWEILSLLPVKSLVRFRCVCKSW-KLTISNPQFMKLHLRRSSAKAAADFADSQVV 254
Query: 84 VLVSPDTAIGGSHFINRF 101
V+ D I S +++F
Sbjct: 255 VMTKRD-HIAHSPLLSKF 305
>BP030985
Length = 402
Score = 26.9 bits (58), Expect = 3.2
Identities = 12/36 (33%), Positives = 21/36 (58%), Gaps = 3/36 (8%)
Frame = +1
Query: 82 YSVLVSPDT---AIGGSHFINRFCLDVQFRNGNPLY 114
Y + + DT +IGG+ FI +F ++ + G+P Y
Sbjct: 52 YFPMAAADTKILSIGGTGFIGKFIVEASLKAGHPTY 159
>TC14398 similar to UP|Q96477 (Q96477) LRR protein, partial (86%)
Length = 982
Score = 26.9 bits (58), Expect = 3.2
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +1
Query: 162 YDNSYLPKLPITIFTCRTLVSLDLHHFSVKG 192
Y+N+ +P + ++L+SLDL+H +V G
Sbjct: 418 YENNIQGTIPEELGNLQSLISLDLYHNNVSG 510
>TC9848 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (19%)
Length = 394
Score = 26.9 bits (58), Expect = 3.2
Identities = 26/95 (27%), Positives = 43/95 (44%), Gaps = 11/95 (11%)
Frame = +1
Query: 12 KASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNN------LWLSLPNIDFNN 65
K +A I LP++ + ILS S +A S+LS + LW S D+++
Sbjct: 4 KPQMATRSIIETLPEECVSEILSHTSPPDACRFSMLSSTLRSAANSDMLWRSFLPSDYSD 183
Query: 66 IKIDS-----IESNSKFIDSVYSVLVSPDTAIGGS 95
I + + S+S F D ++ L +P GG+
Sbjct: 184IISRALNPLFLNSSSSFKD-LFKALCNPLLLDGGT 285
>TC10471 weakly similar to UP|Q84QE1 (Q84QE1) Avr9/Cf-9 rapidly elicited
protein 189, partial (18%)
Length = 459
Score = 26.9 bits (58), Expect = 3.2
Identities = 15/68 (22%), Positives = 33/68 (48%)
Frame = +1
Query: 17 VADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSK 76
+ D + +PD+ L I L++T + S + +RW +D N + S+ + +
Sbjct: 160 IRDYTAEIPDECLAAIFHLLTTTDRKICSAVCRRW-------LRVDGENRRRLSLNAEAS 318
Query: 77 FIDSVYSV 84
+D++ S+
Sbjct: 319 LLDAIPSL 342
>TC18794
Length = 373
Score = 26.6 bits (57), Expect = 4.1
Identities = 11/37 (29%), Positives = 22/37 (58%)
Frame = -1
Query: 206 NSLRLDTFKLYRSIYQVRFIMNTMHVIYFLFLSEILP 242
NS+R+ K+Y ++ + + T+H++ F+S I P
Sbjct: 373 NSVRVYNIKVY*IVH*INALTETVHMMCMYFISAISP 263
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,741,133
Number of Sequences: 28460
Number of extensions: 69874
Number of successful extensions: 576
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 570
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 573
length of query: 242
length of database: 4,897,600
effective HSP length: 88
effective length of query: 154
effective length of database: 2,393,120
effective search space: 368540480
effective search space used: 368540480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC135799.3


