

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
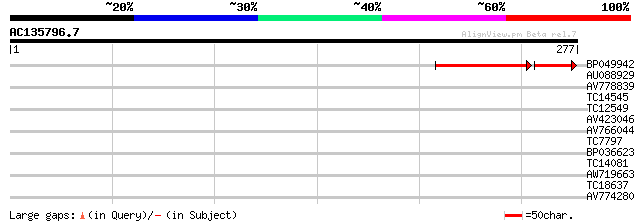
Query= AC135796.7 - phase: 0
(277 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP049942 51 1e-11
AU088929 28 1.3
AV778839 28 1.3
TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial ... 28 1.3
TC12549 similar to GB|AAO11541.1|27363244|BT002625 At1g79930/F19... 28 1.7
AV423046 27 5.0
AV766044 26 6.5
TC7797 similar to UP|TCTP_ORYSA (P35681) Translationally control... 26 6.5
BP036623 26 6.5
TC14081 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate c... 26 6.5
AW719663 26 8.5
TC18637 similar to UP|BAD12326 (BAD12326) NADH dehydrogenase sub... 26 8.5
AV774280 26 8.5
>BP049942
Length = 402
Score = 50.8 bits (120), Expect(2) = 1e-11
Identities = 25/47 (53%), Positives = 32/47 (67%)
Frame = -3
Query: 209 KKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTI 255
KKETE +D N S P+WVL PS+ELE++V Q DG+DGL T+
Sbjct: 400 KKETELKDP*NGSPSSPGPYWVLPPSQELESLVHQGDGHDGLCHHTL 260
Score = 34.3 bits (77), Expect(2) = 1e-11
Identities = 14/21 (66%), Positives = 18/21 (85%)
Frame = -1
Query: 257 EICKIYDLNTQDVVKYEKSTF 277
+IC+IYDL+T DV+KY KS F
Sbjct: 255 KICQIYDLDTLDVIKYGKSLF 193
>AU088929
Length = 720
Score = 28.5 bits (62), Expect = 1.3
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = +2
Query: 174 WILFRSYSHWRALQGS 189
W++F SYS WR ++GS
Sbjct: 533 WLIFSSYSRWRNIKGS 580
>AV778839
Length = 538
Score = 28.5 bits (62), Expect = 1.3
Identities = 17/56 (30%), Positives = 27/56 (47%)
Frame = +1
Query: 22 DHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAELFTDYVANKMNNGWVTL 77
+ ++ SSS+ Q+PST + LW S++ G FN+ N WV+L
Sbjct: 190 EDSMACSSSSVTLFQTPSTKRHLW-SLSGGSSSFNSP-------------NAWVSL 315
>TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial (46%)
Length = 1127
Score = 28.5 bits (62), Expect = 1.3
Identities = 20/71 (28%), Positives = 32/71 (44%), Gaps = 6/71 (8%)
Frame = +3
Query: 29 SSTADFSQSPSTLKQLWKSINTGDKPFNAKAELF--TDYVANKMNNG----WVTLENAPD 82
S A +P + S GDKP+ K ELF + +K+N G + ++ A
Sbjct: 180 SKDAKVDLTPDGVFTFTASAGAGDKPYEVKLELFDKVNVEESKINIGVRSIFSVVQKAES 359
Query: 83 GSFKKKIHGLG 93
G +K+ + G G
Sbjct: 360 GWWKRLLRGEG 392
>TC12549 similar to GB|AAO11541.1|27363244|BT002625 At1g79930/F19K16_11
{Arabidopsis thaliana;}, partial (25%)
Length = 677
Score = 28.1 bits (61), Expect = 1.7
Identities = 11/36 (30%), Positives = 21/36 (57%)
Frame = -1
Query: 137 IAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPF 172
I+ GT++ FF+G+ +L+ +S + P +PF
Sbjct: 386 ISATGTLVFFTFFFGASTLVSVSICTGLSPASGMPF 279
>AV423046
Length = 427
Score = 26.6 bits (57), Expect = 5.0
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = +3
Query: 13 RNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWK 46
+ WCF ++ TLP S D S + + + WK
Sbjct: 36 KEWCFVITV*KTLPECESMMD*SYVVTNVGRYWK 137
>AV766044
Length = 391
Score = 26.2 bits (56), Expect = 6.5
Identities = 15/43 (34%), Positives = 21/43 (47%)
Frame = +2
Query: 126 NARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLP 168
N ++ ++H+ R T IH FF + I SSA PLP
Sbjct: 176 NPKIKHGEVKHVKHRKTQIHISFFIIFIFSISSSSAPQPRPLP 304
>TC7797 similar to UP|TCTP_ORYSA (P35681) Translationally controlled tumor
protein homolog (TCTP), complete
Length = 1031
Score = 26.2 bits (56), Expect = 6.5
Identities = 22/100 (22%), Positives = 40/100 (40%), Gaps = 7/100 (7%)
Frame = +2
Query: 15 WCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDK-------PFNAKAELFTDYVA 67
W ++D + A+ S + Q K ++ D PF+ K +F ++
Sbjct: 167 WVVQGAVDVDIGANPSAEGGGEDEGVDDQAVKVVDIVDTFRLQEQPPFDKK--MFVTFIK 340
Query: 68 NKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFL 107
+ N L+ FKK I G +LLS++ + F+
Sbjct: 341 RYIKNLTPKLDEENQVLFKKHIEGATKYLLSKLSDLQFFV 460
>BP036623
Length = 524
Score = 26.2 bits (56), Expect = 6.5
Identities = 16/65 (24%), Positives = 33/65 (50%), Gaps = 5/65 (7%)
Frame = -2
Query: 198 DGSQSSNTYSGKKETEHEDSENESLG--LDEPHWVLTPSKELENIVRQEDGN---DGLSR 252
D + S N Y+G ++ +H D N + G +P+ + +L++++ + + D SR
Sbjct: 523 DNTLSVNDYNGNEKVDHPDLANNAEGGPYQDPNEIFGELGDLDSLLGLGEADLRGDPFSR 344
Query: 253 GTIEE 257
G +E
Sbjct: 343 GQEDE 329
>TC14081 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate
carboxylase/oxygenase activase, chloroplast precursor
(RuBisCO activase) (RA), partial (98%)
Length = 1625
Score = 26.2 bits (56), Expect = 6.5
Identities = 13/36 (36%), Positives = 23/36 (63%)
Frame = -3
Query: 149 FYGSVSLIPLSSAFSILPLPNVPFFWILFRSYSHWR 184
F GS + PLSS+ +++ + +P FW ++ SHW+
Sbjct: 672 FAGSPAF-PLSSSPALIMMGLIPIFW---KTSSHWK 577
>AW719663
Length = 474
Score = 25.8 bits (55), Expect = 8.5
Identities = 23/79 (29%), Positives = 37/79 (46%), Gaps = 15/79 (18%)
Frame = +1
Query: 210 KETEHEDSENESLGLDEPHWV---LTPSKELENIVRQEDGNDGLSRGTI----EEICKIY 262
+E+EHE E + LDE + V K E ++ +D N G+ + EE K+
Sbjct: 169 EESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDAN-GVDENEVENKPEEHIKLD 345
Query: 263 DLN--------TQDVVKYE 273
D N +QD+V++E
Sbjct: 346 DANGVDENSEHSQDLVEHE 402
>TC18637 similar to UP|BAD12326 (BAD12326) NADH dehydrogenase subunit 2,
partial (5%)
Length = 506
Score = 25.8 bits (55), Expect = 8.5
Identities = 13/29 (44%), Positives = 16/29 (54%), Gaps = 1/29 (3%)
Frame = -3
Query: 148 FFYGSVSLIPLSS-AFSILPLPNVPFFWI 175
+ Y +S LS SILP P V FFW+
Sbjct: 372 YIYSLISNHVLSLLTLSILPFPRVSFFWL 286
>AV774280
Length = 589
Score = 25.8 bits (55), Expect = 8.5
Identities = 12/34 (35%), Positives = 16/34 (46%)
Frame = +3
Query: 159 SSAFSILPLPNVPFFWILFRSYSHWRALQGSEKL 192
S F ++P +WI F + WR L GS L
Sbjct: 48 SPPFLLIPKKERRAWWIRFSTLQSWRRLLGSSTL 149
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,086,067
Number of Sequences: 28460
Number of extensions: 70104
Number of successful extensions: 473
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 470
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 473
length of query: 277
length of database: 4,897,600
effective HSP length: 89
effective length of query: 188
effective length of database: 2,364,660
effective search space: 444556080
effective search space used: 444556080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC135796.7


