

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
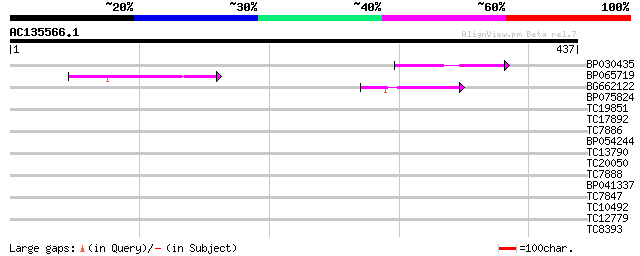
Query= AC135566.1 + phase: 0 /pseudo
(437 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP030435 57 5e-09
BP065719 46 1e-05
BG662122 44 7e-05
BP075824 29 1.3
TC19851 similar to UP|Q9SKK6 (Q9SKK6) At2g25430 protein (At2g254... 28 3.0
TC17892 similar to UP|Q8SXI8 (Q8SXI8) RE28061p, partial (4%) 28 3.0
TC7886 similar to UP|Q9ZWQ8 (Q9ZWQ8) Homolog to plastid-lipid-as... 28 3.0
BP054244 28 3.9
TC13790 weakly similar to UP|Q8VW43 (Q8VW43) Proline dehydrogena... 28 3.9
TC20050 weakly similar to UP|Q9SFG3 (Q9SFG3) F2O10.5 protein (At... 28 3.9
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 27 5.1
BP041337 27 5.1
TC7847 homologue to UP|SUSY_SOYBN (P13708) Sucrose synthase (Su... 27 5.1
TC10492 similar to AAL57709 (AAL57709) AT5g55940/MYN21_5, partia... 27 6.6
TC12779 weakly similar to UP|Q9FMB9 (Q9FMB9) Cysteine-tRNA ligas... 27 8.7
TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenyla... 27 8.7
>BP030435
Length = 533
Score = 57.4 bits (137), Expect = 5e-09
Identities = 32/89 (35%), Positives = 46/89 (50%)
Frame = +1
Query: 297 RVMGQDKDAWCKYHLVQGHNTDDCVHLKREIEKLLLSGKLRGYAKERRHNERQEDKPNPE 356
R+ G+ C YH GH T++C L+RE+ KL+ +GK + N
Sbjct: 73 RISGEPGGE*CVYHQAMGHITEECRTLQREVGKLIATGK-----------PTRIQGGNAP 219
Query: 357 QKHTLHTISGGFAGGGESSNSMKKYARQV 385
+HTI+GGF GGG +S + K+YAR V
Sbjct: 220 VPKGVHTIAGGFGGGGITSAARKRYARAV 306
>BP065719
Length = 567
Score = 45.8 bits (107), Expect = 1e-05
Identities = 30/120 (25%), Positives = 54/120 (45%), Gaps = 2/120 (1%)
Frame = +3
Query: 46 FSAKIWNAPVPDNFKPPHLPTFDGRGDPS--EHVTVFNTRMSVYGVADSLKCKLLAGTFA 103
F ++ + +P +K P F G S EH+ + + ++LK K +
Sbjct: 135 FIEEVLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLT 314
Query: 104 DAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARF 163
A W+ +L S+ + + + F +QF KV L +V++ P+ES+ +YL RF
Sbjct: 315 KNAFTWFTTLAPRSVHTWAQLERIFHEQFF-RGECKVSXKDLASVKRKPAESIDDYLNRF 491
>BG662122
Length = 386
Score = 43.5 bits (101), Expect = 7e-05
Identities = 30/88 (34%), Positives = 44/88 (49%), Gaps = 8/88 (9%)
Frame = +2
Query: 271 LNTRPERILKEVFESKII-------PPPPFSRARVMGQDKDAWCKYHLVQGHNTDDCVHL 323
LN IL++V + ++ PPP G D C+Y H+ DDC L
Sbjct: 101 LNAHLTDILQDVKAAHMVGKSGQS*PPPR------RGIDTTK*CEYRRSVVHDIDDCFTL 262
Query: 324 KREIEKLLLSGKLRGYAK-ERRHNERQE 350
KREIEKL+ G+L+ Y + R+ +RQ+
Sbjct: 263 KREIEKLIKMGRLKQYDRGSRQQGDRQQ 346
>BP075824
Length = 215
Score = 29.3 bits (64), Expect = 1.3
Identities = 14/37 (37%), Positives = 21/37 (55%), Gaps = 3/37 (8%)
Frame = +2
Query: 259 NQNRYVPEHFTPLN---TRPERILKEVFESKIIPPPP 292
++N+ H T LN T ER+L ++ +S PPPP
Sbjct: 101 DKNKITTVHCTELNECCTNTERVLAQLHDSSNTPPPP 211
>TC19851 similar to UP|Q9SKK6 (Q9SKK6) At2g25430 protein
(At2g25430/F13B15.9), partial (30%)
Length = 642
Score = 28.1 bits (61), Expect = 3.0
Identities = 19/66 (28%), Positives = 26/66 (38%)
Frame = +1
Query: 40 EPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLKCKLLA 99
E EP P +I P P+N+ PP P + + P + N R AD +
Sbjct: 13 EEEPVPDMNEIKALPPPENYTPPPQPEPEPKPQPQVQEDLVNLREDAV-TADDQGNRFAX 189
Query: 100 GTFADA 105
FA A
Sbjct: 190 ALFAGA 207
>TC17892 similar to UP|Q8SXI8 (Q8SXI8) RE28061p, partial (4%)
Length = 557
Score = 28.1 bits (61), Expect = 3.0
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 1/55 (1%)
Frame = +2
Query: 28 QEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDG-RGDPSEHVTVFN 81
+ + SHH + P P +A W AP D+++ P + G RG V F+
Sbjct: 239 RNRPPSHHSPAESSAPPPAAAPRWGAP-RDDYRSSGPPRWGGNRGGRDREVNPFD 400
>TC7886 similar to UP|Q9ZWQ8 (Q9ZWQ8) Homolog to plastid-lipid-associated
protein, partial (43%)
Length = 758
Score = 28.1 bits (61), Expect = 3.0
Identities = 13/46 (28%), Positives = 24/46 (51%), Gaps = 4/46 (8%)
Frame = +2
Query: 343 RRHNERQEDKPNPEQKHTLHT----ISGGFAGGGESSNSMKKYARQ 384
R H+E D+P+ ++ + + SGG+ GGGE+ + A +
Sbjct: 209 RTHSETSGDRPSENKRR*MGSGAFLFSGGYGGGGETGGDRDRDAEE 346
>BP054244
Length = 465
Score = 27.7 bits (60), Expect = 3.9
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = -1
Query: 148 VRQGPSESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLWAG-QFNESIAQKPADSMEE 205
++QG ESLR++L +F V++ + +V + + G F S A P+ M+E
Sbjct: 453 LKQGKKESLRDFLTKFYKEATLVTSLDLKVRLHLLCEAIRMGTSFYTSSAXTPSQDMDE 277
>TC13790 weakly similar to UP|Q8VW43 (Q8VW43) Proline dehydrogenase, partial
(3%)
Length = 423
Score = 27.7 bits (60), Expect = 3.9
Identities = 16/63 (25%), Positives = 29/63 (45%), Gaps = 6/63 (9%)
Frame = +3
Query: 10 EQNEALLASVETIQQVQQQEKTDSHHGDLDEPEPQPFS------AKIWNAPVPDNFKPPH 63
+ +E + ++T+QQ + ++ HH D P P S A+I + + + PH
Sbjct: 96 KNHETFMIQIQTMQQQMGEPSSNHHHHDELSPSPSFSSYSSETLAEIAARVIEELRRDPH 275
Query: 64 LPT 66
PT
Sbjct: 276 SPT 284
>TC20050 weakly similar to UP|Q9SFG3 (Q9SFG3) F2O10.5 protein (At3g05990),
partial (25%)
Length = 596
Score = 27.7 bits (60), Expect = 3.9
Identities = 13/41 (31%), Positives = 19/41 (45%), Gaps = 4/41 (9%)
Frame = +2
Query: 30 KTDSHHGDLDE----PEPQPFSAKIWNAPVPDNFKPPHLPT 66
K HH ++ P P+P S + P+PD+ P H T
Sbjct: 50 KPHHHHPEVSSSTVGPPPRPKSTAVHGFPIPDSSPPEHPKT 172
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 27.3 bits (59), Expect = 5.1
Identities = 16/41 (39%), Positives = 20/41 (48%)
Frame = +3
Query: 25 VQQQEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPHLP 65
V + K HG P P+P S K + P P KPPH+P
Sbjct: 270 VHKPPKYPPSHGPEPCPPPKP-SPKPPHYPKPPVVKPPHVP 389
>BP041337
Length = 446
Score = 27.3 bits (59), Expect = 5.1
Identities = 17/50 (34%), Positives = 23/50 (46%)
Frame = +1
Query: 323 LKREIEKLLLSGKLRGYAKERRHNERQEDKPNPEQKHTLHTISGGFAGGG 372
L+R L + R + RH++R ED P P + GFAGGG
Sbjct: 250 LRRRSRAGL*GRRRRRLLRRLRHHDR-ED*PEPASRSVFFFFRIGFAGGG 396
>TC7847 homologue to UP|SUSY_SOYBN (P13708) Sucrose synthase (Sucrose-UDP
glucosyltransferase) (Nodulin-100) , complete
Length = 2797
Score = 27.3 bits (59), Expect = 5.1
Identities = 14/40 (35%), Positives = 19/40 (47%), Gaps = 5/40 (12%)
Frame = -1
Query: 17 ASVETIQQVQQQEKTDS-----HHGDLDEPEPQPFSAKIW 51
A + + +QQQE S H LDE + QP K+W
Sbjct: 430 AETQQVAALQQQEHAHSLSDTPKHQALDEQQEQPMVVKLW 311
>TC10492 similar to AAL57709 (AAL57709) AT5g55940/MYN21_5, partial (65%)
Length = 478
Score = 26.9 bits (58), Expect = 6.6
Identities = 12/44 (27%), Positives = 23/44 (52%)
Frame = -1
Query: 333 SGKLRGYAKERRHNERQEDKPNPEQKHTLHTISGGFAGGGESSN 376
SGK+ Y E R++ E++ + + GG GGG++++
Sbjct: 286 SGKIFLYQNEGDFQLREKSDVRVEERDGIDDLDGGVVGGGDAAD 155
>TC12779 weakly similar to UP|Q9FMB9 (Q9FMB9) Cysteine-tRNA ligase, partial
(45%)
Length = 811
Score = 26.6 bits (57), Expect = 8.7
Identities = 18/45 (40%), Positives = 22/45 (48%)
Frame = -1
Query: 319 DCVHLKREIEKLLLSGKLRGYAKERRHNERQEDKPNPEQKHTLHT 363
+CVH+ + G RGY K+RRH ER D N Q H T
Sbjct: 322 NCVHMSPHSPSV*--GT*RGYQKKRRH-ERDRDG*NRRQ*HHKRT 197
>TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenylated Rab
receptor 2 {Arabidopsis thaliana;}, partial (78%)
Length = 1012
Score = 26.6 bits (57), Expect = 8.7
Identities = 20/92 (21%), Positives = 41/92 (43%), Gaps = 9/92 (9%)
Frame = -3
Query: 158 EYLARFNDSTIKVSNPNQEVFVGAFQNGLWAGQFNESIAQKPADS---------MEEIIA 208
++ +R+ +ST+ + +P+ G A++ +S EEI+
Sbjct: 782 DFTSRYQNSTLPLPDPSDPQSNATGDGRNRGGGDGSRSAEEGEESGGLRGLLLIKEEILR 603
Query: 209 RAECYVKGEESNAEKKAREAKERGNSGGERRN 240
AEC ++ + +AE ++R+ E G GE +
Sbjct: 602 DAECAMEANKGDAEHESRD--EDGTDTGEEND 513
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,091,708
Number of Sequences: 28460
Number of extensions: 121025
Number of successful extensions: 718
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 708
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 716
length of query: 437
length of database: 4,897,600
effective HSP length: 93
effective length of query: 344
effective length of database: 2,250,820
effective search space: 774282080
effective search space used: 774282080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135566.1


