

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135565.5 + phase: 0 /pseudo
(93 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
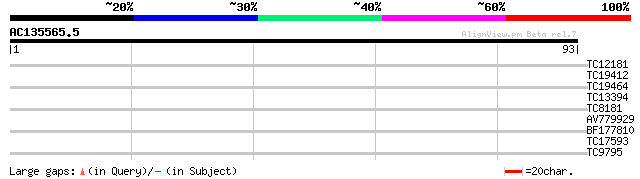
Score E
Sequences producing significant alignments: (bits) Value
TC12181 similar to AAQ89616 (AAQ89616) At3g01370, partial (14%) 30 0.071
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 26 1.0
TC19464 similar to UP|Q84LR1 (Q84LR1) Root-specific metal transp... 25 1.8
TC13394 24 3.9
TC8181 homologue to UP|O23310 (O23310) CCAAT-binding transcripti... 24 3.9
AV779929 24 5.1
BF177810 23 6.7
TC17593 similar to UP|Q8H0T8 (Q8H0T8) Eukaryotic initiation fact... 23 8.7
TC9795 homologue to UP|Q9ATB8 (Q9ATB8) BiP-isoform D (Fragment),... 23 8.7
>TC12181 similar to AAQ89616 (AAQ89616) At3g01370, partial (14%)
Length = 682
Score = 30.0 bits (66), Expect = 0.071
Identities = 9/29 (31%), Positives = 18/29 (62%)
Frame = +2
Query: 65 KVWRRCEVLRLTMNMGLSTANNNADQEEV 93
K+W RCE++++ + G+ N+ EE+
Sbjct: 356 KLWERCEIVKIAVKRGVQNTNSEIMAEEI 442
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 26.2 bits (56), Expect = 1.0
Identities = 8/32 (25%), Positives = 19/32 (59%)
Frame = -2
Query: 47 VVRVATRQQIVSSAVNASKVWRRCEVLRLTMN 78
++ V + + + + +N K+WR+C+ +R N
Sbjct: 188 II*VFPQHRNMKNRMNCRKIWRKCQTIRYRAN 93
>TC19464 similar to UP|Q84LR1 (Q84LR1) Root-specific metal transporter,
partial (22%)
Length = 503
Score = 25.4 bits (54), Expect = 1.8
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = -1
Query: 54 QQIVSSAVNASKVWRRCEVLRLTMNMGLSTANNNADQE 91
QQ++S A ++ ++ CE ++ N LST N +D+E
Sbjct: 242 QQLISVASSSLQISFHCEKPKIENN**LSTCN*ESDEE 129
>TC13394
Length = 511
Score = 24.3 bits (51), Expect = 3.9
Identities = 7/19 (36%), Positives = 13/19 (67%)
Frame = +1
Query: 30 FGGKVFVLVGDFRQVLPVV 48
FGG ++ + R+VLP++
Sbjct: 322 FGGNIYCVYSQIREVLPII 378
>TC8181 homologue to UP|O23310 (O23310) CCAAT-binding transcription factor
subunit A (At4g14540), partial (65%)
Length = 1223
Score = 24.3 bits (51), Expect = 3.9
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +3
Query: 55 QIVSSAVNASKVWRRCEVLRLTMNMGLSTANNNAD 89
Q VSSA AS WR E R T+++G S + D
Sbjct: 285 QRVSSANTASTTWRATESTR-TISVGASISATGLD 386
>AV779929
Length = 484
Score = 23.9 bits (50), Expect = 5.1
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +2
Query: 38 VGDFRQVLPVVRVATRQQIVSSAVNASKVWRR 69
VGD QV P+V + + V S N +VW R
Sbjct: 377 VGD*AQVQPLVSYSQVEARVDSLENQMQVWVR 472
>BF177810
Length = 431
Score = 23.5 bits (49), Expect = 6.7
Identities = 14/43 (32%), Positives = 20/43 (45%)
Frame = +1
Query: 46 PVVRVATRQQIVSSAVNASKVWRRCEVLRLTMNMGLSTANNNA 88
P R T + + AVNA + + L +N G+ NNNA
Sbjct: 97 PTARAVTGV-VTNPAVNAQVAFAKPNGANLPLNNGVPQNNNNA 222
>TC17593 similar to UP|Q8H0T8 (Q8H0T8) Eukaryotic initiation factor 4,
eIF4-like protein, partial (19%)
Length = 704
Score = 23.1 bits (48), Expect = 8.7
Identities = 10/42 (23%), Positives = 21/42 (49%)
Frame = -3
Query: 43 QVLPVVRVATRQQIVSSAVNASKVWRRCEVLRLTMNMGLSTA 84
++ P + R+++ S+ + +W R +L L + STA
Sbjct: 501 EIFPRSIIWIRRKLSSTTTPVTAIWLRISIL*LRWRIAFSTA 376
>TC9795 homologue to UP|Q9ATB8 (Q9ATB8) BiP-isoform D (Fragment), partial
(63%)
Length = 981
Score = 23.1 bits (48), Expect = 8.7
Identities = 10/27 (37%), Positives = 18/27 (66%)
Frame = +2
Query: 48 VRVATRQQIVSSAVNASKVWRRCEVLR 74
+R++TR + ++S + V +RC VLR
Sbjct: 410 IRLSTRMENLTSR*KSRMVRQRCSVLR 490
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.133 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,281,306
Number of Sequences: 28460
Number of extensions: 10365
Number of successful extensions: 49
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 49
length of query: 93
length of database: 4,897,600
effective HSP length: 69
effective length of query: 24
effective length of database: 2,933,860
effective search space: 70412640
effective search space used: 70412640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 48 (23.1 bits)
Medicago: description of AC135565.5


