

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
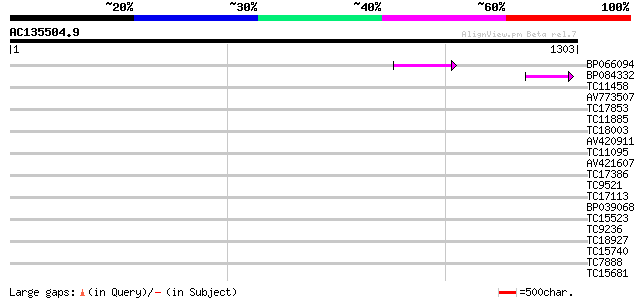
Query= AC135504.9 + phase: 0 /pseudo
(1303 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP066094 90 2e-18
BP084332 74 1e-13
TC11458 38 0.009
AV773507 37 0.020
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 37 0.026
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 34 0.17
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 33 0.22
AV420911 33 0.22
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 33 0.38
AV421607 32 0.50
TC17386 similar to UP|Q9LU46 (Q9LU46) DEAD-box protein abstrakt,... 32 0.85
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 30 1.9
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 30 2.5
BP039068 30 3.2
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 30 3.2
TC9236 similar to UP|Q9SZE9 (Q9SZE9) Formamidase-like protein ,... 29 5.5
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 28 7.2
TC15740 weakly similar to GB|AAM19935.1|20453393|AY097419 AT5g58... 28 7.2
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 28 7.2
TC15681 similar to UP|AAQ14515 (AAQ14515) NADH dehydrogenase sub... 28 9.4
>BP066094
Length = 532
Score = 90.1 bits (222), Expect = 2e-18
Identities = 63/143 (44%), Positives = 85/143 (59%)
Frame = +1
Query: 883 GSNQVTSILRS*SWHNIISNGCQECFS*WCH*RRSVCKTTSWV*GS*AS*PCL*T*EITI 942
GS Q T ++ S S HN S+ C EC S W H R S+C +T *P EI++
Sbjct: 94 GSYQTTDLILSESQHNSSSDEC*ECLSKWLHLRGSLCSSTPRFXR*EEL*PYFQIEEISL 273
Query: 943 WLETSS*SLV**TK*FLN*K*F*KRIS*RNTLQKDS*ERHSDCANIC**YNIWFY*CISL 1002
W E SS S+V* T+ + K* K S N+L ++ + +CAN+C**Y IW * IS+
Sbjct: 274 WSEASSQSMV*ETQLIPSGK*VXKG*SRYNSLLQNLQR*YINCANLC**YYIWIC*SISM 453
Query: 1003 QGIF*VNAG*I*NEHDGRIEVLS 1025
Q +F* +AG*I*N +DGR ++LS
Sbjct: 454 QRVF*DDAG*I*NAYDGRTQILS 522
>BP084332
Length = 368
Score = 74.3 bits (181), Expect = 1e-13
Identities = 47/109 (43%), Positives = 64/109 (58%)
Frame = +1
Query: 1186 NYCYVYSRSRIHFSCKLLYTTTLDETSVGRLSD*C*QYSHFL**YYCYLFVKESNSTF*S 1245
N+C +Y R RI+ S + ++ LDETS G LS+ QY + L**+ C ES TF
Sbjct: 37 NHCTIYCRGRIYLSSNMQHSDALDETSAGGLSNP*EQYPNLL**HCCNFIE*ESYLTFKG 216
Query: 1246 QAY*DQTPLYQRLCSKRNFRYTIH*Y*ASMG*YIYKAFIC*KI*FYKEK 1294
+A+* + LY RLC++ + *Y* SMG Y+YKA I*F+ EK
Sbjct: 217 KAH*GKVSLY*RLCTEGRTSSEVC*Y*PSMGRYLYKALSRG*I*FHSEK 363
>TC11458
Length = 572
Score = 38.1 bits (87), Expect = 0.009
Identities = 25/76 (32%), Positives = 36/76 (46%), Gaps = 4/76 (5%)
Frame = +3
Query: 218 HFIADCPDLQKEKSKSRPKKQSELVKTAGVPK-GKRETKVMVPRQRVFKAHDW---RESF 273
H ++ C ++EK++ R K +L + GVP+ G RE + VP H W R F
Sbjct: 117 HGVSHCRSQEEEKNQ-RESKIRQLGGSRGVPENGTREPSISVP------PHGWGDVRRGF 275
Query: 274 VPYPYHERWRRSEVWW 289
P PYH R+ W
Sbjct: 276 APGPYHAYAHRNSAGW 323
>AV773507
Length = 496
Score = 37.0 bits (84), Expect = 0.020
Identities = 17/40 (42%), Positives = 20/40 (49%)
Frame = +2
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCP 224
GG + G GRG Y + Q CF C +PGH DCP
Sbjct: 284 GGGYSSG---GRGSYGAGDRVGQDDCFECGRPGHRARDCP 394
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 36.6 bits (83), Expect = 0.026
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +2
Query: 198 GYKNSRKEDQKGCFNCKKPGHFIADCP 224
GY SR + CF C KPGHF +CP
Sbjct: 383 GYGGSRGSNGGECFKCGKPGHFARECP 463
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 33.9 bits (76), Expect = 0.17
Identities = 17/51 (33%), Positives = 25/51 (48%), Gaps = 1/51 (1%)
Frame = +2
Query: 177 KKPLESKF-GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCPDL 226
+ P++ K RF Y R + D C NCK+PGHF +CP++
Sbjct: 245 RSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNL-CKNCKRPGHFARECPNV 394
Score = 31.6 bits (70), Expect = 0.85
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +2
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADCPD 225
G G+ S + C+NCK+PGH + CP+
Sbjct: 413 GLPGHIASECTTKSLCWNCKEPGHMASSCPN 505
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 33.5 bits (75), Expect = 0.22
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +1
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GRGG + ED K C+ C +PGHF +C
Sbjct: 124 GRGGGRGRGGEDLK-CYECGEPGHFAREC 207
>AV420911
Length = 418
Score = 33.5 bits (75), Expect = 0.22
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +2
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GRGG + ED K C+ C +PGHF +C
Sbjct: 332 GRGGGRGRGGEDLK-CYECGEPGHFAREC 415
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 32.7 bits (73), Expect = 0.38
Identities = 29/104 (27%), Positives = 44/104 (41%), Gaps = 7/104 (6%)
Frame = +1
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADC-----PDLQKEKSKSRPKKQS 239
GG R G G GG + K C+ C +PGHF +C ++ +S R ++
Sbjct: 319 GGGGRGGGRSGGGGGGSDMK-----CYECGEPGHFARECRMRGGSGRRRSRSPPRFRRSP 483
Query: 240 ELVKTAGVPKGKRETKVMVPRQRVFKAHDWRESF--VPYPYHER 281
+ + P+G+ PR+R H R S+ P PY R
Sbjct: 484 SYGRRSYSPRGRS------PRRRSVSPHG-RGSYSRSPPPYRAR 594
>AV421607
Length = 245
Score = 32.3 bits (72), Expect = 0.50
Identities = 20/75 (26%), Positives = 28/75 (36%), Gaps = 7/75 (9%)
Frame = +3
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKKQSELV-------KTAGV 247
G G S GC+ C +PGH+ DCP S P + ++
Sbjct: 9 GSGPGSGSGSGTATGCYKCGRPGHWSRDCPS-SAPNSNPNPNPNTTTTPNPLLPSSSSFK 185
Query: 248 PKGKRETKVMVPRQR 262
P+ E +PRQR
Sbjct: 186 PRSAVEKPKKLPRQR 230
>TC17386 similar to UP|Q9LU46 (Q9LU46) DEAD-box protein abstrakt, partial
(19%)
Length = 507
Score = 31.6 bits (70), Expect = 0.85
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +2
Query: 208 KGCFNCKKPGHFIADCPDLQKEKSKSRPKKQSELVKTAG 246
KGC C GH I DCP L+ +KS + + + + G
Sbjct: 203 KGCAYCGGLGHRIRDCPKLEHQKSMAIANNRHDYFGSGG 319
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 30.4 bits (67), Expect = 1.9
Identities = 16/39 (41%), Positives = 20/39 (51%)
Frame = +1
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GGR R + +GRG S + CFNC GH+ DC
Sbjct: 331 GGRDRDREYMGRGPPPGSGR-----CFNCGIDGHWARDC 432
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 30.0 bits (66), Expect = 2.5
Identities = 13/33 (39%), Positives = 16/33 (48%), Gaps = 5/33 (15%)
Frame = -1
Query: 275 PYPYHERWR-RSEVWWQPNW----KNHWYRYYW 302
P + E+WR R E WW W WY Y+W
Sbjct: 816 PARWREQWRERREKWWWRRWWWWWS*TWYYYWW 718
>BP039068
Length = 467
Score = 29.6 bits (65), Expect = 3.2
Identities = 13/46 (28%), Positives = 22/46 (47%), Gaps = 1/46 (2%)
Frame = +3
Query: 201 NSRKEDQKGCFNCKKPGHFIADCPDLQ-KEKSKSRPKKQSELVKTA 245
N Q C NC++ GH DCP L + ++ P+ + +T+
Sbjct: 285 NHNNPRQTVCMNCQETGHASNDCPSLTLRSRNAQHPRMNQQSGETS 422
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 29.6 bits (65), Expect = 3.2
Identities = 10/27 (37%), Positives = 13/27 (48%), Gaps = 1/27 (3%)
Frame = +1
Query: 277 PYHERWRRSEVWWQPNWKN-HWYRYYW 302
P WRR WW W + W R++W
Sbjct: 586 PGSRPWRRRRWWWIRRWSS*RWIRWWW 666
>TC9236 similar to UP|Q9SZE9 (Q9SZE9) Formamidase-like protein , partial
(31%)
Length = 586
Score = 28.9 bits (63), Expect = 5.5
Identities = 16/54 (29%), Positives = 28/54 (51%)
Frame = +2
Query: 860 CCTRIQSTRRH*LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS*WCH 913
CC + S + H*LH + + G + +++L S N+ ++GC +C CH
Sbjct: 122 CCVQASSAQCH*LHIQVWLLQRTGVS--SAVLHSL*RKNLFNSGCPQCL---CH 268
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 28.5 bits (62), Expect = 7.2
Identities = 10/17 (58%), Positives = 10/17 (58%)
Frame = -2
Query: 210 CFNCKKPGHFIADCPDL 226
C C K GHF CPDL
Sbjct: 329 CHRCSKKGHFANRCPDL 279
>TC15740 weakly similar to GB|AAM19935.1|20453393|AY097419 AT5g58640/mzn1_90
{Arabidopsis thaliana;}, partial (65%)
Length = 575
Score = 28.5 bits (62), Expect = 7.2
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 3/44 (6%)
Frame = -3
Query: 254 TKVMVPRQRVFKAHDWRESFVPY--PYHERW-RRSEVWWQPNWK 294
T+V +PR+R K H W+ F Y PY+ + R+ Q WK
Sbjct: 534 TQVEIPRRRCNKTHHWKNLFPSYYSPYNTKLNNRNHFA*QTLWK 403
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 28.5 bits (62), Expect = 7.2
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -3
Query: 281 RWRRSEVWWQPNWKNHWYRYYW*FLH 306
RWR +WW NW H+ R W F++
Sbjct: 299 RWR--VLWWFMNWWLHYRRMLWRFMN 228
Score = 28.5 bits (62), Expect = 7.2
Identities = 19/73 (26%), Positives = 31/73 (42%), Gaps = 17/73 (23%)
Frame = -3
Query: 261 QRVFKAHDWRESFVPYPYHERWRRSEVWWQPN-------------WKNHW----YRYYW* 303
+ + + H+W + + +H RW + WW N W NHW R++W*
Sbjct: 506 RHIRRFHNWWFGNMGWFWHIRWFHN--WWFWNMGWFHNWWFWHMRWFNHWRLGIVRWFW* 333
Query: 304 FLHFNK*CVACRW 316
+L + A RW
Sbjct: 332 WLGWWTWFRAMRW 294
>TC15681 similar to UP|AAQ14515 (AAQ14515) NADH dehydrogenase subunit 2,
partial (7%)
Length = 586
Score = 28.1 bits (61), Expect = 9.4
Identities = 11/30 (36%), Positives = 14/30 (46%)
Frame = +2
Query: 273 FVPYPYHERWRRSEVWWQPNWKNHWYRYYW 302
FVP Y S+ WW + W+R YW
Sbjct: 104 FVPLAYRTVEDFSDEWWNLVHSSPWFRDYW 193
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.382 0.170 0.717
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,902,460
Number of Sequences: 28460
Number of extensions: 334960
Number of successful extensions: 5309
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 2215
Number of HSP's successfully gapped in prelim test: 206
Number of HSP's that attempted gapping in prelim test: 2912
Number of HSP's gapped (non-prelim): 2700
length of query: 1303
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1202
effective length of database: 2,023,140
effective search space: 2431814280
effective search space used: 2431814280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135504.9


