

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
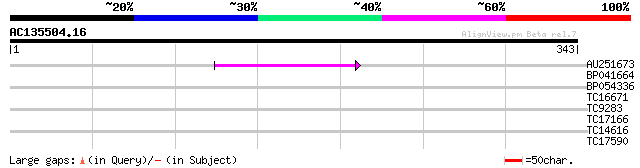
Query= AC135504.16 - phase: 0 /pseudo
(343 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU251673 55 2e-08
BP041664 28 2.9
BP054336 28 2.9
TC16671 similar to PIR|T52104|T52104 GATA-binding transcription ... 27 3.8
TC9283 27 3.8
TC17166 homologue to UP|Q84L15 (Q84L15) Cytosolic glutamine synt... 27 6.5
TC14616 27 6.5
TC17590 similar to UP|Q94BU7 (Q94BU7) AT3g10970/F9F8_21, partial... 26 8.5
>AU251673
Length = 413
Score = 54.7 bits (130), Expect = 2e-08
Identities = 29/89 (32%), Positives = 46/89 (51%), Gaps = 1/89 (1%)
Frame = +2
Query: 125 CSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEI 184
CS+ + V + L A DW+ L TWA F EF+NR+ P+ VR
Sbjct: 5 CSDTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVR 184
Query: 185 EFLELKHGD-MSVTKYAAKFVELAKFYPY 212
+F L+ + M+V++Y+A F L+++ PY
Sbjct: 185 DFERLEQAEGMTVSEYSAHFTHLSRYVPY 271
>BP041664
Length = 487
Score = 27.7 bits (60), Expect = 2.9
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = -1
Query: 32 ECQL*CCLCCCSELLFLM 49
E +L*CC CCC+ L L+
Sbjct: 169 ELRL*CCCCCCTGTLVLI 116
>BP054336
Length = 473
Score = 27.7 bits (60), Expect = 2.9
Identities = 8/15 (53%), Positives = 9/15 (59%)
Frame = +1
Query: 28 PPRIECQL*CCLCCC 42
PP C+ CC CCC
Sbjct: 238 PPNAPCEPCCCCCCC 282
>TC16671 similar to PIR|T52104|T52104 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (20%)
Length = 684
Score = 27.3 bits (59), Expect = 3.8
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -1
Query: 26 VEPPRIECQL*CCLCCCSELLFLM 49
++PP+IE L C CCC + F +
Sbjct: 201 LDPPKIEL*LISCCCCCRCMSFCL 130
>TC9283
Length = 1307
Score = 27.3 bits (59), Expect = 3.8
Identities = 11/42 (26%), Positives = 25/42 (59%)
Frame = +2
Query: 116 VERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDG 157
VER+ R M +E ++ +G H L+ ++ +++ L+++G
Sbjct: 905 VERLCRAMGGAEKVEIEYGNHSLSNRVEEAVQAIIDFLKREG 1030
>TC17166 homologue to UP|Q84L15 (Q84L15) Cytosolic glutamine synthetase,
partial (49%)
Length = 597
Score = 26.6 bits (57), Expect = 6.5
Identities = 13/37 (35%), Positives = 23/37 (62%)
Frame = +1
Query: 106 PDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEA 142
P GT L +++ + ++++ S + K+ GTH L EEA
Sbjct: 223 PSGTMMVLAQIKLLEKIVKSSYIHKLFSGTH-LGEEA 330
>TC14616
Length = 1353
Score = 26.6 bits (57), Expect = 6.5
Identities = 14/40 (35%), Positives = 16/40 (40%), Gaps = 3/40 (7%)
Frame = -1
Query: 76 QPNAAAGN---DGVRMLETFLRNHPPTFKGRYDPDGTQKW 112
QP+ A N DG +NHPPT K R W
Sbjct: 468 QPSTACSNTPQDGPATTRRTEKNHPPTSKSRRSQAAESPW 349
>TC17590 similar to UP|Q94BU7 (Q94BU7) AT3g10970/F9F8_21, partial (21%)
Length = 445
Score = 26.2 bits (56), Expect = 8.5
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = -3
Query: 37 CCLCCCSELLFL 48
CC CCC +LL L
Sbjct: 38 CCCCCCLQLLLL 3
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.355 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,103,185
Number of Sequences: 28460
Number of extensions: 79378
Number of successful extensions: 1524
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1251
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1446
length of query: 343
length of database: 4,897,600
effective HSP length: 91
effective length of query: 252
effective length of database: 2,307,740
effective search space: 581550480
effective search space used: 581550480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC135504.16


