

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135465.2 + phase: 0
(194 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
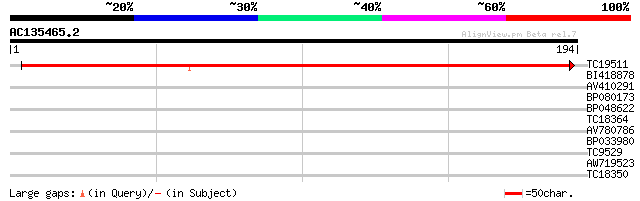
TC19511 similar to UP|Q9ARI4 (Q9ARI4) PII protein, partial (66%) 272 2e-74
BI418878 30 0.36
AV410291 28 1.0
BP080173 27 3.0
BP048622 27 3.0
TC18364 similar to GB|AAP21240.1|30102644|BT006432 At1g30540 {Ar... 26 3.9
AV780786 26 5.2
BP033980 26 5.2
TC9529 similar to UP|O48794 (O48794) F24O1.3, partial (9%) 26 5.2
AW719523 25 8.8
TC18350 similar to UP|O81911 (O81911) T7I23.17 protein, partial ... 25 8.8
>TC19511 similar to UP|Q9ARI4 (Q9ARI4) PII protein, partial (66%)
Length = 631
Score = 272 bits (696), Expect = 2e-74
Identities = 140/190 (73%), Positives = 158/190 (82%), Gaps = 1/190 (0%)
Frame = +1
Query: 5 AKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN-LPD 63
A+ ++F +NF +NE F+ S I G+ S RNV L+ GNA I+P+IRAQ+ +
Sbjct: 1 ARTHMFGVVNFQLNEAPMAFAGSSAILWHHGERSQRNVALRRRGNAMIVPRIRAQSSASE 180
Query: 64 YVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSEFSED 123
YVP+SKFYKVEAILRPWR+P VSS LL MGIRGVTVSDV+GFGAQGGSKERQGGSEFSED
Sbjct: 181 YVPDSKFYKVEAILRPWRVPLVSSALLNMGIRGVTVSDVRGFGAQGGSKERQGGSEFSED 360
Query: 124 NFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGEQAER 183
NFVAKVKMEIVVR DQVEAVI+KI+E ARTGEIGDGKIFLIPVSDVIR+RTGERGEQAER
Sbjct: 361 NFVAKVKMEIVVRNDQVEAVIDKIIEEARTGEIGDGKIFLIPVSDVIRVRTGERGEQAER 540
Query: 184 MAGGLTDALS 193
M GG TD LS
Sbjct: 541 MTGGRTDILS 570
>BI418878
Length = 609
Score = 29.6 bits (65), Expect = 0.36
Identities = 13/49 (26%), Positives = 23/49 (46%)
Frame = +2
Query: 32 KRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPW 80
KR+G HR++ + ++ + Q+ YV KFY +A + W
Sbjct: 443 KRYGRKKHRSIPKPKSAEPDLINQSGHQHAIAYVQGDKFYGAKATINVW 589
>AV410291
Length = 362
Score = 28.1 bits (61), Expect = 1.0
Identities = 12/35 (34%), Positives = 25/35 (71%)
Frame = +3
Query: 13 LNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSN 47
L+F +++ FPF SFS+ ++F SH +++L+++
Sbjct: 123 LSFTLHQILFPFLSFSICARKF---SHLHILLQTS 218
>BP080173
Length = 382
Score = 26.6 bits (57), Expect = 3.0
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +2
Query: 75 AILRPWRIPQVSSGLLK 91
AIL PW+IPQ S+ L+K
Sbjct: 152 AILAPWKIPQSSA*LIK 202
>BP048622
Length = 497
Score = 26.6 bits (57), Expect = 3.0
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = -2
Query: 78 RPWRIPQVSSGLLKMGIRG 96
RPW+ P +SS LK +RG
Sbjct: 439 RPWKAPWMSSDALKKSLRG 383
>TC18364 similar to GB|AAP21240.1|30102644|BT006432 At1g30540 {Arabidopsis
thaliana;}, partial (5%)
Length = 408
Score = 26.2 bits (56), Expect = 3.9
Identities = 17/58 (29%), Positives = 30/58 (51%)
Frame = +1
Query: 137 KDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGEQAERMAGGLTDALSV 194
+++ + +I+ M+ R GEI D + ++PVSD GE G + GG T + +
Sbjct: 208 QEKEQLLIDAEMKRYRNGEIWDFEDDIMPVSD------GETGGVLLGLDGGTTSTVCI 363
>AV780786
Length = 549
Score = 25.8 bits (55), Expect = 5.2
Identities = 14/48 (29%), Positives = 26/48 (54%), Gaps = 1/48 (2%)
Frame = +2
Query: 30 IRKRFGDSSHRNVVLKSNGNAS-ILPKIRAQNLPDYVPESKFYKVEAI 76
+R+R H +++ GNA+ IRA LP YVP+ + + ++ +
Sbjct: 341 LRERVSRVQHLSIIRHRVGNANNTYSGIRAVKLPLYVPQRRRFLLKGL 484
>BP033980
Length = 615
Score = 25.8 bits (55), Expect = 5.2
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 15/71 (21%)
Frame = +2
Query: 12 GLNFHINETQFPFSS-----FSVIRK----------RFGDSSHRNVVLKSNGNASILPKI 56
G + H+ FPF + S++RK F +S N + S+ L I
Sbjct: 329 GADKHVAVLAFPFGTHAPPLLSLVRKIAAEAPEATFSFFSTSRSNASVFSSLKEEELHNI 508
Query: 57 RAQNLPDYVPE 67
+A N+PD +PE
Sbjct: 509 KAYNVPDGLPE 541
>TC9529 similar to UP|O48794 (O48794) F24O1.3, partial (9%)
Length = 614
Score = 25.8 bits (55), Expect = 5.2
Identities = 10/25 (40%), Positives = 18/25 (72%)
Frame = -3
Query: 8 NVFNGLNFHINETQFPFSSFSVIRK 32
++ +G+N+H+ +F FSSF V+ K
Sbjct: 291 SLIHGINYHLLYLKFIFSSFVVLGK 217
>AW719523
Length = 543
Score = 25.0 bits (53), Expect = 8.8
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = +2
Query: 144 INKIMETARTGEIGDGKIFLIPVSD 168
I+++ TAR G+ G +FL+P D
Sbjct: 32 IHRVGRTARLGKQGHAVVFLLPKED 106
>TC18350 similar to UP|O81911 (O81911) T7I23.17 protein, partial (44%)
Length = 628
Score = 25.0 bits (53), Expect = 8.8
Identities = 15/67 (22%), Positives = 32/67 (47%), Gaps = 8/67 (11%)
Frame = -3
Query: 35 GDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYK--------VEAILRPWRIPQVS 86
G SS +++++ +GN ++L + +L + ++ + A LRPW++
Sbjct: 443 GSSSSVSMMMEGSGNLTLLMRRYFMHLASLMHPLSSWREYLYEMPTITAFLRPWKLGGGP 264
Query: 87 SGLLKMG 93
G K+G
Sbjct: 263 VGAWKLG 243
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.137 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,433,497
Number of Sequences: 28460
Number of extensions: 26301
Number of successful extensions: 177
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 176
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 176
length of query: 194
length of database: 4,897,600
effective HSP length: 85
effective length of query: 109
effective length of database: 2,478,500
effective search space: 270156500
effective search space used: 270156500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC135465.2


