

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135465.11 - phase: 0
(295 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
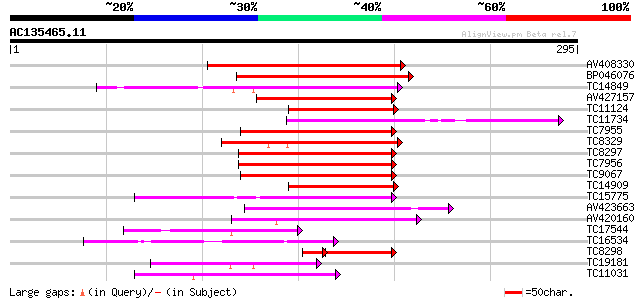
Score E
Sequences producing significant alignments: (bits) Value
AV408330 176 3e-45
BP046076 109 5e-25
TC14849 similar to UP|Q40090 (Q40090) SPF1 protein, partial (37%) 107 2e-24
AV427157 106 4e-24
TC11124 similar to UP|Q8H9D7 (Q8H9D7) WRKY-type DNA binding prot... 94 2e-20
TC11734 similar to UP|WR49_ARATH (Q9FHR7) Probable WRKY transcri... 92 1e-19
TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcrip... 89 1e-18
TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription fact... 88 2e-18
TC8297 similar to UP|Q40828 (Q40828) WRKY3, partial (27%) 85 1e-17
TC7956 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-bi... 85 1e-17
TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcrip... 84 3e-17
TC14909 similar to UP|WR21_ARATH (O04336) Probable WRKY transcri... 84 3e-17
TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcri... 83 6e-17
AV423663 79 7e-16
AV420160 79 9e-16
TC17544 similar to PIR|T04919|T04919 DNA-binding protein homolog... 71 2e-13
TC16534 similar to UP|WR41_ARATH (Q8H0Y8) Probable WRKY transcri... 57 3e-09
TC8298 homologue to UP|Q9SXP4 (Q9SXP4) DNA-binding protein NtWRK... 48 2e-08
TC19181 similar to UP|Q40090 (Q40090) SPF1 protein, partial (19%) 54 4e-08
TC11031 homologue to UP|WR65_ARATH (Q9LP56) Probable WRKY transc... 52 1e-07
>AV408330
Length = 316
Score = 176 bits (447), Expect = 3e-45
Identities = 81/103 (78%), Positives = 91/103 (87%)
Frame = +3
Query: 104 NKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPF 163
++T D +N+ KQLKAKKTNQK+ RE R AFMTKSEVDHLEDGYRWRKYGQKAVKNSPF
Sbjct: 6 DQTEDDEHNKTKKQLKAKKTNQKRQREPRFAFMTKSEVDHLEDGYRWRKYGQKAVKNSPF 185
Query: 164 PRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIM 206
PRSYYRCTS SCNVKK VERS +DP+IVVTTYEG+HTH +P+M
Sbjct: 186 PRSYYRCTSASCNVKKRVERSFTDPSIVVTTYEGQHTHSSPVM 314
>BP046076
Length = 593
Score = 109 bits (273), Expect = 5e-25
Identities = 53/92 (57%), Positives = 64/92 (68%)
Frame = -2
Query: 119 KAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVK 178
K+K ++K RE R F T+S+VD L+DGY+WRKYGQK VKNS PRSYYRCT +C VK
Sbjct: 535 KSKVKVRRKLREPRFCFQTRSDVDVLDDGYKWRKYGQKVVKNSLHPRSYYRCTHSNCRVK 356
Query: 179 KHVERSLSDPTIVVTTYEGKHTHPNPIMSRSS 210
K VER D +V+TTYEG+H H S SS
Sbjct: 355 KRVERLSEDCRMVITTYEGRHNHSPCDESNSS 260
>TC14849 similar to UP|Q40090 (Q40090) SPF1 protein, partial (37%)
Length = 1389
Score = 107 bits (267), Expect = 2e-24
Identities = 65/173 (37%), Positives = 94/173 (53%), Gaps = 14/173 (8%)
Frame = +2
Query: 46 NNYSDSLLDLPVVVKEPPLESDGNGKEYSEVLNSQQQP-ATPNSSSISSASSEAINDEHN 104
N+ S S L +P + +D + Y+ + Q ATP +SSIS + ++ +
Sbjct: 80 NSSSSSSLPIP---HSNHISNDIQDQSYATHGSGQMDSVATPENSSISIGDDDF--EQSS 244
Query: 105 KTVDQTNNQLN------KQLKAKKTNQ-------KKPREARIAFMTKSEVDHLEDGYRWR 151
+ ++L+ K+ K + N+ + RE R+ T S++D L+DGYRWR
Sbjct: 245 QKCRSGGDELDEDEPDSKRWKVEGENEGISAPGSRTVREPRVVVQTTSDIDILDDGYRWR 424
Query: 152 KYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNP 204
KYGQK VK +P PRSYY+CT C V+KHVER+ D V+TTYEGKH H P
Sbjct: 425 KYGQKVVKGNPNPRSYYKCTHPGCPVRKHVERASHDLRAVITTYEGKHNHDVP 583
Score = 33.5 bits (75), Expect = 0.044
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = +2
Query: 183 RSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNY 234
RSL D I Y+G H HP P +R ++ + SL P + +N DQ+Y
Sbjct: 2 RSL-DGQITEIVYKGTHNHPKPQATRRNSSSSSSLPIPHSNHISNDIQDQSY 154
>AV427157
Length = 379
Score = 106 bits (265), Expect = 4e-24
Identities = 47/73 (64%), Positives = 55/73 (74%)
Frame = +3
Query: 129 REARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDP 188
RE R+ T SEVD L+DGYRWRKYGQK VK +P PRSYY+CT+ C V+KHVER+ D
Sbjct: 159 REPRVVVQTTSEVDILDDGYRWRKYGQKVVKGNPNPRSYYKCTNAGCMVRKHVERASHDL 338
Query: 189 TIVVTTYEGKHTH 201
V+TTYEGKH H
Sbjct: 339 KSVITTYEGKHNH 377
>TC11124 similar to UP|Q8H9D7 (Q8H9D7) WRKY-type DNA binding protein,
partial (45%)
Length = 517
Score = 94.4 bits (233), Expect = 2e-20
Identities = 42/57 (73%), Positives = 45/57 (78%)
Frame = +1
Query: 146 DGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHP 202
DGYRWRKYGQKAVKN+ FPRSYYRCT C+VKK V R D +VVTTYEG HTHP
Sbjct: 1 DGYRWRKYGQKAVKNNKFPRSYYRCTHHGCHVKKPVPRLTQDEGVVVTTYEGVHTHP 171
>TC11734 similar to UP|WR49_ARATH (Q9FHR7) Probable WRKY transcription
factor 49 (WRKY DNA-binding protein 49), partial (28%)
Length = 1009
Score = 91.7 bits (226), Expect = 1e-19
Identities = 53/145 (36%), Positives = 73/145 (49%), Gaps = 1/145 (0%)
Frame = +3
Query: 145 EDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTH-PN 203
+DGY+WRKYGQK++KNSP PRSYYRCT+ C+ KK VERS DP ++ TYEG H H
Sbjct: 333 DDGYKWRKYGQKSIKNSPNPRSYYRCTNPRCSAKKQVERSNEDPETLIITYEGLHLHFAY 512
Query: 204 PIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMN 263
P ++ S PP + + F S Q Q Q + N +
Sbjct: 513 PCFLVGQPQQSQS--HPPVK-KSKFTSSQ-----VQAQAQAQDFIVNEAQASASVGLTSP 668
Query: 264 ATFSQDRPLCNPRVQDNGLLQDVVP 288
+ + + + GLL+D+VP
Sbjct: 669 TLLNSPQDMAQQNLGSQGLLEDMVP 743
>TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (27%)
Length = 579
Score = 88.6 bits (218), Expect = 1e-18
Identities = 37/82 (45%), Positives = 53/82 (64%), Gaps = 1/82 (1%)
Frame = +3
Query: 121 KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSV-SCNVKK 179
KK + + R+ ++ D D Y WRKYGQK +K SP+PR YY+C++V C +K
Sbjct: 18 KKRKNRVKKTVRVPAISSKTADIPPDEYSWRKYGQKPIKGSPYPRGYYKCSTVRGCPARK 197
Query: 180 HVERSLSDPTIVVTTYEGKHTH 201
HVER+ DPT+++ TYEG+H H
Sbjct: 198 HVERATDDPTMLIVTYEGEHRH 263
>TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription factor 1,
partial (41%)
Length = 1717
Score = 87.8 bits (216), Expect = 2e-18
Identities = 43/104 (41%), Positives = 66/104 (63%), Gaps = 10/104 (9%)
Frame = +1
Query: 111 NNQLNKQLKAKKTNQKKPREARI------AFMTKSEVDH---LEDGYRWRKYGQKAVKNS 161
NN ++ + + KKPRE I A++ D ++DGY+WRKYGQK +++
Sbjct: 454 NNGNSESSSTDEDSFKKPREETIKAKISRAYVRTEATDTGFVVKDGYQWRKYGQKVTRDN 633
Query: 162 PFPRSYYRCT-SVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNP 204
P PR+Y++C+ + SC VKK V+RS+ D +++V TYEG+H HP P
Sbjct: 634 PCPRAYFKCSFAPSCPVKKKVQRSVDDQSVLVATYEGEHNHPQP 765
>TC8297 similar to UP|Q40828 (Q40828) WRKY3, partial (27%)
Length = 617
Score = 85.1 bits (209), Expect = 1e-17
Identities = 36/83 (43%), Positives = 53/83 (63%), Gaps = 1/83 (1%)
Frame = +1
Query: 120 AKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTS-VSCNVK 178
+KK + R R+ ++ D D ++WRKYGQK +K SP+PR YY+C+S C +
Sbjct: 94 SKKKKLRVKRTIRVPAISSKTADIPTDDFQWRKYGQKPIKGSPYPRGYYKCSSEKGCPAR 273
Query: 179 KHVERSLSDPTIVVTTYEGKHTH 201
KHVER+ DP +++ TYEG+H H
Sbjct: 274 KHVERAQDDPNMLIVTYEGEHRH 342
>TC7956 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-binding
protein {Arabidopsis thaliana;} , partial (45%)
Length = 1257
Score = 85.1 bits (209), Expect = 1e-17
Identities = 36/83 (43%), Positives = 53/83 (63%), Gaps = 1/83 (1%)
Frame = +3
Query: 120 AKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSV-SCNVK 178
+KK + + R+ ++ D D Y WRKYGQK +K SP+PR YY+C++V C +
Sbjct: 576 SKKRKSRVKQTIRVPAVSSKIADIPPDEYSWRKYGQKPIKGSPYPRGYYKCSTVRGCPAR 755
Query: 179 KHVERSLSDPTIVVTTYEGKHTH 201
KHVER+ DP +++ TYEG+H H
Sbjct: 756 KHVERAQDDPNMLIVTYEGEHRH 824
>TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcription
factor 15 (WRKY DNA-binding protein 15), partial (31%)
Length = 660
Score = 84.0 bits (206), Expect = 3e-17
Identities = 38/82 (46%), Positives = 50/82 (60%), Gaps = 1/82 (1%)
Frame = +3
Query: 121 KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSV-SCNVKK 179
K + R R+ ++ D D Y WRKYGQK +K SP PR YY+C+SV C +K
Sbjct: 51 KSRKMRLKRVVRVPAISLKMADIPPDDYSWRKYGQKPIKGSPHPRGYYKCSSVRGCPARK 230
Query: 180 HVERSLSDPTIVVTTYEGKHTH 201
HVER+L D ++V TYEG+H H
Sbjct: 231 HVERALDDAAMLVVTYEGEHNH 296
>TC14909 similar to UP|WR21_ARATH (O04336) Probable WRKY transcription
factor 21 (WRKY DNA-binding protein 21), partial (22%)
Length = 590
Score = 84.0 bits (206), Expect = 3e-17
Identities = 34/58 (58%), Positives = 44/58 (75%), Gaps = 1/58 (1%)
Frame = +3
Query: 146 DGYRWRKYGQKAVKNSPFPRSYYRCTSV-SCNVKKHVERSLSDPTIVVTTYEGKHTHP 202
D Y WRKYGQK +K SP PR YY+C+S+ C +KHVER L +PT+++ TYEG+H HP
Sbjct: 63 DDYSWRKYGQKPIKGSPHPRGYYKCSSMRGCPARKHVERCLEEPTMLLVTYEGEHNHP 236
>TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcription
factor 17 (WRKY DNA-binding protein 17), partial (53%)
Length = 1284
Score = 82.8 bits (203), Expect = 6e-17
Identities = 47/137 (34%), Positives = 71/137 (51%), Gaps = 1/137 (0%)
Frame = +3
Query: 66 SDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQ 125
S NGK S + + A S+ S+ EH+ +N + + K +K
Sbjct: 510 SVSNGKLGSSIFLTPAVSAGGKPPLSSTTPSKKRCHEHSGDPSGSNTKCHCT-KRRKNRV 686
Query: 126 KKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSV-SCNVKKHVERS 184
KK R+ ++ D D Y WRKYGQK +K +P+PR YY+C++V C +KHVER+
Sbjct: 687 KKT--IRVPAVSSKVADIPADEYSWRKYGQKPIKGTPYPRGYYKCSTVRGCPARKHVERA 860
Query: 185 LSDPTIVVTTYEGKHTH 201
DP +++ TYE +H H
Sbjct: 861 TDDPAMLIVTYEDEHDH 911
>AV423663
Length = 447
Score = 79.3 bits (194), Expect = 7e-16
Identities = 44/111 (39%), Positives = 61/111 (54%), Gaps = 2/111 (1%)
Frame = +1
Query: 123 TNQKKPREARIAFMTKSEVDHLE-DGYRWRKYGQKAVKNSPFPRSYYRC-TSVSCNVKKH 180
T + K R+ + + + ++L D + WRKYGQK +K SP+PR YYRC +S C +K
Sbjct: 10 TPRSKRRKNQFKKVCQVSAENLSSDIWAWRKYGQKPIKGSPYPRGYYRCSSSKGCLARKQ 189
Query: 181 VERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASD 231
VER+ S+P + + TY G+H HP P S AGS P T A D
Sbjct: 190 VERNKSEPAMFIVTYTGEHNHPAPTHRNS---LAGSTRQKPLTPETATAGD 333
>AV420160
Length = 417
Score = 79.0 bits (193), Expect = 9e-16
Identities = 45/112 (40%), Positives = 60/112 (53%), Gaps = 13/112 (11%)
Frame = +1
Query: 116 KQLKAKKTNQKKPREARIAFMT-----------KSEVDHLEDGYRWRKYGQKAVKNSPFP 164
+ +K + + KK RE + +T K E D + WRKYGQK +K SP+P
Sbjct: 34 EDIKTEAPSPKKRREMKKRVVTLPIGDVDGSKSKGETYPPSDSWAWRKYGQKPIKGSPYP 213
Query: 165 RSYYRC-TSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPI-MSRSSAVRA 214
R YYRC +S C +K VERS DPT ++ TY +H H P+ S SSA A
Sbjct: 214 RGYYRCSSSKGCPARKQVERSRVDPTKLMVTYSYEHNHSLPVPKSHSSAASA 369
>TC17544 similar to PIR|T04919|T04919 DNA-binding protein homolog T9A21.10 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(14%)
Length = 578
Score = 70.9 bits (172), Expect = 2e-13
Identities = 45/98 (45%), Positives = 56/98 (56%), Gaps = 5/98 (5%)
Frame = +1
Query: 60 KEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQL----- 114
K P + G G E L + T NSS +S+SS+A +E + + NNQ+
Sbjct: 295 KPPEGDVAGGGGGGGETLAT----TTLNSSISASSSSDAGAEEDSGKKKKDNNQVKEEGN 462
Query: 115 NKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRK 152
N K KK +KK +E R AFMTKSEVDHLEDGYRWRK
Sbjct: 463 NSGNKDKKKGEKKQKEPRFAFMTKSEVDHLEDGYRWRK 576
>TC16534 similar to UP|WR41_ARATH (Q8H0Y8) Probable WRKY transcription
factor 41 (WRKY DNA-binding protein 41), partial (20%)
Length = 697
Score = 57.4 bits (137), Expect = 3e-09
Identities = 42/133 (31%), Positives = 63/133 (46%)
Frame = +1
Query: 39 LGVQQNMNNYSDSLLDLPVVVKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEA 98
L VQ+ +++Y +L L L++ NG + L S+ P + N S + A
Sbjct: 310 LQVQRILSSYEKALQILRWNESTSKLQTT-NG---ATTLLSESPPLSANGSPMREEIDWA 477
Query: 99 INDEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAV 158
I D + + + K +KT K + R+ E H +DGY WRKYGQK +
Sbjct: 478 IQD---------HQEAKRDSKKRKTMPKWIDQIRVVSENGLEGPH-DDGYNWRKYGQKDI 627
Query: 159 KNSPFPRSYYRCT 171
+ +PRSYYRCT
Sbjct: 628 LGAKYPRSYYRCT 666
>TC8298 homologue to UP|Q9SXP4 (Q9SXP4) DNA-binding protein NtWRKY3,
partial (16%)
Length = 514
Score = 48.1 bits (113), Expect(2) = 2e-08
Identities = 19/38 (50%), Positives = 27/38 (71%), Gaps = 1/38 (2%)
Frame = +3
Query: 165 RSYYRCTSVS-CNVKKHVERSLSDPTIVVTTYEGKHTH 201
R YY+C+S C +KHVER+ DP +++ TYEG+H H
Sbjct: 159 RGYYKCSSEKGCPARKHVERAQDDPNMLIVTYEGEHRH 272
Score = 26.2 bits (56), Expect(2) = 2e-08
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +1
Query: 153 YGQKAVKNSPFPR 165
YGQK +K SP+PR
Sbjct: 1 YGQKPIKGSPYPR 39
>TC19181 similar to UP|Q40090 (Q40090) SPF1 protein, partial (19%)
Length = 402
Score = 53.5 bits (127), Expect = 4e-08
Identities = 30/99 (30%), Positives = 51/99 (51%), Gaps = 10/99 (10%)
Frame = +3
Query: 74 SEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQ---LNKQLKAKKTNQ----- 125
+ V ATP +SS+S + +K+ + ++ K+ + + N+
Sbjct: 102 THVSGQMDSVATPENSSLSMGGDDFEQSSMSKSGGEELDEDEPHGKKWRMEGENEGMSAA 281
Query: 126 --KKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSP 162
+ +E+R+ T S++D L+DGYRWRKYGQK VK +P
Sbjct: 282 GSRTVKESRVVVQTTSDIDILDDGYRWRKYGQKVVKGNP 398
>TC11031 homologue to UP|WR65_ARATH (Q9LP56) Probable WRKY transcription
factor 65 (WRKY DNA-binding protein 65), partial (15%)
Length = 453
Score = 52.0 bits (123), Expect = 1e-07
Identities = 33/110 (30%), Positives = 55/110 (50%), Gaps = 3/110 (2%)
Frame = +3
Query: 66 SDGNGKEYSEVLNSQQQPATPNSSSISSA--SSEAINDEHNKTVDQTNNQLNKQLKAKKT 123
S+ + +E+L + A + +I+ + SS A N + T T++ + +K
Sbjct: 102 SNNRHRYINELLEEDAEVAHHHQENIADSPPSSTAFNIDGLVTSSSTSSAKRSRRAIQKR 281
Query: 124 NQKKPREARIAFMTKSEVDHL-EDGYRWRKYGQKAVKNSPFPRSYYRCTS 172
+ P + K+E + D + WRKYGQK +K SP+PR YYRC+S
Sbjct: 282 VVQIPIKETEGSRLKAENNTPPSDSWAWRKYGQKPIKGSPYPRGYYRCSS 431
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,112,744
Number of Sequences: 28460
Number of extensions: 73865
Number of successful extensions: 422
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 408
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 412
length of query: 295
length of database: 4,897,600
effective HSP length: 90
effective length of query: 205
effective length of database: 2,336,200
effective search space: 478921000
effective search space used: 478921000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC135465.11


