

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
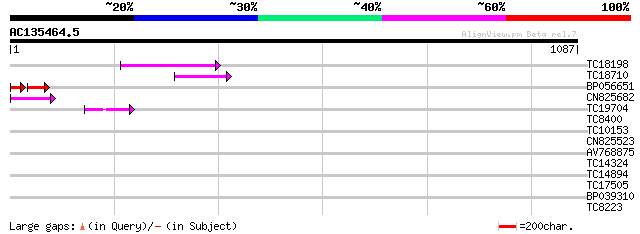
Query= AC135464.5 - phase: 0 /pseudo
(1087 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 98 6e-21
TC18710 78 7e-15
BP056651 48 7e-09
CN825682 52 4e-07
TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA po... 50 2e-06
TC8400 similar to UP|PIRL_LYCES (Q9SEE4) Pirin-like protein, par... 32 0.54
TC10153 UP|NUKC_LOTJA (Q9BBT7) NAD(P)H-quinone oxidoreductase ch... 30 2.1
CN825523 30 2.1
AV768875 30 2.1
TC14324 weakly similar to UP|Q8LEE9 (Q8LEE9) Arabinogalactan pro... 28 6.0
TC14894 weakly similar to UP|Q9LD82 (Q9LD82) ESTs AU029648, part... 28 6.0
TC17505 similar to UP|Q9SB33 (Q9SB33) SRG1-like protein, partial... 28 7.8
BP039310 28 7.8
TC8223 UP|AAQ87021 (AAQ87021) VDAC1.3, complete 28 7.8
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 98.2 bits (243), Expect = 6e-21
Identities = 64/194 (32%), Positives = 102/194 (51%), Gaps = 2/194 (1%)
Frame = -3
Query: 212 RLLWLKDGDANSKYFHSVLSGRRRRNSIISLL-ADSNLVEGVQPIRNAVLSHFRDHFAAR 270
R+ LK GD N+++FH+ RR N I+ + A VEG + + A +++F D +++
Sbjct: 590 RIK*LK*GDHNTRFFHASTIQRRDFNRILKIKDAHGIWVEGQKKVNEAAVNYFSDIYSSS 411
Query: 271 HTSRVGIDNLPF-KCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFW 329
+ P K ++ L S DE+K AV+ K+PG DG+N F ++
Sbjct: 410 PLQYLRECLAPIPKLVTDRLNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQQNR 231
Query: 330 DELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKV 389
+ +K DV I DF +G+L +N T + LI KI + I FRPIS +YK++SK+
Sbjct: 230 EIVKNDVNNAILDFFDHGELPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKI 51
Query: 390 LANRLKLVLGKIIS 403
+ RLK + +IS
Sbjct: 50 IVARLKPDMVNLIS 9
>TC18710
Length = 843
Score = 78.2 bits (191), Expect = 7e-15
Identities = 43/109 (39%), Positives = 62/109 (56%)
Frame = +2
Query: 316 GPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRP 375
G DG+N F ++ D +K+ V I F ++ IN T + LI K+ P+ IN FRP
Sbjct: 188 GMDGMNGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETLVTLIPKVPHPESINQFRP 367
Query: 376 ISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANE 424
IS LYK++SK+ RLK + +IS Q+ F++ RQI D +LI E
Sbjct: 368 ISGCTFLYKVISKIFVARLKDAMPDLISPMQSGFIQGRQIQDNLLIVQE 514
>BP056651
Length = 564
Score = 47.8 bits (112), Expect(2) = 7e-09
Identities = 19/42 (45%), Positives = 28/42 (66%)
Frame = +3
Query: 35 LIDLSLCGRRFTWFKGDGNAMSRIDRFLLSEEWCLQWPNSFQ 76
L D+ L GR++TW + +G A SR+DRFL+S +W W + Q
Sbjct: 432 LDDIPLVGRKYTWVRSNGKARSRLDRFLVSHQWNTSWXDCAQ 557
Score = 30.0 bits (66), Expect(2) = 7e-09
Identities = 15/32 (46%), Positives = 21/32 (64%), Gaps = 2/32 (6%)
Frame = +1
Query: 1 DFNAVRYAEERKSS--RGISGVIDYHHFSDFI 30
DFN+VR +EER+ + G SG + F+DFI
Sbjct: 325 DFNSVRRSEERRGTHGEGSSGAGEMEEFNDFI 420
>CN825682
Length = 626
Score = 52.4 bits (124), Expect = 4e-07
Identities = 32/91 (35%), Positives = 48/91 (52%), Gaps = 3/91 (3%)
Frame = -3
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGNAM---SR 57
DFN + + +E++ +R F DF+ +L+DL L G RFTW N + R
Sbjct: 285 DFNEILHHQEKEGARD-QDPTRIQVFRDFLDGANLMDLELKGCRFTWLSNLRNGLVTKER 109
Query: 58 IDRFLLSEEWCLQWPNSFQVALLRGLSDHCP 88
+DR L++ W L +PN+ +AL SDH P
Sbjct: 108 LDRVLVNWTWRLDYPNATAMALPIISSDHSP 16
>TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA
polymerase-like protein, partial (3%)
Length = 530
Score = 50.1 bits (118), Expect = 2e-06
Identities = 32/96 (33%), Positives = 46/96 (47%)
Frame = -3
Query: 144 KTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLHSLSRVN 203
K AL EWH + S +I LK + + + G+ + E++ L L R
Sbjct: 435 KLALTEWHHQNFSRADKEIAELKNQLEHLQNYSNSGE----DWQEIKQRKMRLKDLWRQE 268
Query: 204 SSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSI 239
Q+SR+ WLK GD N+K+FH+ RR RN I
Sbjct: 267 ELFWSQRSRIKWLKSGDKNTKFFHASTVQRRERNRI 160
>TC8400 similar to UP|PIRL_LYCES (Q9SEE4) Pirin-like protein, partial (93%)
Length = 1352
Score = 32.0 bits (71), Expect = 0.54
Identities = 23/85 (27%), Positives = 40/85 (47%), Gaps = 6/85 (7%)
Frame = +1
Query: 246 SNLVEGVQPIRNAVLSHFRDHFAARHTSRVGIDNLPFKCLSHTDGSGLTKPFSADEVKS- 304
+N+ + P+RN ++S DH A +T R+ + + K DG+ + + E+K+
Sbjct: 115 TNIRRSLPPVRNIIMSSESDHSPAFNTPRLVLKKVLAKSQREGDGAVVRRGIGRSELKNL 294
Query: 305 ---AVWDCDSYKSPG--PDGVNFGF 324
+ D S PG PD + GF
Sbjct: 295 DPFLMLDHFSVSPPGGFPDHPHRGF 369
>TC10153 UP|NUKC_LOTJA (Q9BBT7) NAD(P)H-quinone oxidoreductase chain K,
chloroplast (NAD(P)H dehydrogenase, chain K)
(NADH-plastoquinone oxidoreductase subunit K) , complete
Length = 1317
Score = 30.0 bits (66), Expect = 2.1
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +1
Query: 905 LLHWSLQTFLWETLYGISRCRLRFPFLLG 933
L +WS + LW LYG S C + F L+G
Sbjct: 85 LSNWSRLSSLWPLLYGTSCCFIEFASLIG 171
>CN825523
Length = 611
Score = 30.0 bits (66), Expect = 2.1
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +1
Query: 736 VGGCWWIEADFGIGSW*RGM 755
+G CWW D GIG+W +G+
Sbjct: 193 LGSCWWK*EDKGIGAWNQGL 252
>AV768875
Length = 232
Score = 30.0 bits (66), Expect = 2.1
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +1
Query: 26 FSDFIVDNSLIDLSLCGRRFTWFKGDGNAM 55
F+ + D +LIDL + G +FTWF+ N +
Sbjct: 142 FATVLDDCNLIDLGMVGGKFTWFRKRNNRL 231
>TC14324 weakly similar to UP|Q8LEE9 (Q8LEE9) Arabinogalactan protein-like,
partial (63%)
Length = 1172
Score = 28.5 bits (62), Expect = 6.0
Identities = 21/63 (33%), Positives = 33/63 (52%), Gaps = 1/63 (1%)
Frame = +3
Query: 159 PGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLH-SLSRVNSSILWQQSRLLWLK 217
P K + KK+ AD D +G GD SA ++ R + L ++ V + W Q ++ W K
Sbjct: 687 PEKAKASKKKSAD-DTEGPAGDDSAAVSVKQRLVWGPLPVAMVVVAAFSSW*QWKIEWFK 863
Query: 218 DGD 220
DG+
Sbjct: 864 DGE 872
>TC14894 weakly similar to UP|Q9LD82 (Q9LD82) ESTs AU029648, partial (84%)
Length = 982
Score = 28.5 bits (62), Expect = 6.0
Identities = 19/52 (36%), Positives = 27/52 (51%), Gaps = 3/52 (5%)
Frame = -2
Query: 41 CGRRFTWFKGDGNAMSRIDRFL-LSEEWCLQWPN--SFQVALLRGLSDHCPL 89
C R F + N ++ ++FL L +E CL+W N S Q+ G HCPL
Sbjct: 318 CSRSFLEY----N*IASRNQFL*LQQEKCLEWGNLRSTQLNYSDGFHSHCPL 175
>TC17505 similar to UP|Q9SB33 (Q9SB33) SRG1-like protein, partial (12%)
Length = 538
Score = 28.1 bits (61), Expect = 7.8
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -1
Query: 895 IVWEAPIFILLLHWSLQTFL 914
I W P F+LL HW + FL
Sbjct: 424 IFWRLPKFLLLFHWKTKEFL 365
>BP039310
Length = 584
Score = 28.1 bits (61), Expect = 7.8
Identities = 20/45 (44%), Positives = 21/45 (46%), Gaps = 3/45 (6%)
Frame = -3
Query: 737 GGCWWIEADFGIG---SW*RGMGRRLGGWR*GAGVFHLGGEK*RR 778
GGC W FG+G SW R L G G G GGEK RR
Sbjct: 156 GGCSWS*GRFGVGSRRSWAAE*ARNLAGI--GGGRHGGGGEKRRR 28
>TC8223 UP|AAQ87021 (AAQ87021) VDAC1.3, complete
Length = 1261
Score = 28.1 bits (61), Expect = 7.8
Identities = 21/59 (35%), Positives = 25/59 (41%)
Frame = +3
Query: 959 VSQGAVWKRLHLIFLFIAPLLVLCGILSGLGSECQVLTLLLLRPISFSLFTLQVTQRLV 1017
VS +W L IF I PL L + S L LLL I +L T +T LV
Sbjct: 426 VSLEIIWSLLGQIFHLILPLATLPSTMQD*TSRMLTLLHLLL*MIKVTLLTPHITMLLV 602
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.344 0.155 0.538
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,291,363
Number of Sequences: 28460
Number of extensions: 369693
Number of successful extensions: 3403
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 3340
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3402
length of query: 1087
length of database: 4,897,600
effective HSP length: 100
effective length of query: 987
effective length of database: 2,051,600
effective search space: 2024929200
effective search space used: 2024929200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135464.5


