

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
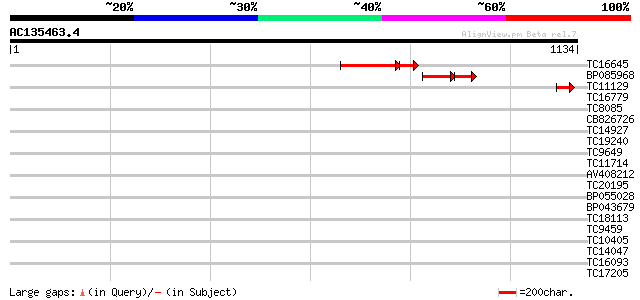
Query= AC135463.4 - phase: 2 /pseudo
(1134 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16645 110 4e-33
BP085968 64 3e-19
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 49 6e-06
TC16779 similar to UP|Q9LEY9 (Q9LEY9) Nhp2-like protein (AT5g081... 39 0.004
TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meioti... 34 0.15
CB826726 33 0.33
TC14927 weakly similar to GB|BAC16200.1|22831356|AP005486 transp... 32 0.74
TC19240 similar to UP|Q94BY4 (Q94BY4) AT5g35570/K2K18_1, partial... 31 0.96
TC9649 similar to UP|Q95TP4 (Q95TP4) LD33277p, partial (4%) 31 0.96
TC11714 similar to UP|Q16861 (Q16861) Super cysteine rich protei... 30 1.6
AV408212 30 1.6
TC20195 similar to UP|Q8IQS0 (Q8IQS0) CG32193-PA, partial (5%) 30 2.1
BP055028 30 2.1
BP043679 30 2.8
TC18113 27 3.2
TC9459 similar to AAQ23982 (AAQ23982) Transcription factor Rap21... 29 3.7
TC10405 similar to UP|AAP51013 (AAP51013) 26K protein, partial (9%) 29 3.7
TC14047 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {... 29 3.7
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 29 4.8
TC17205 weakly similar to PIR|S50830|S50830 Machado-Joseph disea... 28 6.2
>TC16645
Length = 596
Score = 110 bits (276), Expect(2) = 4e-33
Identities = 54/119 (45%), Positives = 77/119 (64%)
Frame = +1
Query: 662 MDANRNYSQSWNLTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLL 721
M+AN+ Y + NLTY +FP +F Y+ W + S+GRL F+ PG E Y MR+LL
Sbjct: 7 MNANKKYQEGRNLTYAEFPSEFLYHQHTTKWVPKQKRFSLGRLPFIAPGMGENYYMRVLL 186
Query: 722 NRQIGCTSFEDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVF 780
Q GC SF+ I+TV G VY ++ +AC A+GLL +DR+++D IS GSG+ +RK+F
Sbjct: 187 TMQKGCDSFKSIKTVKGVVYPTFHDACEAMGLLEDDREYVDGISISSEFGSGTQLRKLF 363
Score = 48.9 bits (115), Expect(2) = 4e-33
Identities = 19/37 (51%), Positives = 29/37 (78%)
Frame = +2
Query: 780 FTSLLLTSCMSDPLNVWNQTWHILSEGILYERRRTLN 816
FT +L+T+ +S P VW ++W +L +GILY+RR+TLN
Sbjct: 362 FTRMLMTNLISRPQEVWRKSWSLLCDGILYDRRKTLN 472
>BP085968
Length = 341
Score = 63.5 bits (153), Expect(2) = 3e-19
Identities = 32/65 (49%), Positives = 45/65 (69%)
Frame = +2
Query: 826 QLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLL 885
Q Y++I++ V ++ +F+ G+GG GKTF+WN S RS+G +VLNVASS IAS L
Sbjct: 20 QSKVYKQIMSSVWSDDGGFYFLYGFGGSGKTFVWNTWSSGLRSQGLMVLNVASSGIASWL 199
Query: 886 LPRGR 890
LP G+
Sbjct: 200 LPGGK 214
Score = 49.7 bits (117), Expect(2) = 3e-19
Identities = 24/43 (55%), Positives = 31/43 (71%)
Frame = +3
Query: 890 RTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAP 932
RTAHS+FSI + + + S C I++ KA+LL AS IIWDEAP
Sbjct: 213 RTAHSRFSIHISINDISTCNIKQGSQKAELLQKASSIIWDEAP 341
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 48.5 bits (114), Expect = 6e-06
Identities = 22/36 (61%), Positives = 27/36 (74%)
Frame = +2
Query: 1094 PTLELVEKVNDYVLSLVPGEEKEYLSCDTVLKCDEE 1129
PTLE VEKVN+++L L+PG EYLS DT K DE+
Sbjct: 458 PTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDED 565
>TC16779 similar to UP|Q9LEY9 (Q9LEY9) Nhp2-like protein
(AT5g08180/T22D6_120), partial (17%)
Length = 850
Score = 39.3 bits (90), Expect = 0.004
Identities = 17/61 (27%), Positives = 30/61 (48%)
Frame = +2
Query: 409 VLMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCCYKMMFMLL*DDFYDVKIMSLYDDIYD 468
V ++ C CC ++F +CC C Y ++F++ Y L* ++ L+ I+D
Sbjct: 662 VTFLLLPFSCCCCLLILFYLCCFCLYFLLFVVFLYVFSCARL*SHLGEIVPEILFLSIHD 841
Query: 469 L 469
L
Sbjct: 842 L 844
Score = 38.5 bits (88), Expect = 0.006
Identities = 12/34 (35%), Positives = 20/34 (58%), Gaps = 1/34 (2%)
Frame = +2
Query: 417 CCLCCYKMMFM-MCCLCCYKMMFMMCCYKMMFML 449
CC CC + + C CC ++F +CC+ + F+L
Sbjct: 647 CCCCCVTFLLLPFSCCCCLLILFYLCCFCLYFLL 748
>TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meiotic
recombination protein)-like protein {Arabidopsis
thaliana;} , partial (8%)
Length = 500
Score = 33.9 bits (76), Expect = 0.15
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = +1
Query: 789 MSDPLNVWNQTWHILSEGILYERRRTLNSP 818
M++P +VWN TW +L+ I +E R TL P
Sbjct: 115 MTNPDHVWNITWKLLANDIQHEYRGTLTQP 204
>CB826726
Length = 505
Score = 32.7 bits (73), Expect = 0.33
Identities = 11/33 (33%), Positives = 16/33 (48%)
Frame = -2
Query: 411 MMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCCY 443
++ + CC CC K + CC CC CC+
Sbjct: 465 LLSHCCCCCCCAKFLCT*CCCCC-----CCCCW 382
>TC14927 weakly similar to GB|BAC16200.1|22831356|AP005486 transposase-like
{Oryza sativa (japonica cultivar-group);} , partial
(11%)
Length = 539
Score = 31.6 bits (70), Expect = 0.74
Identities = 10/34 (29%), Positives = 18/34 (52%)
Frame = +1
Query: 409 VLMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCC 442
V+ Y++CC+C Y +CC ++ + CC
Sbjct: 361 VVFAFYLVCCVC-YLFFHFLCCFLFISVVALFCC 459
>TC19240 similar to UP|Q94BY4 (Q94BY4) AT5g35570/K2K18_1, partial (12%)
Length = 554
Score = 31.2 bits (69), Expect = 0.96
Identities = 10/21 (47%), Positives = 10/21 (47%)
Frame = -1
Query: 414 YMMCCLCCYKMMFMMCCLCCY 434
Y C CCY CC CCY
Sbjct: 263 YSESCCCCYCRCCYRCCCCCY 201
>TC9649 similar to UP|Q95TP4 (Q95TP4) LD33277p, partial (4%)
Length = 754
Score = 31.2 bits (69), Expect = 0.96
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = -2
Query: 409 VLMMMYMMCCLCCYKMMFMMCCLCCY 434
++ MY CC+CC CC CC+
Sbjct: 201 IMAAMYCCCCICC-----CCCCSCCW 139
>TC11714 similar to UP|Q16861 (Q16861) Super cysteine rich protein
(Fragment), partial (48%)
Length = 519
Score = 30.4 bits (67), Expect = 1.6
Identities = 8/18 (44%), Positives = 10/18 (55%)
Frame = +2
Query: 417 CCLCCYKMMFMMCCLCCY 434
CC CC+ CC CC+
Sbjct: 83 CCCCCFSC*SCCCCCCCF 136
>AV408212
Length = 362
Score = 30.4 bits (67), Expect = 1.6
Identities = 11/32 (34%), Positives = 15/32 (46%), Gaps = 3/32 (9%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCCYKMMFM---MCCYKM 445
CC CC CC CC++ + +CC M
Sbjct: 142 CCCCCCCCCCCCCCCCCWRAACLCNRICCICM 47
Score = 29.3 bits (64), Expect = 3.7
Identities = 9/26 (34%), Positives = 13/26 (49%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCC 442
CC CC++ + +CC M CC
Sbjct: 106 CCCCCWRAACLCNRICCICMCCCCCC 29
Score = 28.1 bits (61), Expect = 8.2
Identities = 10/26 (38%), Positives = 11/26 (41%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCC 442
CC CC + CLC MCC
Sbjct: 118 CCCCCCCCCWRAACLCNRICCICMCC 41
>TC20195 similar to UP|Q8IQS0 (Q8IQS0) CG32193-PA, partial (5%)
Length = 591
Score = 30.0 bits (66), Expect = 2.1
Identities = 15/58 (25%), Positives = 30/58 (50%), Gaps = 1/58 (1%)
Frame = +1
Query: 394 LLSC*VVLYPHVVNEVLMM-MYMMCCLCCYKMMFMMCCLCCYKMMFMMCCYKMMFMLL 450
+LS V++Y V +L+ + +MCC CCY + L + F C + +++++
Sbjct: 322 MLSLLVLIYAAAVLALLLCDVAVMCCCCCYSKWRLFLLLLVAVLPFSCC*FAPIYVVV 495
Score = 28.1 bits (61), Expect = 8.2
Identities = 16/56 (28%), Positives = 24/56 (42%), Gaps = 15/56 (26%)
Frame = +2
Query: 408 EVLMMMYMMCCLCCYKMMF---------MMCCLCC-----YKMMF-MMCCYKMMFM 448
E L CCLCC +F + C CC Y ++ ++CCY M+ +
Sbjct: 224 ESLFFCCRWCCLCCC*YVFDAGAGAGAVLAGC*CCPCWC*YTLLLSLLCCYVMLLL 391
>BP055028
Length = 530
Score = 30.0 bits (66), Expect = 2.1
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = -1
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMC 441
CC CC CC CCY++ +C
Sbjct: 185 CCCCC-------CCCCCYELE*WLC 132
>BP043679
Length = 501
Score = 29.6 bits (65), Expect = 2.8
Identities = 10/27 (37%), Positives = 14/27 (51%), Gaps = 2/27 (7%)
Frame = +1
Query: 418 CLCCYKMMFMMCCLC--CYKMMFMMCC 442
C CC++ F CC C C + + CC
Sbjct: 214 CQCCWRWWFSCCCSCPSCCCIWWRFCC 294
>TC18113
Length = 543
Score = 26.6 bits (57), Expect(2) = 3.2
Identities = 7/14 (50%), Positives = 10/14 (71%)
Frame = +3
Query: 419 LCCYKMMFMMCCLC 432
LCC +F++CC C
Sbjct: 210 LCCSNWLFVLCCFC 251
Score = 21.2 bits (43), Expect(2) = 3.2
Identities = 6/22 (27%), Positives = 14/22 (63%)
Frame = +2
Query: 404 HVVNEVLMMMYMMCCLCCYKMM 425
HV++ V + ++ +CC C ++
Sbjct: 152 HVLSMVEIDIFFLCCSCLCSLL 217
>TC9459 similar to AAQ23982 (AAQ23982) Transcription factor Rap212, partial
(12%)
Length = 434
Score = 29.3 bits (64), Expect = 3.7
Identities = 10/21 (47%), Positives = 10/21 (47%)
Frame = -1
Query: 414 YMMCCLCCYKMMFMMCCLCCY 434
Y CC CC CC CCY
Sbjct: 305 YCYCCCCC-------CCCCCY 264
>TC10405 similar to UP|AAP51013 (AAP51013) 26K protein, partial (9%)
Length = 590
Score = 29.3 bits (64), Expect = 3.7
Identities = 10/28 (35%), Positives = 16/28 (56%)
Frame = +3
Query: 410 LMMMYMMCCLCCYKMMFMMCCLCCYKMM 437
L ++ ++CC C Y + + C CYK M
Sbjct: 414 LAIV*VICCCCLYWLAVFIVCFFCYKSM 497
>TC14047 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (67%)
Length = 842
Score = 29.3 bits (64), Expect = 3.7
Identities = 17/40 (42%), Positives = 23/40 (57%)
Frame = -3
Query: 768 VVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGI 807
VVVG GS+I K+FTS +PL VW +L+ G+
Sbjct: 714 VVVGQGSAIFKLFTS------KDEPLLVWWDALFVLNFGL 613
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 28.9 bits (63), Expect = 4.8
Identities = 8/17 (47%), Positives = 9/17 (52%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCC 433
CC+CC CC CC
Sbjct: 215 CCICCCGCCIGCCCCCC 165
Score = 28.5 bits (62), Expect = 6.2
Identities = 10/26 (38%), Positives = 11/26 (41%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCC 442
CC CC CC+CC CC
Sbjct: 233 CCSCC------CCCICCCGCCIGCCC 174
>TC17205 weakly similar to PIR|S50830|S50830 Machado-Joseph disease MJD1a
protein - human {Homo sapiens;}, partial (7%)
Length = 528
Score = 28.5 bits (62), Expect = 6.2
Identities = 9/17 (52%), Positives = 10/17 (57%)
Frame = -3
Query: 417 CCLCCYKMMFMMCCLCC 433
CC CCY + CC CC
Sbjct: 361 CCCCCY---WYRCCCCC 320
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.337 0.148 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,436,767
Number of Sequences: 28460
Number of extensions: 301168
Number of successful extensions: 3287
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 2746
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3167
length of query: 1134
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1034
effective length of database: 2,051,600
effective search space: 2121354400
effective search space used: 2121354400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135463.4


